Abstract
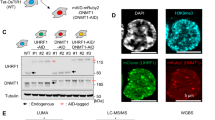
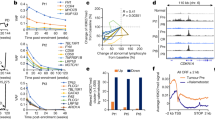
Epigenetic deregulation of gene expression has a role in the initiation and progression of prostate cancer (PCa). The histone methyltransferase MMSET/WHSC1 (Multiple Myeloma SET domain) is overexpressed in a number of metastatic tumors, but its mechanism of action has not been defined. In this work, we found that PCa cell lines expressed significantly higher levels of MMSET compared with immortalized, non-transformed prostate cells. Knockdown experiments showed that, in metastatic PCa cell lines, dimethylation of lysine 36 and trimethylation of lysine 27 on histone H3 (H3K36me2 and H3K27me3, respectively) depended on MMSET expression, whereas depletion of MMSET in benign prostatic cells did not affect chromatin modifications. Knockdown of MMSET in DU145 and PC-3 tumor cells decreased cell proliferation, colony formation in soft agar and strikingly diminished cell migration and invasion. Conversely, overexpression of MMSET in immortalized, non-transformed RWPE-1 cells promoted cell migration and invasion, accompanied by an epithelial–mesenchymal transition (EMT). Among a panel of EMT-promoting genes analyzed, TWIST1 expression was strongly activated in response to MMSET. Chromatin immunoprecipitation analysis demonstrated that MMSET binds to the TWIST1 locus and leads to an increase in H3K36me2, suggesting a direct role of MMSET in the regulation of this gene. Depletion of TWIST1 in MMSET-overexpressing RWPE-1 cells blocked cell invasion and EMT, indicating that TWIST1 was a critical target of MMSET, responsible for the acquisition of an invasive phenotype. Collectively, these data suggest that MMSET has a role in PCa pathogenesis and progression through epigenetic regulation of metastasis-related genes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Jemal A, Siegel R, Xu J, Ward E . Cancer statistics. CA Cancer J Clin 2010; 60: 277–300.
Jeronimo C, Bastian PJ, Bjartell A, Carbone GM, Catto JW, Clark SJ et al. Epigenetics in prostate cancer: biologic and clinical relevance. Eur Urol 2011; 60: 753–766.
Perry AS, Watson RW, Lawler M, Hollywood D . The epigenome as a therapeutic target in prostate cancer. Nat Rev Urol 2010; 7: 668–680.
Schulz WA, Hatina J . Epigenetics of prostate cancer: beyond DNA methylation. J Cell Mol Med 2006; 10: 100–125.
Yegnasubramanian S, Kowalski J, Gonzalgo ML, Zahurak M, Piantadosi S, Walsh PC et al. Hypermethylation of CpG islands in primary and metastatic human prostate cancer. Cancer Res 2004; 64: 1975–1986.
Jenuwein T, Allis CD . Translating the histone code. Science 2001; 293: 1074–1080.
Kouzarides T . Chromatin modifications and their function. Cell 2007; 128: 693–705.
Lennartsson A, Ekwall K . Histone modification patterns and epigenetic codes. Biochim Biophys Acta 2009; 1790: 863–868.
Mellor J . The dynamics of chromatin remodeling at promoters. Mol Cell 2005; 19: 147–157.
Halkidou K, Gaughan L, Cook S, Leung HY, Neal DE, Robson CN . Upregulation and nuclear recruitment of HDAC1 in hormone refractory prostate cancer. Prostate 2004; 59: 177–189.
Kahl P, Gullotti L, Heukamp LC, Wolf S, Friedrichs N, Vorreuther R et al. Androgen receptor coactivators lysine-specific histone demethylase 1 and four and a half LIM domain protein 2 predict risk of prostate cancer recurrence. Cancer Res 2006; 66: 11341–11347.
