Abstract
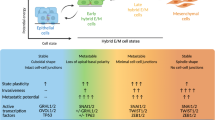
Epithelial-mesenchymal (EMT) and mesenchymal-epithelial (MET) transitions occur in the development of human tumorigenesis and are part of the natural history of the process to adapt to the changing microenvironment. In this setting, the miR-200 family is recognized as a master regulator of the epithelial phenotype by targeting ZEB1 and ZEB2, two important transcriptional repressors of the cell adherence (E-cadherin) and polarity (CRB3 and LGL2) genes. Recently, the putative DNA methylation associated inactivation of various miR-200 members has been described in cancer. Herein, we show that the miR-200ba429 and miR-200c141 transcripts undergo a dynamic epigenetic regulation linked to EMT or MET phenotypes in tumor progression. The 5′-CpG islands of both miR-200 loci were found unmethylated and coupled to the expression of the corresponding miRNAs in human cancer cell lines with epithelial features, such as low levels of ZEB1/ZEB2 and high expression of E-cadherin, CRB3 and LGL2, while CpG island hypermethylation-associated silencing was observed in transformed cells with mesenchymal characteristics. The recovery of miR-200ba429 and miR-200c141 expression by stable transfection in the hypermethylated cells restored the epithelial markers and inhibited migration in cell culture and tumoral growth and metastasis formation in nude mice. We also discovered, using both cell culture and animal models, that the miR-200 epigenetic silencing is not an static and fixed process but it can be shifted to hypermethylated or unmethylated 5′-CpG island status corresponding to the EMT and MET phenotypes, respectively. In fact, careful laser microdissection in human primary colorectal tumorigenesis unveiled that in normal colon mucosa crypts (epithelia) and stroma (mesenchyma) already are unmethylated and methylated at these loci, respectively; and that the colorectal tumors undergo selective miR-200 hypermethylation of their epithelial component. These findings indicate that the epigenetic silencing plasticity of the miR-200 family contributes to the evolving and adapting phenotypes of human tumors.
Similar content being viewed by others
Introduction
About 85% of human tumors are derived from epithelial cells (Edwards et al., 2010). The epithelial-mesenchymal transition (EMT) involves molecular reprogramming of the cells that gradually lose their epithelial features while acquiring mesenchymal attributes that promote cellular detachment from the primary tumor, invasion of adjacent stroma, entry into the systemic circulation and extravasation (Thiery, 2002; Yang and Weinberg, 2008; Acloque et al., 2009; Kalluri and Weinberg, 2009; Polyak and Weinberg, 2009; Thiery et al., 2009; Zeisberg and Neilson, 2009). This might be a reversible process and the cancer cells could eventually undergo mesenchymal-epithelial transition (MET) phenotype, then proliferate and form distant secondary tumors (Brabletz et al., 2001; Chaffer et al., 2006; Chaffer et al., 2007; Hugo et al., 2007; Polyak and Weinberg, 2009; Thiery et al., 2009; Aokage et al., 2010). Thus, epithelial and mesenchymal transitions are considered to be hallmarks of tumor progression and also have a central role in metastasis development, the leading cause of cancer-associated mortality.
From a molecular standpoint, most signaling pathways that regulate EMT converge on the control of E-cadherin, a cell adhesion protein crucial for maintaining epithelial structure (Peinado et al., 2007). Thus, the accurate regulation of transcriptional repressors of this gene, such as ZEB1 (TCF8/ZFHX1A), ZEB2 (SIP1/ZFHX1B), Snail (SNAI1), Slug (SNAI2) and other factors is crucial for normal epithelium and its dysregulation in cancer is associated with EMT (Peinado et al., 2007). In addition, lack of epithelial organization involves disruption of cell polarity (Etienne-Manneville, 2008; Moreno-Bueno et al., 2008; Bryant and Mostov, 2008; Huang and Muthuswamy, 2010). The E-cadherin repressor ZEB1 also represses the transcription of some polarity genes, including CRB3 (Crumbs3), MPP5 (Pals1), INADL (Patj) and LLGL2 (Lgl2) (Aigner et al., 2007; Spaderna et al., 2008).
If ZEB1 and ZEB2 have a central role in EMT, their activity must be finely regulated. MicroRNAs (miRNAs) are small non-coding RNAs that regulate gene expression by targeting messenger RNA (mRNA) transcripts (Medina and Slack, 2008; Agami, 2010). It is at this level that a miRNA family, miR-200, has important EMT regulatory activity in which it directly targets and inhibits ZEB1 and ZEB2 (Hurteau et al., 2006; Christoffersen et al., 2007; Hurteau et al., 2007; Burk et al., 2008; Gregory et al., 2008; Korpal et al., 2008; Park et al., 2008). Although several mechanisms may help control miR-200 activity, such as the existence of a double-negative feedback loop between the miR-200 family and ZEB factors based on the presence of E-boxes in miR-200 genes (Bracken et al., 2008; Burk et al., 2008), we focused on a common disruption in cancer cells: the epigenetic inactivation by CpG island hypermethylation surrounding the transcription start sites (Jones and Baylin, 2007; Esteller, 2008). Promoter hypermethylation and associated silencing of classical tumor suppressor genes is a molecular hallmark of human tumors (Jones and Baylin, 2007; Esteller, 2008), and this epigenetic lesion has recently been extended to the DNA methylation-associated silencing of miRNAs with growth-inhibitory features (Saito et al., 2006; Lujambio et al., 2007; Lujambio et al., 2008; Toyota et al., 2008; Huang et al., 2009). In this regard, recent reports have suggested the presence of hypermethylation-associated silencing of some miR-200 family members (Ceppi et al., 2010; Neves et al., 2010; Vrba et al., 2010; Wiklund et al., 2010).
The reversibility of epigenetic mechanisms might be a strategy for modulating sequences with distinct expression patterns during tumor progression. This might occur with miR-200 in association with acquiring initial mesenchymal attributes (EMT) and, once the cancer cells have reached a secondary site, regaining some epithelial properties (MET) for colonization and establishment of a secondary tumor. Herein, we show that the 5′-CpG island hypermethylation-associated silencing of the miR-200b/200a/429 and miR-200c/141 polycistronic transcripts is a dynamic process that mediates the shifts between EMT and MET phenotypes and contributes to cancer progression and metastasis formation in human tumorigenesis.
Results
CpG island hypermethylation-associated silencing of the miR-200 family in cancer cells
In order to evaluate the possible epigenetic regulation of the miR-200 family by CpG island methylation in transformed cells, we first screened a panel of cancer cell lines representing the most common human tumor types: colon (RKO and HCT116), lung (H157, EBC1 and H441) and breast (MDAMB231 and MCF7). Genetically, the miR-200 family is grouped in two polycistronic units: miR-200b/200a/429 (200ba429) and miR-200c/141 (200c141), clustered in chromosomes 1 and 12, respectively (Figure 1a). The miR-200b/200a/429 and miR-200c/141 transcription start sites have been previously determined (Bracken et al., 2008; Neves et al., 2010) and are located within canonical CpG islands (Figure 1a). The CpG islands of both clusters were densely methylated in RKO, H157 and MDAMB231, whereas HCT116, H441, EBC1 and MCF7 were found to be unmethylated (Figure 1b). Methylation-specific PCR analyses confirmed the bisulfite genomic sequencing data (Supplementary Figure 1).
