Abstract
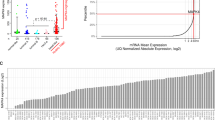
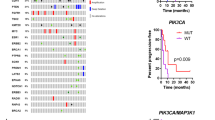
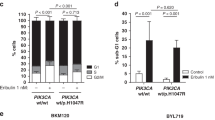
The phosphatidylinositol 3-kinase (PI3K) pathway is commonly activated in breast cancers due to frequent mutations in PIK3CA, loss of expression of PTEN or over-expression of receptor tyrosine kinases. PI3K pathway activation leads to stimulation of the key growth and proliferation regulatory kinase mammalian target of rapamycin (mTOR), which can be inhibited by rapamycin analogues and by kinase inhibitors; the effectiveness of these drugs in breast cancer treatment is currently being tested in clinical trials. To identify the molecular determinants of response to inhibitors that target mTOR via different mechanisms in breast cancer cells, we investigated the effects of pharmacological inhibition of mTOR using the allosteric mTORC1 inhibitor everolimus and the active-site mTORC1/mTORC2 kinase inhibitor PP242 on a panel of 31 breast cancer cell lines. We demonstrate here that breast cancer cells harbouring PIK3CA mutations are selectively sensitive to mTOR allosteric and kinase inhibitors. However, cells with PTEN loss of function are not sensitive to these drugs, suggesting that the functional consequences of these two mechanisms of activation of the mTOR pathway are quite distinct. In addition, a subset of HER2-amplified cell lines showed increased sensitivity to PP242, but not to everolimus, irrespective of the PIK3CA/PTEN status. These selective sensitivities were confirmed in more physiologically relevant three-dimensional cell culture models. Our findings provide a rationale to guide selection of breast cancer patients who may benefit from mTOR inhibitor therapy and highlight the importance of accurately assessing the expression of PTEN protein and not just its mutational status.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Adelaide J, Finetti P, Bekhouche I, Repellini L, Geneix J, Sircoulomb F et al. (2007). Integrated profiling of basal and luminal breast cancers. Cancer Res 67: 11565–11575.
American Type Culture Collection Standards Development Organization Workgroup ASN-000 (2010). Cell line misidentification: the beginning of the end. Nat Rev Cancer 10: 441–448.
Atkins MB, Yasothan U, Kirkpatrick P . (2009). Everolimus. Nat Rev Drug Discov 8: 535–536.
Bachman KE, Argani P, Samuels Y, Silliman N, Ptak J, Szabo S et al. (2004). The PIK3CA gene is mutated with high frequency in human breast cancers. Cancer Biol Ther 3: 772–775.
Brachmann SM, Hofmann I, Schnell C, Fritsch C, Wee S, Lane H et al. (2009). Specific apoptosis induction by the dual PI3K/mTor inhibitor NVP-BEZ235 in HER2 amplified and PIK3CA mutant breast cancer cells. Proc Natl Acad Sci USA 106: 22299–22304.
Brown EJ, Albers MW, Shin TB, Ichikawa K, Keith CT, Lane WS et al. (1994). A mammalian protein targeted by G1-arresting rapamycin-receptor complex. Nature 369: 756–758.
Carracedo A, Ma L, Teruya-Feldstein J, Rojo F, Salmena L, Alimonti A et al. (2008). Inhibition of mTORC1 leads to MAPK pathway activation through a PI3K-dependent feedback loop in human cancer. J Clin Invest 118: 3065–3074.
Chan S, Scheulen ME, Johnston S, Mross K, Cardoso F, Dittrich C et al. (2005). Phase II study of temsirolimus (CCI-779), a novel inhibitor of mTOR, in heavily pretreated patients with locally advanced or metastatic breast cancer. J Clin Oncol 23: 5314–5322.
Cobleigh MA, Langmuir VK, Sledge GW, Miller KD, Haney L, Novotny WF et al. (2003). A phase I/II dose-escalation trial of bevacizumab in previously treated metastatic breast cancer. Semin Oncol 30: 117–124.
Crowder RJ, Phommaly C, Tao Y, Hoog J, Luo J, Perou CM et al. (2009). PIK3CA and PIK3CB inhibition produce synthetic lethality when combined with estrogen deprivation in estrogen receptor-positive breast cancer. Cancer Res 69: 3955–3962.
Dan S, Okamura M, Seki M, Yamazaki K, Sugita H, Okui M et al. (2010). Correlating phosphatidylinositol 3-kinase inhibitor efficacy with signaling pathway status: in silico and biological evaluations. Cancer Res 70: 4982–4994.