Metzger E, Wissmann M, Yin N, Muller JM, Schneider R, Peters AH et al. LSD1 demethylates repressive histone marks to promote androgen-receptor-dependent transcription. Nature 2005; 437: 436–439.
Varambally S, Dhanasekaran SM, Zhou M, Barrette TR, Kumar-Sinha C, Sanda MG et al. The polycomb group protein EZH2 is involved in progression of prostate cancer. Nature 2002; 419: 624–629.
Bianco-Miotto T, Chiam K, Buchanan G, Jindal S, Day TK, Thomas M et al. Global levels of specific histone modifications and an epigenetic gene signature predict prostate cancer progression and development. Cancer Epidemiol Biomarkers Prev 19: 2611–2622.
Seligson DB, Horvath S, Shi T, Yu H, Tze S, Grunstein M et al. Global histone modification patterns predict risk of prostate cancer recurrence. Nature 2005; 435: 1262–1266.
Stec I, Wright TJ, van Ommen GJ, de Boer PA, van Haeringen A, Moorman AF et al. WHSC1, a 90 kb SET domain-containing gene, expressed in early development and homologous to a Drosophila dysmorphy gene maps in the Wolf-Hirschhorn syndrome critical region and is fused to IgH in t(4;14) multiple myeloma. Hum Mol Genet 1998; 7: 1071–1082.
Volkel P, Angrand PO . The control of histone lysine methylation in epigenetic regulation. Biochimie 2007; 89: 1–20.
Kim JY, Kee HJ, Choe NW, Kim SM, Eom GH, Baek HJ et al. Multiple-myeloma-related WHSC1/MMSET isoform RE-IIBP is a histone methyltransferase with transcriptional repression activity. Mol Cell Biol 2008; 28: 2023–2034.
Martinez-Garcia E, Popovic R, Min DJ, Sweet SM, Thomas PM, Zamdborg L et al. The MMSET histone methyl transferase switches global histone methylation and alters gene expression in t(4;14) multiple myeloma cells. Blood 117: 211–220.
Li Y, Trojer P, Xu CF, Cheung P, Kuo A, Drury WJ et al. The target of the NSD family of histone lysine methyltransferases depends on the nature of the substrate. J Biol Chem 2009; 284: 34283–34295.
Chesi M, Nardini E, Lim RS, Smith KD, Kuehl WM, Bergsagel PL . The t(4;14) translocation in myeloma dysregulates both FGFR3 and a novel gene, MMSET, resulting in IgH/MMSET hybrid transcripts. Blood 1998; 92: 3025–3034.
Garlisi CG, Uss AS, Xiao H, Tian F, Sheridan KE, Wang L et al. A unique mRNA initiated within a middle intron of WHSC1/MMSET encodes a DNA binding protein that suppresses human IL-5 transcription. Am J Respir Cell Mol Biol 2001; 24: 90–98.
Keats JJ, Maxwell CA, Taylor BJ, Hendzel MJ, Chesi M, Bergsagel PL et al. Overexpression of transcripts originating from the MMSET locus characterizes all t(4;14)(p16;q32)-positive multiple myeloma patients. Blood 2005; 105: 4060–4069.
Brito JL, Walker B, Jenner M, Dickens NJ, Brown NJ, Ross FM et al. MMSET deregulation affects cell cycle progression and adhesion regulons in t(4;14) myeloma plasma cells. Haematologica 2009; 94: 78–86.
Lauring J, Abukhdeir AM, Konishi H, Garay JP, Gustin JP, Wang Q et al. The multiple myeloma associated MMSET gene contributes to cellular adhesion, clonogenic growth, and tumorigenicity. Blood 2008; 111: 856–864.
Marango J, Shimoyama M, Nishio H, Meyer JA, Min DJ, Sirulnik A et al. The MMSET protein is a histone methyltransferase with characteristics of a transcriptional corepressor. Blood 2008; 111: 3145–3154.