CpG island hypermethylation-associated silencing of the miR-200 family in human cancer cell lines. (a) Schematic depiction of miR-200ba429 and miR-200c141 genomic loci. CpG islands (blue boxes) and transcription start sites (arrows) are shown. (b) Bisulfite genomic sequencing analysis of miR-200ba429 and miR-200c141 CpG islands in cancer cell lines. Six single clones are represented for each cell line. Location of bisulfite genomic sequencing PCR primers (black arrows) and CpG dinucleotides (vertical lines) are shown. Presence of unmethylated or methylated CpGs is indicated by white or black squares, respectively. (c) Expression of mature miRNAs of the miR-200 family in cancer cell lines, evaluated by qRT–PCR. Data represent relative quantification (mean±s.d.). (d) Restored mature miR-200 expression upon treatment with DNA-demethylating agent 5-aza-2′-deoxycytidine (aza) in miR-200 CpG island methylated cell lines (Student's t-test P<0.05 in all experiments).
To evaluate the impact of CpG methylation in transcription, we analyzed the expression of miR-200 family members. We observed that all the hypermethylated cancer cell lines had significant loss of both mature (Figure 1c) and pri-miRNA (Supplementary Figure 1) expression for the miR-200 family relative to the CpG island unmethylated cell lines derived from the same tumor type. Treatment of the miR-200 family hypermethylated cancer cell lines with the DNA-demethylating agent 5′-aza-2′-deoxycytidine restored the expression of the miRNAs at the mature (Figure 1d) and pri-miRNA (Supplementary Figure 1) stages.
Epigenetic silencing of the miR-200 family is associated with upregulation of ZEB1/ZEB2 and downregulation of their cell-adhesion and polarity-target genes
We next wondered about the impact of miR-200 epigenetic silencing on the expression of ZEB1 and ZEB2 and on the downstream cell-adhesion and polarity-target genes, such as E-cadherin (CDH1), CRB3 and LGL2. We observed that miR-200 family CpG island hypermethylated cancer cell lines had higher levels of expression of ZEB1 and ZEB2, quantified by qRT–PCR, than did unmethylated cells, in which these transcripts were inhibited (Supplementary Figure 2). To reinforce the results of the analysis of the consequences of miR-200 methylation for the entire pathway, we found that miR-200 hypermethylation and ZEB1/ZEB2 upregulation were associated with downregulation in CDH1, CRB3 and LGL2 (Supplementary Figure 2). The high level of expression of ZEB1 and the converse minimal levels of E-cadherin in miR-200-hypermethylated cancer cell lines, a characteristic feature in the EMT phenotype, were confirmed by western blot (Supplementary Figure 2).
We also reconstituted miR-200 activity by cloning each cluster (200ba429 and 200c141) followed by its stable transfection in the CpG island-hypermethylated cancer cell lines H157 and MDA-MB-231 (Supplementary Figure 2). We observed that stable transfection of the 200ba429 or 200c141 pri-miRNA clusters in both miR-200-hypermethylated cell lines caused the downregulation of their target genes, ZEB1 and ZEB2 (Supplementary Figure 2), and an increase in the expression of the downstream cell-adherence and polarity genes, E-cadherin, CRB3 and LGL2 (Supplementary Figure 2).
Reconstitution of miR-200 activity inhibits cellular migration, tumor growth and metastasis formation
Previous data indicate that miR-200 expression reduces motility and invasiveness while its inhibition increases migration capability (Burk et al., 2008; Gregory et al., 2008; Korpal et al., 2008; Gibbons et al., 2009). We sought to investigate these activities in the context of miR-200 epigenetic silencing. We first evaluated the effects of the miR-200 transfection in the hypermethylated cancer cell line RKO using the wound-healing assay. We observed that, in both cell lines, stable transfection of the clusters 200ba429, 200c141 or both together pri-miRNAs caused a significant reduction in the migration rate (Fisher's exact test P<0.01) (Figure 2a). The inhibition of cell migration was not observed before 24 h, and the reduced migration rate observed later was not associated with diminished cell proliferation determined by the MTT assay (Supplementary Figure 3).
Reconstitution of miR-200 levels inhibits cellular migration, tumor growth and metastasis formation. (a) Stable transfection of pri-miR-200ba429, pri-miR-200c141 and both clusters (pri-miR-200ba429 and -200c141) in the CpG island-hypermethylated cancer cell lines RKO caused decreased migration capability revealed by wound-healing assay. Accurate measures of the wounds were taken at 0, 24, 48 and 72 h to calculate the migration rate. Data are means±s.d. from two independent experiments (Fisher's exact test P<0.01 vs empty vector in all experiments). Illustrative examples of the time-coursed wound-healing assays are shown in the upper panel. (b) Tumor development of xenograft mouse models was monitored during 45 days after subcutaneous injection of empty vector or pri-miR-200c141 stably transfected RKO cells. Restoring of miR-200c141 in the hypermethylated RKO cells led to a significant reduction of tumor volume (Student's t-test P=0.02) and weight (Student's t-test P=0.04). Representative tumors are shown in the upper panel. (c) Decreased metastatic potential by forced miR-200c141 expression in methylated RKO cells. The presence of metastases was evaluated after 2 months of intrasplenic injection of pri-miR-200c141 or empty-vector RKO-transfected cells. Illustrative examples are shown in the left panel. Metastases are indicated by arrows.
Next, we performed in vivo functional experiments in nude mice. First, to test the tumorigenic capability we established xenograft models by subcutaneous injection of cancer cells transfected with the pri-miRNA 200c141 cluster, or the corresponding empty vector, into each flank of the mice. Notably, transfection of miR-200c141 in the miR-200-hypermethylated RKO cells significantly reduced their tumorigenicity as measured by tumor mass (Student's t-test P=0.04) and volume (Student's t-test P=0.02) (Figure 2b). We also tested the possible effect of miR-200 reconstitution in hypermethylated cancer cells upon metastatic potential. Following intrasplenic injection of miR-200-hypermethylated RKO cells stably transfected with the pri-miR-200c141 cluster, the development of hepatic metastases was analyzed. We found that, after 8 weeks, the mice injected with the miR-200c141 reconstituted-RKO cells presented liver metastases in only 25% (2/8) of cases, while the RKO-empty vector-transfected cells developed metastases in 67% (6/9) of the mice (Figure 2c).