Dancey J . (2010). mTOR signaling and drug development in cancer. Nat Rev Clin Oncol 7: 209–219.
Del Bufalo D, Ciuffreda L, Trisciuoglio D, Desideri M, Cognetti F, Zupi G et al. (2006). Antiangiogenic potential of the Mammalian target of rapamycin inhibitor temsirolimus. Cancer Res 66: 5549–5554.
Di Nicolantonio F, Arena S, Tabernero J, Grosso S, Molinari F, Macarulla T et al. (2010). Deregulation of the PI3K and KRAS signaling pathways in human cancer cells determines their response to everolimus. J Clin Invest 120: 2858–2866.
Ding L, Ellis MJ, Li S, Larson DE, Chen K, Wallis JW et al. (2010). Genome remodelling in a basal-like breast cancer metastasis and xenograft. Nature 464: 999–1005.
Efeyan A, Sabatini DM . (2010). mTOR and cancer: many loops in one pathway. Curr Opin Cell Biol 22: 169–176.
Ellard SL, Clemons M, Gelmon KA, Norris B, Kennecke H, Chia S et al. (2009). Randomized phase II study comparing two schedules of everolimus in patients with recurrent/metastatic breast cancer: NCIC Clinical Trials Group IND.163. J Clin Oncol 27: 4536–4541.
Engelman JA . (2009). Targeting PI3K signalling in cancer: opportunities, challenges and limitations. Nat Rev Cancer 9: 550–562.
Engelman JA, Luo J, Cantley LC . (2006). The evolution of phosphatidylinositol 3-kinases as regulators of growth and metabolism. Nat Rev Genet 7: 606–619.
Favata MF, Horiuchi KY, Manos EJ, Daulerio AJ, Stradley DA, Feeser WS et al. (1998). Identification of a novel inhibitor of mitogen-activated protein kinase kinase. J Biol Chem 273: 18623–18632.
Feldman ME, Apsel B, Uotila A, Loewith R, Knight ZA, Ruggero D et al. (2009). Active-site inhibitors of mTOR target rapamycin-resistant outputs of mTORC1 and mTORC2. PLoS Biol 7: e38.
Forbes SA, Bhamra G, Bamford S, Dawson E, Kok C, Clements J et al. (2008). The catalogue of somatic mutations in cancer (COSMIC). Curr Protoc Hum Genet Suppl 57: 10.11.1–10.11.26.
Guba M, von Breitenbuch P, Steinbauer M, Koehl G, Flegel S, Hornung M et al. (2002). Rapamycin inhibits primary and metastatic tumor growth by antiangiogenesis: involvement of vascular endothelial growth factor. Nat Med 8: 128–135.
Guertin DA, Sabatini DM . (2007). Defining the role of mTOR in cancer. Cancer Cell 12: 9–22.
Gymnopoulos M, Elsliger MA, Vogt PK . (2007). Rare cancer-specific mutations in PIK3CA show gain of function. Proc Natl Acad Sci USA 104: 5569–5574.
Hennessy BT, Smith DL, Ram PT, Lu Y, Mills GB . (2005). Exploiting the PI3K/AKT pathway for cancer drug discovery. Nat Rev Drug Discov 4: 988–1004.
Hollestelle A, Elstrodt F, Nagel JH, Kallemeijn WW, Schutte M . (2007). Phosphatidylinositol-3-OH kinase or RAS pathway mutations in human breast cancer cell lines. Mol Cancer Res 5: 195–201.
Hu X, Stern HM, Ge L, O'Brien C, Haydu L, Honchell CD et al. (2009). Genetic alterations and oncogenic pathways associated with breast cancer subtypes. Mol Cancer Res 7: 511–522.
Huse JT, Brennan C, Hambardzumyan D, Wee B, Pena J, Rouhanifard SH et al. (2009). The PTEN-regulating microRNA miR-26a is amplified in high-grade glioma and facilitates gliomagenesis in vivo. Genes Dev 23: 1327–1337.
Janes MR, Limon JJ, So L, Chen J, Lim RJ, Chavez MA et al. (2010). Effective and selective targeting of leukemia cells using a TORC1/2 kinase inhibitor. Nat Med 16: 205–213.
Janku F, Tsimberidou AM, Garrido-Laguna I, Wang X, Luthra R, Hong DS et al. (2011). PIK3CA mutations in patients with advanced cancers treated with PI3K/AKT/mTOR axis inhibitor. Mol Cancer Ther (e-pub ahead of print; doi:10.1158/1535-7163.MCT-10-0994).