Hudlebusch HR, Santoni-Rugiu E, Simon R, Ralfkiaer E, Rossing HH, Johansen JV et al. The histone methyltransferase and putative oncoprotein MMSET is overexpressed in a large variety of human tumors. Clin Cancer Res 17: 2919–2933.
Hudlebusch HR, Skotte J, Santoni-Rugiu E, Zimling ZG, Lees MJ, Simon R et al. MMSET is highly expressed and associated with aggressiveness in neuroblastoma. Cancer Res 71: 4226–4235.
Kassambara A, Klein B, Moreaux JMMSET . is overexpressed in cancers: link with tumor aggressiveness. Biochem Biophys Res Commun 2009; 379: 840–845.
Toyokawa G, Cho HS, Masuda K, Yamane Y, Yoshimatsu M, Hayami S et al. Histone lysine methyltransferase wolf-hirschhorn syndrome candidate 1 is involved in human carcinogenesis through regulation of the wnt pathway. Neoplasia 13: 887–898.
Li J, Yin C, Okamoto H, Mushlin H, Balgley BM, Lee CS et al. Identification of a novel proliferation-related protein, WHSC1 4a, in human gliomas. Neuro Oncol 2008; 10: 45–51.
Okabe H, Satoh S, Kato T, Kitahara O, Yanagawa R, Yamaoka Y et al. Genome-wide analysis of gene expression in human hepatocellular carcinomas using cDNA microarray: identification of genes involved in viral carcinogenesis and tumor progression. Cancer Res 2001; 61: 2129–2137.
Kwok WK, Ling MT, Lee TW, Lau TC, Zhou C, Zhang X et al. Up-regulation of TWIST in prostate cancer and its implication as a therapeutic target. Cancer Res 2005; 65: 5153–5162.
Yang J, Mani SA, Donaher JL, Ramaswamy S, Itzykson RA, Come C et al. Twist, a master regulator of morphogenesis, plays an essential role in tumor metastasis. Cell 2004; 117: 927–939.
Thiery JP, Acloque H, Huang RY, Nieto MA . Epithelial-mesenchymal transitions in development and disease. Cell 2009; 139: 871–890.
Brabletz T, Jung A, Spaderna S, Hlubek F, Kirchner T . Opinion: migrating cancer stem cells - an integrated concept of malignant tumour progression. Nat Rev Cancer 2005; 5: 744–749.
Lapointe J, Li C, Higgins JP, van de Rijn M, Bair E, Montgomery K et al. Gene expression profiling identifies clinically relevant subtypes of prostate cancer. Proc Natl Acad Sci USA 2004; 101: 811–816.
Varambally S, Yu J, Laxman B, Rhodes DR, Mehra R, Tomlins SA et al. Integrative genomic and proteomic analysis of prostate cancer reveals signatures of metastatic progression. Cancer Cell 2005; 8: 393–406.
Yu YP, Landsittel D, Jing L, Nelson J, Ren B, Liu L et al. Gene expression alterations in prostate cancer predicting tumor aggression and preceding development of malignancy. J Clin Oncol 2004; 22: 2790–2799.
Lombardi L, Poretti G, Mattioli M, Fabris S, Agnelli L, Bicciato S et al. Molecular characterization of human multiple myeloma cell lines by integrative genomics: insights into the biology of the disease. Genes Chromosomes Cancer 2007; 46: 226–238.
Agnelli L, Bicciato S, Mattioli M, Fabris S, Intini D, Verdelli D et al. Molecular classification of multiple myeloma: a distinct transcriptional profile characterizes patients expressing CCND1 and negative for 14q32 translocations. J Clin Oncol 2005; 23: 7296–7306.
Berger MF, Lawrence MS, Demichelis F, Drier Y, Cibulskis K, Sivachenko AY et al. The genomic complexity of primary human prostate cancer. Nature 2011; 470: 214–220.