Dynamic epigenetic silencing of the miRNA-200 family during in vitro-induced EMT and MET
Once we had established how the CpG island hypermethylation-associated silencing of miR-200 enhanced the tumorigenic features and re-expression of these transcripts inhibited the malignant phenotype, we began to study the dynamics of the process. In view of the increasing evidences of the importance of EMT and MET in the progression of the primary and metastatic disease (Brabletz et al., 2001; Thiery, 2002; Chaffer et al., 2006, 2007; Hugo et al., 2007; Yang and Weinberg, 2008; Acloque et al., 2009; Thiery et al., 2009; Kalluri and Weinberg, 2009; Polyak and Weinberg, 2009; Zeisberg and Neilson, 2009; Aokage et al., 2010) and the critical roles of the miR-200 family in these processes (Hurteau et al., 2006; Christoffersen et al., 2007; Hurteau et al., 2007; Burk et al., 2008; Gregory et al., 2008; Korpal et al., 2008; Park et al., 2008), it is plausible that the establishment of the DNA methylation patterns in the 5′-CpG islands of the miR-200 might be a reversible process that allows cellular plasticity and adaptation to new microenvironments.
To explore this possibility, we first used a well-known model of induced EMT, the treatment with transforming growth factor β (TGFβ) of Madin–Darby canine kidney cells (MDCK) (Moreno-Bueno et al., 2009). TGFβ is a potent inductor of EMT through several pathways, including the Smad pathway (Zavadil and Bottinger, 2005; Pardali and Moustakas, 2007; Xu et al., 2009). This is a reversible process and the cells can eventually undergo MET upon withdrawal of TGFβ from the culture medium (Figure 3a). Bisulfite genomic sequencing of multiple clones of the 5′-CpG islands of the miR-200ba429 and miR-200c141 demonstrated that, in agreement with the observed epithelial phenotype, these CpG sites are unmethylated in untreated MDCK cells (Figure 3b). The unmethylated CpG islands are associated with strong expression of the miR-200 (Figure 3c). However, upon induction of EMT by TGFβ treatment, there is a progressive gain of CpG methylation in the miR-200ba429 and miR-200c141 CpG islands (Figure 3b) in association with a reduced expression of the miR-200 (Figure 3c). Most notably, when these cells then underwent a change to a MET phenotype after TGFβ withdrawal from the medium (Figure 3a), the previously methylated CpG islands recovered the original unmethylated CpG status (Figure 3b), and the expression of the miR-200 was restored (Figure 3c). We also performed quantitative chromatin immunoprecipitation studies for three histone modification marks in the miR-200ba429 and miR-200c141 CpG islands in the EMT/MET MDCK model: two marks of active transcription (H3K4 trimethylation and H3K36 trimethylation) and one of silencing (H3K27 trimethylation). We observed that the DNA unmethylated CpG islands of the miR-200 loci were enriched in active marks (H3K4 trimethylation and H3K36 trimethylation) and depleted of the transcriptional silencing H3K27 trimethylation mark (Supplementary Figure 4). In contrast, the gain of CpG island methylation of the miR-200ba429 and miR-200c141 loci was associated with an increase in the silencing mark (H3K27 trimethylation) and a depletion of the active marks (H3K4 trimethylation and H3K36 trimethylation) (Supplementary Figure 4). MDCK cells that underwent EMT upon TGFβ use at the time points of 10, 20 and 30 days were also treated with 5′-aza-2′-deoxycytidine during 72 h. We observed that upon the use of DNA-demethylating agent, the miR-200ba429 and miR-200c141 CpG islands underwent hypomethylation events associated with a restoration of the expression of the corresponding miRNA transcripts (Figure 3d and Supplementary Figure 5). The epigenetic reactivation of the described miR-200 loci was associated with the acquisition of epithelial features, such as the downregulation of ZEB1 and the upregulation of E-cadherin (Supplementary Figure 5).
Dynamic epigenetic regulation of miR-200 expression during in vitro induced EMT and MET. (a) The epithelial MDCK cells underwent morphological changes toward a mesenchymal phenotype (EMT) during TGFβ treatment that was reverted after TGFβ withdrawal (mesenchymal to epithelial transition, MET). (b) Progressive gain of CpG methylation in both clusters of miR-200 family during TGFβ-induced EMT, revealed by bisulfite genomic sequencing. The original unmethylated status was re-established in cells undergoing MET after TGFβ withdrawal. In all, 20 single clones were evaluated in each experimental condition. Presence of unmethylated or methylated CpGs is indicated by white or black squares, respectively. (c) Decreased expression of mature miR-200 family members during TGFβ treatment (Student's t-test P<0.05), which was recovered after TGFβ withdrawal, analyzed by qRT–PCR. (d) Restored expression of miR-200 in TGFβ-treated cells is achieved upon treatment with DNA-demethylating agent 5-aza-2′-deoxycytidine (aza), evaluated by qRT–PCR of mature miR-200 family members (Student's t-test P<0.05).
The role of dynamic miR-200 epigenetic silencing in the establishment of EMT and MET was confirmed in two additional experimental lines. First, the described miR-200 hypermethylation-associated inactivation in the TGFβ-induced EMT was accompanied by increased ZEB1 expression, loss of E-cadherin and the acquisition of a spindle-like shape, a mesenchymal phenotype marked by Vimentin (Supplementary Figure 6). Second, the subsequent acquisition of a MET phenotype upon TGFβ depletion was not only characterized by the hypomethylation and re-expression of the aforementioned miR-200ba429 and miR-200c141 clusters, but also by a depletion of ZEB1 and Vimentin expression and the enhancement of the E-cadherin signal (Supplementary Figure 6). The ZEB1 and E-cadherin promoter CpG islands were found unmethylated in all the different stages of EMT and MET studied in MDCK cells (Supplementary Figure 7).
Dynamic epigenetic silencing of the miRNA-200 family during in vivo metastasis formation
The observation of the reversible miR-200 CpG island methylation patterns in cell culture during EMT and MET transitions, described above, prompted us to consider whether these dynamic epigenetic changes also took place in in vivo models of metastases where cancer cells also undergo important phenotypic changes. To address this matter, we used two mouse models: a paired human primary/metastasis breast cancer cell line obtained from the animal system and the analyses of liver metastases generated in nude mice by the intrasplenic injection of a human colorectal cancer cell line.
