Abstract
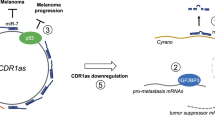
MicroRNAs (miRNAs) are small ∼22nt single stranded RNAs that negatively regulate protein expression by binding to partially complementary sequences in the 3′ untranslated region (3′ UTRs) of target gene messenger RNAs (mRNA). Recently, mutations have been identified in both miRNAs and target genes that disrupt regulatory relationships, contribute to oncogenesis and serve as biomarkers for cancer risk. KIT, an established oncogene with a multifaceted role in melanogenesis and melanoma pathogenesis, has recently been shown to be upregulated in some melanomas, and is also a target of the miRNA miR-221. Here, we describe a genetic variant in the 3′ UTR of the KIT oncogene that correlates with a greater than fourfold increased risk of acral melanoma. This KIT variant results in a mismatch in the seed region of a miR-221 complementary site and reporter data suggests that this mismatch can result in increased expression of the KIT oncogene. Consistent with the hypothesis that this is a functional variant, KIT mRNA and protein levels are both increased in the majority of samples harboring the KIT variant. This work identifies a novel genetic marker for increased heritable risk of melanoma.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Alexeev V, Yoon K . (2006). Distinctive role of the cKit receptor tyrosine kinase signaling in mammalian melanocytes. J Invest Dermatol 126: 1102–1110.
Anderson MA, Gusella JF . (1984). Use of cyclosporin A in establishing Epstein-Barr virus-transformed human lymphoblastoid cell lines. In vitro 20: 856–858.
Ashida A, Takata M, Murata H, Kido K, Saida T . (2009). Pathological activation of KIT in metastatic tumors of acral and mucosal melanomas. Int J Cancer 124: 862–868.
Chin LJ, Ratner E, Leng S, Zhai R, Nallur S, Babar I et al. (2008). A SNP in a let-7 microRNA complementary site in the KRAS 3′ untranslated region increases non-small cell lung cancer risk. Cancer Res 68: 8535–8540.
Curtin JA, Busam K, Pinkel D, Bastian BC . (2006). Somatic activation of KIT in distinct subtypes of melanoma. J Clin Oncol 24: 4340–4346.
Demetri GD, von Mehren M, Blanke CD, Van den Abbeele AD, Eisenberg B, Roberts PJ et al. (2002). Efficacy and safety of imatinib mesylate in advanced gastrointestinal stromal tumors. N Engl J Med 347: 472–480.
Felicetti F, Errico MC, Bottero L, Segnalini P, Stoppacciaro A, Biffoni M et al. (2008). The promyelocytic leukemia zinc finger-microRNA-221/-222 pathway controls melanoma progression through multiple oncogenic mechanisms. Cancer Res 68: 2745–2754.
Funasaka Y, Boulton T, Cobb M, Yarden Y, Fan B, Lyman SD et al. (1992). c-Kit-kinase induces a cascade of protein tyrosine phosphorylation in normal human melanocytes in response to mast cell growth factor and stimulates mitogen-activated protein kinase but is down-regulated in melanomas. Mol Biol Cell 3: 197–209.
Gershenwald JE, Soong SJ, Balch CM . (2010). TNM staging system for cutaneous melanoma.and beyond. Ann Surg Oncol 17: 1475–1477.
Godshalk SE, Bhaduri-McIntosh S, Slack FJ . (2008). Epstein-Barr virus-mediated dysregulation of human microRNA expression. Cell Cycle 7: 3595–3600.
Halaban R, Krauthammer M, Pelizzola M, Cheng E, Kovacs D, Sznol M et al. (2009). PLoS One 4: e4563.
He H, Jazdzewski K, Li W, Liyanarachchi S, Nagy R, Volinia S et al. (2005). The role of microRNA genes in papillary thyroid carcinoma. Proc Natl Acad Sci USA 102: 19075–19080.
Hodi FS, Friedlander P, Corless CL, Heinrich MC, Mac Rae S, Kruse A et al. (2008). Major response to imatinib mesylate in KIT-mutated melanoma. J Clin Oncol 26: 2046–2051.
Huang S, Jean D, Luca M, Tainsky MA, Bar-Eli M . (1998). Loss of AP-2 results in downregulation of c-KIT and enhancement of melanoma tumorigenicity and metastasis. Embo J 17: 4358–4369.
Huang S, Luca M, Gutman M, McConkey DJ, Langley KE, Lyman SD et al. (1996). Enforced c-KIT expression renders highly metastatic human melanoma cells susceptible to stem cell factor-induced apoptosis and inhibits their tumorigenic and metastatic potential. Oncogene 13: 2339–2347.
Igoucheva O, Alexeev V . (2009). MicroRNA-dependent regulation of cKit in cutaneous melanoma. Biochem Biophys Res Commun 379: 790–794.
Jazdzewski K, Liyanarachchi S, Swierniak M, Pachucki J, Ringel MD, Jarzab B et al. (2009). Polymorphic mature microRNAs from passenger strand of pre-miR-146a contribute to thyroid cancer. Proc Natl Acad Sci USA 106: 1502–1505.
Jiang X, Zhou J, Yuen NK, Corless CL, Heinrich MC, Fletcher JA et al. (2008). Imatinib targeting of KIT-mutant oncoprotein in melanoma. Clin Cancer Res 14: 7726–7732.
Koga Y, Pelizzola M, Cheng E, Krauthammer M, Sznol M, Ariyan S et al. (2009). Genome Res 19: 1462–1470.