Wang GG, Cai L, Pasillas MP, Kamps MP . NUP98-NSD1 links H3K36 methylation to Hox-A gene activation and leukaemogenesis. Nat Cell Biol 2007; 9: 804–812.
Morishita M, di Luccio E . Cancers and the NSD family of histone lysine methyltransferases. Biochim Biophys Acta 2011; 1816: 158–163.
Bergsagel PL, Kuehl WM . Molecular pathogenesis and a consequent classification of multiple myeloma. J Clin Oncol 2005; 23: 6333–6338.
Zhuo WL, Wang Y, Zhuo XL, Zhang YS, Chen ZT . Short interfering RNA directed against TWIST, a novel zinc finger transcription factor, increases A549 cell sensitivity to cisplatin via MAPK/mitochondrial pathway. Biochem Biophys Res Commun 2008; 369: 1098–1102.
Yang MH, Hsu DS, Wang HW, Wang HJ, Lan HY, Yang WH et al. Bmi1 is essential in Twist1-induced epithelial-mesenchymal transition. Nat Cell Biol 12: 982–992.
Vedadi M, Barsyte-Lovejoy D, Liu F, Rival-Gervier S, Allali-Hassani A, Labrie V et al. A chemical probe selectively inhibits G9a and GLP methyltransferase activity in cells. Nat Chem Biol 7: 566–574.
Kubicek S, O'Sullivan RJ, August EM, Hickey ER, Zhang Q, Teodoro ML et al. Reversal of H3K9me2 by a small-molecule inhibitor for the G9a histone methyltransferase. Mol Cell 2007; 25: 473–481.
Daigle SR, Olhava EJ, Therkelsen CA, Majer CR, Sneeringer CJ, Song J et al. Selective killing of mixed lineage leukemia cells by a potent small-molecule DOT1L inhibitor. Cancer Cell 20: 53–65.
Garcia BA, Mollah S, Ueberheide BM, Busby SA, Muratore TL, Shabanowitz J et al. Chemical derivatization of histones for facilitated analysis by mass spectrometry. Nat Protoc 2007; 2: 933–938.
MacLean B, Tomazela DM, Shulman N, Chambers M, Finney GL, Frewen B et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26: 966–968.
Acknowledgements
This work was supported by a Fundacion Alfonso Martin Escudero fellowship (T.E.), a Ruth Kirschstein National Research Service Award F32HL099177 (R.P.), the Associazione Italiana Ricerca sul Cancro (AIRC) (A.N.), NIH K99/R00CA129565 (J.Y.), the U.S. Department of Defense W81XWH-09-1-0193 (J.Y.), a Research Scholar Award RSG-12-085-01 from the American Cancer Society (J.Y.), R01GM067193 (N.L.K.), RO1CA123204 (J.D.L.) a Leukemia and Lymphoma Society Specialized Center of Research Award (J.D.L.) and Physical Sciences Oncology Center grant U54CA143869 (J.D.L. and N.L.K.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Ezponda, T., Popovic, R., Shah, M. et al. The histone methyltransferase MMSET/WHSC1 activates TWIST1 to promote an epithelial–mesenchymal transition and invasive properties of prostate cancer. Oncogene 32, 2882–2890 (2013). https://doi.org/10.1038/onc.2012.297
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2012.297
Keywords
This article is cited by
-
Epigenetic regulation of hybrid epithelial-mesenchymal cell states in cancer
Oncogene (2023)
-
Targeting SphK1/2 by SKI-178 inhibits prostate cancer cell growth
Cell Death & Disease (2023)
-
O-GlcNAc modified-TIP60/KAT5 is required for PCK1 deficiency-induced HCC metastasis
Oncogene (2021)
-
Identification of histone methyltransferase NSD2 as an important oncogenic gene in colorectal cancer
Cell Death & Disease (2021)
-
Histone methyltransferase WHSC1 inhibits colorectal cancer cell apoptosis via targeting anti-apoptotic BCL2
Cell Death Discovery (2021)