In the first case, we used an available model to analyze the EMT process occurring in metastasis, the paired breast cancer cell lines 468PT and 468LN (Vantyghem et al., 2005). The 468PT cell line was generated by stable transfection of the human breast adenocarcinoma cell line MDA-MB-468 with the pEGFP-C2 expression vector, whereas 468LN was derived from lung metastases dissected from a nude mouse following orthotopic injection with 468PT cells (Vantyghem et al., 2005). In contrast to the epithelioid phenotype of the parental 468PT cell line, 468LN are spindle-shaped cells, morphological changes associated with EMT. The 468LN cells produce extensive spontaneous lymph node metastases following orthotopic injection in nude mice, in contrast to the poorly metastatic MDA-MB-468 (Vantyghem et al., 2005). In agreement with the data obtained from the MDCK model, we observed that although miR-200ba429 and miR-200c141 loci both featured unmethylated 5′-CpG islands in the epithelial breast cancer cell line 468PT (Figure 4a), associated with a high level of expression of the transcripts (Figure 4b), the spindle-shaped metastatic 468LN cells underwent a dense CpG island hypermethylation for both miR-200 clusters (Figure 4a) that was linked to transcriptional silencing (Figure 4b). The use of a DNA-demethylating agent in the 468LN cell line was able to restore the expression of the miR-200ba429 and miR-200c141 transcripts (Figure 4c). The CpG island hypermethylation-associated silencing of the miR-200 in the metastatic 468LN was associated with the acquisition of an EMT phenotype with upregulation of the ZEB1 and Vimentin proteins and downregulation of E-cadherin (Supplementary Figure 8), relative to the miR-200 unmethylated parental epithelial cell line 468PT. Upon transfection of miR-200ba429, miR-200c141 or both loci together, the exogenous re-expression of these miR-200 loci transcripts reversed the phenotype of the mesenchymal-like 468LN cells: they acquired an epithelioid phenotype with ZEB and Vimentin downregulation and upregulation of the ZEB target CRB3 (Supplementary Figure 9). These changes were not associated with any change in the DNA methylation status of the endogenous miR-200ba429 and miR-200c141 CpG islands that stayed hypermethylated (Supplementary Figure 9).
Dynamic epigenetic silencing of the miRNA-200 family during in vivo metastasis formation. (a) CpG hypermethylation of both miR-200 clusters in the lung metastasis-derived 468LN cell line, in contrast to unmethylated status in the parental 468PT epithelial breast cancer cell line, analyzed by bisulfite genomic sequencing. In all, 20 single clones are represented for each cell line. Presence of unmethylated or methylated CpGs is indicated by white or black squares, respectively. (b) Significant reduction of miR-200 expression in metastatic 468LN cells (Student's t-test P<0.01). Data represent relative quantification of miRNAs, using the parental 468PT cell line as reference. (c) Restored expression of miR-200 in 468LN cells upon DNA-demethylating treatment with 5-aza-2′-deoxycytidine (aza), detected by qRT–PCR (Student's t-test P<0.05). (d) CpG island hypermethylation of miR-200c141 cluster during hepatic metastasis formation in mice. Spleen injection of 2 × 106 human colon cancer SW620 cells was able to generate hepatic metastases in 9 out of 15 mice after 8 weeks. Illustrative examples of metastases are shown (white arrows). (e) miR-200c141 CpG island methylation analysis of the hepatic metastases derived from SW620 shows hypermethylation of the cluster in two of them (HM3 and HM4), in contrast to the unmethylated status of the original SW620 cell line (CL). Results were obtained by bisulfite genomic sequencing analysis. Presence of unmethylated or methylated CpGs is indicated by white or black squares, respectively. (f) Significant decrease of pri-miR-200c141 in a CpG-hypermethylated hepatic metastasis (HM3) in comparison with the unmethylated metastases and original SW620 cell line (Student's t-test P<0.01). Data represent means±s.d. and were obtained by qRT–PCR. (g) Gain of expression of ZEB1, ZEB2 and Vimentin, and repression of E-cadherin in a miR-200c141-methylated hepatic metastasis (HM3). Human (hGAPDH) or mouse GAPDH (mGAPDH) were used as endogenous controls in semiquantitative RT–PCR. CL: original SW620 cell line; mNL: mouse normal liver.
In the second approach, we generated a mouse model of hepatic metastasis by intrasplenic injection of the human colorectal cancer cell line SW620. We injected 2 × 106 cells into 15 animals and verified the occurrence of metastasis in liver 8 weeks later. SW620 was able to generate macroscopic liver metastasis after 2 months in 9 of the 15 (60%) mice (Figure 4d). The miR-200ba429 5′-CpG island remained unmethylated in all cases (Supplementary Figure 10). Strikingly, in two of the mice carrying the hepatic metastases, the miR-200c141 epigenetic pattern was distinct from that observed in the parental cell line: if the original SW620 had an unmethylated CpG island (Figure 4e and Supplementary Figure 11) associated with expression of the transcript (Figure 4f), the liver metastases of SW620 had experienced dense miR-200c141 CpG island methylation (Figure 4e and Supplementary Figure 11) and loss of expression (Figure 4f) in association with the gain of EMT features such as ZEB1, ZEB2 and Vimentin overexpression and E-cadherin depletion (Figure 4g). These latter results in colorectal dissemination, together with the paired 468PT/468LN breast cancer experiment described above, highlight the role of epigenetic plasticity in miR-200 activities and later tumor propagation.
Differential miR-200 CpG island methylation in epithelial and stromal cells of human colonic mucosa
Having demonstrated that miR-200 CpG island methylation-associated silencing was characteristic of the mesenchymal phenotypes, while the unmethylated and expressing status of the CpG island was linked to epithelial features, we investigated whether these different DNA methylation patterns were also observed in human primary tissues. We took normal colon mucosa as a model. Bisulfite genomic sequencing of multiple clones (Figure 5a) and methylation-specific PCR analyses (Supplementary Figure 11) of the miR-200ba429 and miR-200c141 5′-CpG islands in primary total normal colorectal mucosa samples revealed a mixture of hypermethylated and unmethylated alleles in all the studied cases (n=10). These results may reflect the existence of a heterogeneous population within the sample: methylated/silenced and unmethylated/expressing. The stromal and epithelial components of the normal colon mucosa were good candidates in each respective category. Thus, we mechanically isolated the Lieberkühn colorectal crypts (the epithelial component) (Figure 5b) and the lamina propria-stromal cells (the mesenchymal component), and proceeded with the DNA methylation analyses. Normal epithelial cells from the colorectal crypts had unmethylated 5′-CpG islands for both miR-200ba429 and miR-200c141 (Figure 5c) and expressed the corresponding transcripts (Figure 5d), whereas lamina propria cells were hypermethylated at the miR-200 CpG islands (Figure 5c) and lacked expression of these miRNAs (Figure 5d). Consistent with the described observations, miR-200ba429 and miR-200c141 unmethylated crypt cells had minimal ZEB1 and ZEB2 expression, while the miR-200-hypermethylated stromal cells in the colon mucosa showed upregulated ZEB1/ZEB2 transcripts (Figure 5e). Thus, 5′-CpG island methylation of the miR-200 loci is an important component in the compartmental distinction between cell lineages in normal colon mucosa.