Kalinsky K, Jacks LM, Heguy A, Patil S, Drobnjak M, Bhanot UK et al. (2009). PIK3CA mutation associates with improved outcome in breast cancer. Clin Cancer Res 15: 5049–5059.
Kwitkowski VE, Prowell TM, Ibrahim A, Farrell AT, Justice R, Mitchell SS et al. (2010). FDA approval summary: temsirolimus as treatment for advanced renal cell carcinoma. Oncologist 15: 428–435.
Lee GY, Kenny PA, Lee EH, Bissell MJ . (2007). Three-dimensional culture models of normal and malignant breast epithelial cells. Nat Methods 4: 359–365.
Levine DA, Bogomolniy F, Yee CJ, Lash A, Barakat RR, Borgen PI et al. (2005). Frequent mutation of the PIK3CA gene in ovarian and breast cancers. Clin Cancer Res 11: 2875–2878.
Liu P, Cheng H, Roberts TM, Zhao JJ . (2009). Targeting the phosphoinositide 3-kinase pathway in cancer. Nat Rev Drug Discov 8: 627–644.
Loi S, Haibe-Kains B, Majjaj S, Lallemand F, Durbecq V, Larsimont D et al. (2010). PIK3CA mutations associated with gene signature of low mTORC1 signaling and better outcomes in estrogen receptor-positive breast cancer. Proc Natl Acad Sci USA 107: 10208–10213.
Pallares J, Bussaglia E, Martinez-Guitarte JL, Dolcet X, Llobet D, Rue M et al. (2005). Immunohistochemical analysis of PTEN in endometrial carcinoma: a tissue microarray study with a comparison of four commercial antibodies in correlation with molecular abnormalities. Mod Pathol 18: 719–727.
Pampaloni F, Reynaud EG, Stelzer EH . (2007). The third dimension bridges the gap between cell culture and live tissue. Nat Rev Mol Cell Biol 8: 839–845.
Perez-Tenorio G, Alkhori L, Olsson B, Waltersson MA, Nordenskjold B, Rutqvist LE et al. (2007). PIK3CA mutations and PTEN loss correlate with similar prognostic factors and are not mutually exclusive in breast cancer. Clin Cancer Res 13: 3577–3584.
Pickl M, Ries CH . (2009). Comparison of 3D and 2D tumor models reveals enhanced HER2 activation in 3D associated with an increased response to trastuzumab. Oncogene 28: 461–468.
Poliseno L, Salmena L, Zhang J, Carver B, Haveman WJ, Pandolfi PP . (2010). A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature 465: 1033–1038.
Roidl A, Foo P, Wong W, Mann C, Bechtold S, Berger HJ et al. (2010). The FGFR4 Y367C mutant is a dominant oncogene in MDA-MB453 breast cancer cells. Oncogene 29: 1543–1552.
Saal LH, Holm K, Maurer M, Memeo L, Su T, Wang X et al. (2005). PIK3CA mutations correlate with hormone receptors, node metastasis, and ERBB2, and are mutually exclusive with PTEN loss in human breast carcinoma. Cancer Res 65: 2554–2559.
Sabatini DM, Erdjument-Bromage H, Lui M, Tempst P, Snyder SH . (1994). RAFT1: a mammalian protein that binds to FKBP12 in a rapamycin-dependent fashion and is homologous to yeast TORs. Cell 78: 35–43.
Samuels Y, Wang Z, Bardelli A, Silliman N, Ptak J, Szabo S et al. (2004). High frequency of mutations of the PIK3CA gene in human cancers. Science 304: 554.
Serebriiskii I, Castello-Cros R, Lamb A, Golemis EA, Cukierman E . (2008). Fibroblast-derived 3D matrix differentially regulates the growth and drug-responsiveness of human cancer cells. Matrix Biol 27: 573–585.
Serra V, Markman B, Scaltriti M, Eichhorn PJ, Valero V, Guzman M et al. (2008). NVP-BEZ235, a dual PI3K/mTOR inhibitor, prevents PI3K signaling and inhibits the growth of cancer cells with activating PI3K mutations. Cancer Res 68: 8022–8030.
She QB, Chandarlapaty S, Ye Q, Lobo J, Haskell KM, Leander KR et al. (2008). Breast tumor cells with PI3K mutation or HER2 amplification are selectively addicted to Akt signaling. PLoS One 3: e3065.
Stemke-Hale K, Gonzalez-Angulo AM, Lluch A, Neve RM, Kuo WL, Davies M et al. (2008). An integrative genomic and proteomic analysis of PIK3CA, PTEN, and AKT mutations in breast cancer. Cancer Res 68: 6084–6091.