Lassam N, Bickford S . (1992). Loss of c-kit expression in cultured melanoma cells. Oncogene 7: 51–56.
Lee CT, Risom T, Strauss WM . (2007). Evolutionary conservation of microRNA regulatory circuits: an examination of microRNA gene complexity and conserved microRNA-target interactions through metazoan phylogeny. DNA Cell Biol 26: 209–218.
Lutzky J, Bauer J, Bastian BC . (2008). Dose-dependent, complete response to imatinib of a metastatic mucosal melanoma with a K642E KIT mutation. Pigment Cell Melanoma Res 21: 492–493.
Medina PP, Slack FJ . (2008). microRNAs and cancer: an overview. Cell Cycle 7: 2485–2492.
Meyle KD, Guldberg P . (2009). Genetic risk factors for melanoma. Hum Genet 126: 499–510.
Mishra PJ, Bertino JR . (2009). MicroRNA polymorphisms: the future of pharmacogenomics, molecular epidemiology and individualized medicine. Pharmacogenomics 10: 399–416.
Monsel G, Ortonne N, Bagot M, Bensussan A, Dumaz N. . (2009). c-Kit mutants require hypoxia-inducible factor 1alpha to transform melanocytes. Oncogene 29: 227–236.
Natali PG, Nicotra MR, Winkler AB, Cavaliere R, Bigotti A, Ullrich A . (1992). Progression of human cutaneous melanoma is associated with loss of expression of c-kit proto-oncogene receptor. Int J Cancer 52: 197–201.
Rajeevan H, Cheung KH, Gadagkar R, Stein S, Soundararajan U, Kidd JR et al. (2005). ALFRED: an allele frequency database for microevolutionary studies. Evol Bioinform Online 1: 1–10.
Ratner E, Lu L, Boeke M, Barnett R, Nallur S, Chin LJ et al. (2010). A KRAS-variant in ovarian cancer acts as a genetic marker of cancer risk. Cancer Res 70: 6509–6515.
Rigel DS . (2010). Trends in dermatology: melanoma incidence. Arch Dermatol 146: 318.
Saetrom P, Biesinger J, Li SM, Smith D, Thomas LF, Majzoub K et al. (2009). A risk variant in an miR-125b binding site in BMPR1B is associated with breast cancer pathogenesis. Cancer Res 69: 7459–7465.
Sayers EW, Barrett T, Benson DA, Bryant SH, Canese K, Chetvernin V et al. (2009). Database resources of the National Center for Biotechnology Information. Nucleic Acids Res 37: D5–15.
Sherry ST, Ward MH, Kholodov M, Baker J, Phan L, Smigielski EM et al. (2001). dbSNP: the NCBI database of genetic variation. Nucleic Acids Res 29: 308–311.
Smalley KS, Contractor R, Nguyen TK, Xiao M, Edwards R, Muthusamy V et al. (2008). Identification of a novel subgroup of melanomas with KIT/cyclin-dependent kinase-4 overexpression. Cancer Res 68: 5743–5752.
Smalley KS, Nathanson KL, Flaherty KT . (2009a). Genetic subgrouping of melanoma reveals new opportunities for targeted therapy. Cancer Res 69: 3241–3244.
Smalley KS, Sondak VK, Weber JS . (2009b). c-KIT signaling as the driving oncogenic event in sub-groups of melanomas. Histol Histopathol 24: 643–650.
Speed WC, Kang SP, Tuck DP, Harris LN, Kidd KK. (2009). Global variation in CYP2C8–CYP2C9 functional haplotypes. Pharmacogenomics J 9: 283–290.
Stefani G . (2007). Roles of microRNAs and their targets in cancer. Expert Opin Biol Ther 7: 1833–1840.
Went PT, Dirnhofer S, Bundi M, Mirlacher M, Schraml P, Mangialaio S et al. (2004). Prevalence of KIT expression in human tumors. J Clin Oncol 22: 4514–4522.
Zakut R, Perlis R, Eliyahu S, Yarden Y, Givol D, Lyman SD et al. (1993). KIT ligand (mast cell growth factor) inhibits the growth of KIT-expressing melanoma cells. Oncogene 8: 2221–2229.
Acknowledgements
This work was supported by Yale SPORE in Skin Cancer funded by the National Cancer Institute grant number 1 P50 CA121974 (R Halaban, PI) and by a generous gift from Milstein–Meyer Center for Melanoma Research. JW is supported by K08 CA124484. AMM is supported by the CTSA Grant number UL1 RR024139 from the National Center for Research Resources (NCRR), a component of the National Institutes of Health (NIH), and NIH roadmap for Medical Research. The testing of the samples from the several world populations was supported by 1 P01 GM057672 (KK Kidd, PI).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Godshalk, S., Paranjape, T., Nallur, S. et al. A Variant in a MicroRNA complementary site in the 3′ UTR of the KIT oncogene increases risk of acral melanoma. Oncogene 30, 1542–1550 (2011). https://doi.org/10.1038/onc.2010.536
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2010.536
Keywords
This article is cited by
-
Emerging Biomarkers in Cutaneous Melanoma
Molecular Diagnosis & Therapy (2018)
-
A human 3′UTR clone collection to study post-transcriptional gene regulation
BMC Genomics (2015)
-
MicroRNA-mediated regulation of KRAS in cancer
Journal of Hematology & Oncology (2014)
-
Control by a hair’s breadth: the role of microRNAs in the skin
Cellular and Molecular Life Sciences (2013)