Differential miR-200 CpG island methylation in epithelial and stromal cells of normal human colonic mucosa. (a) miR-200ba429 and miR-200c141 5′-CpG island methylation analyses by bisulfite genomic sequencing in primary total normal colon mucosa samples demonstrated a mixture of unmethylated and methylated alleles. Illustrative examples are shown. (b) Epithelial and stromal components of human colonic mucosa were mechanically isolated. Illustrative microscope photographs in conventional bright-field illumination (left panel) or under phase contrast (right panel) of epithelial colonic crypts are shown. (c) Lack of DNA methylation in miR-200 CpG islands in epithelial component (colonic crypts), but hypermethylation in the surrounded lamina propria-stromal cells derived from normal colonic mucosa samples (NM), studied by bisulfite genomic sequencing. Illustrative examples are shown. (d) High miR-200 expression levels in unmethylated colonic crypts and loss of expression in methylated lamina propria cells, evaluated by qRT–PCR of primary miRNAs (Student's t-test P<0.01). (e) Inhibition of ZEB1 and ZEB2 mRNA expression in miR-200 unmethylated crypts (c), in contrast to high expression in miR-200-methylated lamina propria (LP) cells. GAPDH was used as endogenous control in semiquantitative RT–PCRs.
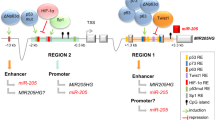
The next question to address was how the specific identified miR-200 CpG island methylation pattern that distinguishes the epithelia from the mesenchymal component of the normal colon mucosa could be disrupted in human colorectal tumorigenesis. Thus, we used laser microbeam microdissection and pressure catapulting to analyze the four tissue components found in 25 colorectal cancer patients: normal epithelium (the crypts) and stroma neighboring the colorectal tumor and, within the malignant pathological mass, its tumor epithelium (TE) and tumor stroma (Figure 6a). Bisulfite genomic sequencing demonstrated that the miR-200-hypermethylated 5′-CpG islands of the stroma of normal colorectal mucosa continued to be densely methylated in the stroma within colorectal tumors (Figure 6b) with minimal miRNA expression (Figure 6c). However, the 5′-CpG island of miR-200c141 underwent a hypermethylation event in the tumor epithelial cells relative to the unmethylated status of epithelium from the normal mucosa in 40% (10/25) of colorectal cancer patients (Figure 6b). miR-200c141 CpG island hypermethylation in the TE was associated with the transcriptional silencing of the transcript (Figure 6c). Following immunostaining for the epithelial marker E-cadherin in the 25 studied TE samples, among the cases that showed minimal expression of E-cadherin, miR-200c-141 CpG island hypermethylation was observed in six of eight (75%) cases (Supplementary Figure 12). None of the 25 analyzed TE samples showed E-cadherin CpG island hypermethylation (Supplementary Figure 12). miR-200c-141 CpG island hypermethylation was not observed in pre-neoplastic colorectal lesions (five microdissected adenomas) (Supplementary Figure 13). It is worth noting that the miR-200ba429 CpG island did not undergo dense methylation in the TE in the studied samples (Supplementary Figure 14), which suggests that different tissue and cancer-type activities might exist within the miR-200 family.
CpG island hypermethylation-associated silencing of miR-200c141 in human colorectal cancer. (a) Isolation of epithelial and stromal cells from normal and tumor colonic mucosa using laser microbeam microdissection and pressure catapulting. (b) Aberrant miR-200c141 CpG island hypermethylation in tumor epithelial cells (TE) in comparison with corresponding unmethylated normal epithelial crypts (NE), evaluated by bisulfite genomic sequencing. Both normal stroma (NS) and tumor stroma (TS) samples shown dense methylation. (c) Decreased expression of mature miR-200c and miR-141 in hypermethylated TE samples in contrast to high expression in paired normal epithelium samples. Both NS and TS samples showed low levels of miR-200c141. Data represent mean±s.d., obtained by qRT–PCR (Student's t-test P<0.05).
Discussion
The dynamic nature of the contribution of the DNA methylation mark to the ‘survival skills’ of cancer cells within the human body host is emerging as a critical tool for accomplishing this goal. Although many external and microenvironmental agents could be involved in shifting DNA methylation patterns, it is interesting that TGFβ seems to have a role in one case, that of the MDCK system. TGFβ induced EMT, and its withdrawal caused MET, linked with the corresponding 5′-CpG island methylation changes at miR-200ba429 and miR-200c141. Interestingly, an analogous de novo DNA methylation event in response to TGFβ has recently been reported as being an inactivation mechanism of β-4 integrin in NMuMG mouse mammary gland epithelial cells undergoing EMT (Yang et al., 2009). Likewise, acquisition of CpG methylation in several gene promoters, such as Twist, has been described upon serum-induced EMT in immortalized human mammary epithelial-expressing oncogenic ras cells (Dumont et al., 2008). Thus, local microenvironment and signals from tumor-associated stroma, including TGFβ that appear to be important for the induction and functional activation of EMT-related pathways (Zavadil and Bottinger, 2005; Pardali and Moustakas, 2007; Xu et al., 2009), might also mediate these effects through the action of epigenetic switches such as miR-200 5′-CpG island methylation events. The finding of possible triggers and controllers of the observed miR-200 loci DNA methylation dynamics in EMT would be extremely interesting. TGFβ has been involved in the establishment of DNA methylation patterns in cancer cells (reviewed by Chou et al., 2010) and in the activity of two epigenetic proteins, MTA1 (Pakala et al., 2011) and HDAC6 (Shan et al., 2008), in EMT. In addition, other histone modifiers, such as LSD1 (Lin et al., 2010), CREB-binding protein (Abell et al., 2011) and SIRT1 (Eades et al., 2011), also contribute to EMT.
Our study provides further clues about the paradoxical role of the miR-200 family during tumor progression and metastasis. During early tumor development, from the 5′-unmethylated miR-200 CpG islands the transcripts are produced and initially ‘secure’ the epithelial phenotype. However, the tumors progress and gain CpG promoter hypermethylation with a lack of miR-200 expression that promotes metastasis. In this regard, lower levels of miR-200c have been associated with a higher frequency of lymph node metastases in non-small cell lung cancer patients (Ceppi et al., 2010), and miR-200 downregulation is observed in metastatic lymph nodes (Baffa et al., 2009). However, once the micrometastases have been established, CpG island demethylation and gain of miR-200 expression could act as a macrometastasis promoter at the secondary sites. The experimental models for studying the complex multistep metastasis process are limited. Studies of isogenic cancer cell lines have shown that sub-lines overexpressing miR-200 (Dykxhoorn et al., 2009) or with more epithelial phenotypes (Chaffer et al., 2006) more efficiently produced local colonization growth and macrometastases than those with low miR-200 levels or mesenchymal-like phenotypes. Thus, if metastatic tumors tend to recapitulate the traits of the primary tumor from which they arose, including epithelial features favorable to tumor growth in the secondary organ, CpG hypermethylation of the miR-200 family could be a transitional state. Under such circumstances, epigenetic silencing of these master regulators of the epithelial phenotype could initially facilitate the dissemination of cancer cells, although restoring unmethylated miR-200 CpG island status and undergoing MET could be advantageous once the cells have reached the secondary site.