Tomlinson GE, Chen TT, Stastny VA, Virmani AK, Spillman MA, Tonk V et al. (1998). Characterization of a breast cancer cell line derived from a germ-line BRCA1 mutation carrier. Cancer Res 58: 3237–3242.
Torbett NE, Luna-Moran A, Knight ZA, Houk A, Moasser M, Weiss W et al. (2008). A chemical screen in diverse breast cancer cell lines reveals genetic enhancers and suppressors of sensitivity to PI3K isoform-selective inhibition. Biochem J 415: 97–110.
Turner N, Lambros MB, Horlings HM, Pearson A, Sharpe R, Natrajan R et al. (2010). Integrative molecular profiling of triple negative breast cancers identifies amplicon drivers and potential therapeutic targets. Oncogene 29: 2013–2023.
Vasudevan KM, Barbie DA, Davies MA, Rabinovsky R, McNear CJ, Kim JJ et al. (2009). AKT-independent signaling downstream of oncogenic PIK3CA mutations in human cancer. Cancer Cell 16: 21–32.
Wang X, Jiang X . (2008). Post-translational regulation of PTEN. Oncogene 27: 5454–5463.
Weaver VM, Lelievre S, Lakins JN, Chrenek MA, Jones JC, Giancotti F et al. (2002). beta4 integrin-dependent formation of polarized three-dimensional architecture confers resistance to apoptosis in normal and malignant mammary epithelium. Cancer Cell 2: 205–216.
Wee S, Wiederschain D, Maira SM, Loo A, Miller C, deBeaumont R et al. (2008). PTEN-deficient cancers depend on PIK3CB. Proc Natl Acad Sci USA 105: 13057–13062.
Weigelt B, Bissell MJ . (2008). Unraveling the microenvironmental influences on the normal mammary gland and breast cancer. Semin Cancer Biol 18: 311–321.
Weigelt B, Lo AT, Park CC, Gray JW, Bissell MJ . (2010). HER2 signaling pathway activation and response of breast cancer cells to HER2-targeting agents is dependent strongly on the 3D microenvironment. Breast Cancer Res Treat 122: 35–43.
Yamada KM, Cukierman E . (2007). Modeling tissue morphogenesis and cancer in 3D. Cell 130: 601–610.
Yeh TC, Marsh V, Bernat BA, Ballard J, Colwell H, Evans RJ et al. (2007). Biological characterization of ARRY-142886 (AZD6244), a potent, highly selective mitogen-activated protein kinase kinase 1/2 inhibitor. Clin Cancer Res 13: 1576–1583.
Yuan TL, Cantley LC . (2008). PI3K pathway alterations in cancer: variations on a theme. Oncogene 27: 5497–5510.
Zhang S, Yu D . (2010). PI(3)king apart PTEN's role in cancer. Clin Cancer Res 16: 4325–4330.
Acknowledgements
We thank Miriam Molina Arcas, Ralph Fritsch, Elza de Bruin, Charles Swanton (CRUK London Research Institute) and Maryou Lambros and Jorge Reis-Filho (Breakthrough Breast Cancer Centre, London, UK) for helpful discussions, technical advice or critical reading of the manuscript, members of the LRI Equipment Park for sequencing and of the LRI FACS facility for cell-cycle analysis. We thank Morri Feldman and Kevan Shokat (UCSF) for providing PP242. This work was funded by Cancer Research UK.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Weigelt, B., Warne, P. & Downward, J. PIK3CA mutation, but not PTEN loss of function, determines the sensitivity of breast cancer cells to mTOR inhibitory drugs. Oncogene 30, 3222–3233 (2011). https://doi.org/10.1038/onc.2011.42
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2011.42
Keywords
This article is cited by
-
RB loss determines selective resistance and novel vulnerabilities in ER-positive breast cancer models
Oncogene (2022)
-
Feedback activation of STAT3 limits the response to PI3K/AKT/mTOR inhibitors in PTEN-deficient cancer cells
Oncogenesis (2021)
-
PTEN suppresses epithelial–mesenchymal transition and cancer stem cell activity by downregulating Abi1
Scientific Reports (2020)
-
WDHD1 is essential for the survival of PTEN-inactive triple-negative breast cancer
Cell Death & Disease (2020)
-
Comparison of Three Real-Time PCR Assays for the Detection of PIK3CA Somatic Mutations in Formalin-Fixed Paraffin Embedded Tissues of Patients with Breast Carcinomas
Pathology & Oncology Research (2019)