Many aspects remain to be investigated in future research. The first is the recognition of the presence of different miR-200 5′-CpG island DNA methylation patterns according to the epithelial or mesenchymal origin of the cell type. This finding is supported by the distinct miR-200c141 methylation profile observed in mammary epithelial cells and isogenic fibroblasts (Vrba et al., 2010). In addition, a miR-200 downregulation has been reported in the mesenchymal component of endometrial carcinosarcomas in comparison with the epithelial part, confirming the role of miR-200 in EMT in human tumors (Castilla et al., 2011). These results indicate the need for careful microdissection experiments if we want to exploit the potential of miR-200 CpG island hypermethylation as a biomarker of tumor behavior for future clinical purposes. Second, our results might provide a molecular base for new uses of pharmacological compounds with DNA-demethylating activity in the treatment of cancer patients (Mack, 2006; Issa, 2007; Esteller, 2008; Kaiser, 2010). Drug-induced miR-200 CpG island demethylation, which restores the expression of these transcripts, might inhibit tumor progression and micrometastasis formation mediated by EMT, as suggested by our in vitro and in vivo results. This area warrants further pre-clinical research, because of the potential benefits it offers for the treatment of cancer patients with metastatic disease, where extremely few therapeutic options are currently available.
Materials and methods
Cell lines and DNA methylation analysis
Cell lines and culture conditions are detailed in Supplementary Methods. For epigenetic drug treatments, cells were treated with 2 μM 5-aza-2′-deoxycytidine (Sigma, St Louis, MO, USA) for 72 h. For TGFβ treatments, recombinant human TGFβ (R&D systems, Minneapolis, MN, USA) was added to the culture medium at a final concentration of 5 ng/ml. CpG islands were identified in silico using Methyl Primer Express v1.0 software (Applied Biosystems, Carlsbad, CA, USA). DNA methylation status was established by bisulfite genomic sequencing of multiple clones and methylation-specific PCR. Primers are listed in Supplementary Table 1.
RNA extraction and real-time qRT–PCR
Total RNA was extracted using Trizol (Invitrogen, San Diego, CA, USA) and DNase-treated with Turbo DNA-free (Ambion, Austin, TX, USA). mRNA expression was analyzed using SYBR Green (Applied Biosystems). For miRNA quantification, TaqMan Pri-miRNA or MicroRNA Assays (Applied Biosystems) (Chen et al., 2005) were used according to the manufacture's protocol. Further information is described in Supplementary Methods. Primers and TaqMan assays are listed in Supplementary Table 1.
Protein extraction, western blot and immunofluorescence
Cell pellets were lysed in RIPA buffer containing complete protease inhibitor cocktail (Roche, Mannheim, Germany). Proteins were quantified with DC Protein Assay (Bio-Rad, Hercules, CA, USA) and 50 μg of whole-cell lysates were separated by SDS–PAGE. Blots were probed using anti-ZEB1 antibody (sc-10572, Santa Cruz Biotechnology, Santa Cruz, CA, USA), anti-Vimentin (CBL-202, Chemicon-Millipore, Billerica, MA, USA) or anti-E-cadherin (ab-1416, Abcam, Cambridge, UK). Proteins were detected using horseradish peroxidase-conjugated secondary antibodies. For immunofluorescence, cells were cultured directly on coverslips and double immunostainings were performed as detailed in Supplementary Methods.
miR-200 cloning and stable transfection
The genomic regions surrounding the pri-miRNA sequences of miR-200 family members were amplified using primers listed in Supplementary Table 1. Products were directionally cloned in pSilencer 4.1-CMV puro (Ambion). Cells seeded in six-well plates were transfected using Lipofectamine 2000 (Invitrogen), according to the manufacturer's instructions.
Wound-healing assay
Control and miR-200 stably transfected cells were grown to confluence, and a wound was made through the monolayer using a p1000 tip. Accurate measures of the wounds were taken during the time course to calculate the migration rate according to the equation: percentage wound healing=((wound length at 0 h)−(wound length at 24, 48 or 72 h))/(wound length at 0 h) × 100. Two independent experiments were performed.
Mouse xenograft and metastasis models
Five-week-old male athymic nu/nu mice (Charles River, Wilmington, MA, USA), housed under specific pathogen-free conditions, were used in this study. For xenografts, a total of 3.5 × 106 cells were subcutaneously injected in each flank of the mouse. The right flank was used for RKO control cells (empty vector) and the left for the miR-200c141 stably transfected RKO cells. Tumor development was monitored every 5 days and the tumor width (W) and length (L) were measured. Tumor volume was estimated according to the formula V=π/6 × L × W2. Mice were killed 45 days after injection, and tumors were then excised and weighed. For the experimental metastasis assay, mice were anesthetized, the spleens were exposed through a cut on the left flank and 2 × 106 control (empty vector) or miR-200c141 stably transfected RKO cells were injected into the middle of spleen. In order to minimize local tumor growth, the spleens were resected 48 h after injection. Hepatic metastasis were examined 8 weeks later, analyzed macroscopically and by H&E tissue staining. In all, 10 animals were tested for each experimental condition. All experiments with mice were approved by the IDIBELL Animal Care and Use Committee.
Colonic crypt isolation from fresh tissue
Colonic tissues were obtained from surgical resection of colorectal cancer patients at the Bellvitge Hospital, Barcelona, Spain. Written informed consents were obtained from all patients. Colonic crypts were isolated by a previously described method (Fujimoto et al., 2002). Lamina propria and crypt samples were snap-frozen in liquid nitrogen and stored at −80 °C before DNA and RNA extraction.
Laser microdissection from paraffin-embedded and frozen tissues
Tissues were provided by the Department of Pathology, Hospital São João, Porto, Portugal. All patients gave written consent and the hospital Ethics Committee approved the study. Laser microbeam microdissection and pressure catapulting process is detailed in Supplementary Methods. A minimum of 1,500 cell equivalents were isolated. DNA and RNA were extracted using a RecoverAll Total Nucleic Acid Isolation Kit for FFPE Tissues (Ambion).
Statistical analyses
Experiments were performed in triplicate and Fisher's exact test or Student's two-tailed t-test were used to identify statistical significance of differences between group means.
References
Abell AN, Jordan NV, Huang W, Prat A, Midland AA, Johnson NL et al. (2011). MAP3K4/CBP-regulated H2B acetylation controls epithelial-mesenchymal transition in trophoblast stem cells. Cell Stem Cell 8: 525–537.
Acloque H, Adams MS, Fishwick K, Bronner-Fraser M, Nieto MA . (2009). Epithelial-mesenchymal transitions: the importance of changing cell state in development and disease. J Clin Invest 119: 1438–1449.
Agami R . (2010). microRNAs, RNA binding proteins and cancer. Eur J Clin Invest 40: 370–374.
Aigner K, Dampier B, Descovich L, Mikula M, Sultan A, Schreiber M et al. (2007). The transcription factor ZEB1 (deltaEF1) promotes tumour cell dedifferentiation by repressing master regulators of epithelial polarity. Oncogene 26: 6979–6988.
Aokage K, Ishii G, Ohtaki Y, Yamaguchi Y, Hishida T, Yoshida J et al. (2010). Dynamic molecular changes associated with epithelial-mesenchymal transition and subsequent mesenchymal-epithelial transition in the early phase of metastatic tumor formation. Int J Cancer 128: 1585–1595.
Baffa R, Fassan M, Volinia S, O′Hara B, Liu CG, Palazzo JP et al. (2009). MicroRNA expression profiling of human metastatic cancers identifies cancer gene targets. J Pathol 219: 214–221.
Brabletz T, Jung A, Reu S, Porzner M, Hlubek F, Kunz-Schughart LA et al. (2001). Variable beta-catenin expression in colorectal cancers indicates tumor progression driven by the tumor environment. Proc Natl Acad Sci USA 98: 10356–10361.
Bracken CP, Gregory PA, Kolesnikoff N, Bert AG, Wang J, Shannon MF et al. (2008). A double-negative feedback loop between ZEB1-SIP1 and the microRNA-200 family regulates epithelial-mesenchymal transition. Cancer Res 68: 7846–7854.
Bryant DM, Mostov KE . (2008). From cells to organs: building polarized tissue. Nat Rev Mol Cell Biol 9: 887–901.
Burk U, Schubert J, Wellner U, Schmalhofer O, Vincan E, Spaderna S et al. (2008). A reciprocal repression between ZEB1 and members of the miR-200 family promotes EMT and invasion in cancer cells. EMBO Rep 9: 582–589.
Castilla MA, Moreno-Bueno G, Romero-Perez L, De Vijver KV, Biscuola M, Lopez-Garcia MA et al. (2011). Micro-RNA signature of the epithelial-mesenchymal transition in endometrial carcinosarcoma. J Pathol 223: 72–80.
Ceppi P, Mudduluru G, Kumarswamy R, Rapa I, Scagliotti GV, Papotti M et al. (2010). Loss of miR-200c expression induces an aggressive, invasive, and chemoresistant phenotype in non-small cell lung cancer. Mol Cancer Res 8: 1207–1216.
Chaffer CL, Brennan JP, Slavin JL, Blick T, Thompson EW, Williams ED . (2006). Mesenchymal-to-epithelial transition facilitates bladder cancer metastasis: role of fibroblast growth factor receptor-2. Cancer Res 66: 11271–11278.
Chaffer CL, Thompson EW, Williams ED . (2007). Mesenchymal to epithelial transition in development and disease. Cells Tissues Organs 185: 7–19.
Chen C, Ridzon DA, Broomer AJ, Zhou Z, Lee DH, Nguyen JT et al. (2005). Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res 33: e179.
Chou JL, Chen LY, Lai HC, Chan MW . (2010). TGF-beta: friend or foe? The role of TGF-beta/SMAD signaling in epigenetic silencing of ovarian cancer and its implication in epigenetic therapy. Expert Opin Ther Targets 14: 1213–1223.
Christoffersen NR, Silahtaroglu A, Orom UA, Kauppinen S, Lund AH . (2007). miR-200b mediates post-transcriptional repression of ZFHX1B. Rna 13: 1172–1178.
Dumont N, Wilson MB, Crawford YG, Reynolds PA, Sigaroudinia M, Tlsty TD . (2008). Sustained induction of epithelial to mesenchymal transition activates DNA methylation of genes silenced in basal-like breast cancers. Proc Natl Acad Sci USA 105: 14867–14872.
Dykxhoorn DM, Wu Y, Xie H, Yu F, Lal A, Petrocca F et al. (2009). miR-200 enhances mouse breast cancer cell colonization to form distant metastases. PLoS One 4: e7181.
Eades G, Yao Y, Yang M, Zhang Y, Chumsri S, Zhou Q . (2011). MiR-200a regulates SIRT1 and EMT-like transformation in mammary epithelial cells. J Biol Chem 286: 25992–26002.
Edwards BK, Ward E, Kohler BA, Eheman C, Zauber AG, Anderson RN et al. (2010). Annual report to the nation on the status of cancer, 1975-2006, featuring colorectal cancer trends and impact of interventions (risk factors, screening, and treatment) to reduce future rates. Cancer 116: 544–573.
Esteller M . (2008). Epigenetics in cancer. N Engl J Med 358: 1148–1159.
Etienne-Manneville S . (2008). Polarity proteins in migration and invasion. Oncogene 27: 6970–6980.
Fujimoto K, Beauchamp RD, Whitehead RH . (2002). Identification and isolation of candidate human colonic clonogenic cells based on cell surface integrin expression. Gastroenterology 123: 1941–1948.
Gibbons DL, Lin W, Creighton CJ, Rizvi ZH, Gregory PA, Goodall GJ et al. (2009). Contextual extracellular cues promote tumor cell EMT and metastasis by regulating miR-200 family expression. Genes Dev 23: 2140–2151.
Gregory PA, Bert AG, Paterson EL, Barry SC, Tsykin A, Farshid G et al. (2008). The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat Cell Biol 10: 593–601.
Huang L, Muthuswamy SK . (2010). Polarity protein alterations in carcinoma: a focus on emerging roles for polarity regulators. Curr Opin Genet Dev 20: 41–50.
Huang YW, Liu JC, Deatherage DE, Luo J, Mutch DG, Goodfellow PJ et al. (2009). Epigenetic repression of microRNA-129-2 leads to overexpression of SOX4 oncogene in endometrial cancer. Cancer Res 69: 9038–9046.
Hugo H, Ackland ML, Blick T, Lawrence MG, Clements JA, Williams ED et al. (2007). Epithelial--mesenchymal and mesenchymal--epithelial transitions in carcinoma progression. J Cell Physiol 213: 374–383.
Hurteau GJ, Carlson JA, Spivack SD, Brock GJ . (2007). Overexpression of the microRNA hsa-miR-200c leads to reduced expression of transcription factor 8 and increased expression of E-cadherin. Cancer Res 67: 7972–7976.
Hurteau GJ, Spivack SD, Brock GJ . (2006). Potential mRNA degradation targets of hsa-miR-200c, identified using informatics and qRT-PCR. Cell Cycle 5: 1951–1956.
Issa JP . (2007). DNA methylation as a therapeutic target in cancer. Clin Cancer Res 13: 1634–1637.
Jones PA, Baylin SB . (2007). The epigenomics of cancer. Cell 128: 683–692.
Kaiser J . (2010). Epigenetic drugs take on cancer. Science 330: 576–578.
Kalluri R, Weinberg RA . (2009). The basics of epithelial-mesenchymal transition. J Clin Invest 119: 1420–1428.
Korpal M, Lee ES, Hu G, Kang Y . (2008). The miR-200 family inhibits epithelial-mesenchymal transition and cancer cell migration by direct targeting of E-cadherin transcriptional repressors ZEB1 and ZEB2. J Biol Chem 283: 14910–14914.
Lin T, Ponn A, Hu X, Law BK, Lu J . (2010). Requirement of the histone demethylase LSD1 in Snai1-mediated transcriptional repression during epithelial-mesenchymal transition. Oncogene 29: 4896–4904.
Lujambio A, Calin GA, Villanueva A, Ropero S, Sanchez-Cespedes M, Blanco D et al. (2008). A microRNA DNA methylation signature for human cancer metastasis. Proc Natl Acad Sci USA 105: 13556–13561.
Lujambio A, Ropero S, Ballestar E, Fraga MF, Cerrato C, Setien F et al. (2007). Genetic unmasking of an epigenetically silenced microRNA in human cancer cells. Cancer Res 67: 1424–1429.
Mack GS . (2006). Epigenetic cancer therapy makes headway. J Natl Cancer Inst 98: 1443–1444.
Medina PP, Slack FJ . (2008). microRNAs and cancer: an overview. Cell Cycle 7: 2485–2492.
Moreno-Bueno G, Peinado H, Molina P, Olmeda D, Cubillo E, Santos V et al. (2009). The morphological and molecular features of the epithelial-to-mesenchymal transition. Nat Protoc 4: 1591–1613.
Moreno-Bueno G, Portillo F, Cano A . (2008). Transcriptional regulation of cell polarity in EMT and cancer. Oncogene 27: 6958–6969.
Neves R, Scheel C, Weinhold S, Honisch E, Iwaniuk KM, Trompeter HI et al. (2010). Role of DNA methylation in miR-200c/141 cluster silencing in invasive breast cancer cells. BMC Res Notes 3: 219.
Pakala SB, Singh K, Reddy SD, Ohshiro K, Li DQ, Mishra L et al. (2011). TGF-beta1 signaling targets metastasis-associated protein 1, a new effector in epithelial cells. Oncogene 30: 2230–2241.
Pardali K, Moustakas A . (2007). Actions of TGF-beta as tumor suppressor and pro-metastatic factor in human cancer. Biochim Biophys Acta 1775: 21–62.
Park SM, Gaur AB, Lengyel E, Peter ME . (2008). The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev 22: 894–907.
Peinado H, Olmeda D, Cano A . (2007). Snail, Zeb and bHLH factors in tumour progression: an alliance against the epithelial phenotype? Nat Rev Cancer 7: 415–428.
Polyak K, Weinberg RA . (2009). Transitions between epithelial and mesenchymal states: acquisition of malignant and stem cell traits. Nat Rev Cancer 9: 265–273.
Saito Y, Liang G, Egger G, Friedman JM, Chuang JC, Coetzee GA et al. (2006). Specific activation of microRNA-127 with downregulation of the proto-oncogene BCL6 by chromatin-modifying drugs in human cancer cells. Cancer Cell 9: 435–443.
Shan B, Yao TP, Nguyen HT, Zhuo Y, Levy DR, Klingsberg RC et al. (2008). Requirement of HDAC6 for transforming growth factor-beta1-induced epithelial-mesenchymal transition. J Biol Chem 283: 21065–21073.
Spaderna S, Schmalhofer O, Wahlbuhl M, Dimmler A, Bauer K, Sultan A et al. (2008). The transcriptional repressor ZEB1 promotes metastasis and loss of cell polarity in cancer. Cancer Res 68: 537–544.
Thiery JP, Acloque H, Huang RY, Nieto MA . (2009). Epithelial-mesenchymal transitions in development and disease. Cell 139: 871–890.
Thiery JP . (2002). Epithelial-mesenchymal transitions in tumour progression. Nat Rev Cancer 2: 442–454.
Toyota M, Suzuki H, Sasaki Y, Maruyama R, Imai K, Shinomura Y et al. (2008). Epigenetic silencing of microRNA-34b/c and B-cell translocation gene 4 is associated with CpG island methylation in colorectal cancer. Cancer Res 68: 4123–4132.
Vantyghem SA, Allan AL, Postenka CO, Al-Katib W, Keeney M, Tuck AB et al. (2005). A new model for lymphatic metastasis: development of a variant of the MDA-MB-468 human breast cancer cell line that aggressively metastasizes to lymph nodes. Clin Exp Metastasis 22: 351–361.
Vrba L, Jensen TJ, Garbe JC, Heimark RL, Cress AE, Dickinson S et al. (2010). Role for DNA methylation in the regulation of miR-200c and miR-141 expression in normal and cancer cells. PLoS One 5: e8697.
Wiklund ED, Bramsen JB, Hulf T, Dyrskjot L, Ramanathan R, Hansen TB et al. (2010). Coordinated epigenetic repression of the miR-200 family and miR-205 in invasive bladder cancer. Int J Cancer 128: 1327–1334.
Xu J, Lamouille S, Derynck R . (2009). TGF-beta-induced epithelial to mesenchymal transition. Cell Res 19: 156–172.
Yang J, Weinberg RA . (2008). Epithelial-mesenchymal transition: at the crossroads of development and tumor metastasis. Dev Cell 14: 818–829.
Yang X, Pursell B, Lu S, Chang TK, Mercurio AM . (2009). Regulation of beta 4-integrin expression by epigenetic modifications in the mammary gland and during the epithelial-to-mesenchymal transition. J Cell Sci 122: 2473–2480.
Zavadil J, Bottinger EP . (2005). TGF-beta and epithelial-to-mesenchymal transitions. Oncogene 24: 5764–5774.
Zeisberg M, Neilson EG . (2009). Biomarkers for epithelial-mesenchymal transitions. J Clin Invest 119: 1429–1437.
Acknowledgements
We thank Dr Balazs Balint for bioinformatic support. This study was supported by Fondo de Investigaciones Sanitarias Grant PI08-1345, Olga Torres Foundation Grant, Consolider Grant MEC09-05, Dr Josef Steiner Cancer Research Foundation Award, Lilly Foundation Biomedical Research Award, European Research Council Advanced Grant EPINORC and the Health Department of the Catalan Government (Generalitat de Catalunya). VD is supported by Instituto de Salud Carlos III, Sara Borrell postdoctoral contract. ME is an Institucio Catalana de Recerca i Estudis Avançats (ICREA) Research Professor.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Davalos, V., Moutinho, C., Villanueva, A. et al. Dynamic epigenetic regulation of the microRNA-200 family mediates epithelial and mesenchymal transitions in human tumorigenesis. Oncogene 31, 2062–2074 (2012). https://doi.org/10.1038/onc.2011.383
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2011.383
Keywords
This article is cited by
-
The epithelial–mesenchymal plasticity landscape: principles of design and mechanisms of regulation
Nature Reviews Genetics (2023)
-
The miR-200 family in normal mammary gland development
BMC Developmental Biology (2021)
-
LncRNA-NNT-AS1 contributes to the progression of glioma by miR-582-5p/EZH2 axis
Cytotechnology (2021)
-
Epigenetic regulation in human cancer: the potential role of epi-drug in cancer therapy
Molecular Cancer (2020)
-
Epigenetics of colorectal cancer: biomarker and therapeutic potential
Nature Reviews Gastroenterology & Hepatology (2020)