Abstract
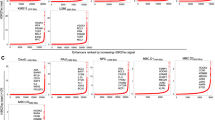
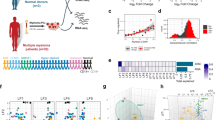
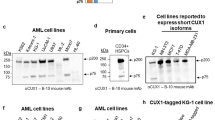
Dysregulation of cyclin D2 contributes to the pathogenesis of multiple myeloma, and can occur through translocations that activate MAF/MAFB or MMSET/FGFR3. However, cyclin D2 induction can also be seen in the absence of such translocations, such as in patients with hyperdiploid disease, through unknown mechanisms. In UniGene cluster data-mining and ECgene analysis, we found that zinc-finger with KRAB and SCAN domains 3 (ZKSCAN3), a novel transcription factor, is overrepresented in this malignancy, and three consensus ZKSCAN3 binding sites were found in the cyclin D2 promoter. Analysis of a panel of myeloma cell lines, primary patient samples and datasets from Oncomine and the Multiple Myeloma Genomics Portal (MMGP) revealed expression of ZKSCAN3 messenger RNA (mRNA) in a majority of samples. Studies of cell lines by western blotting, and of primary tissue microarrays by immunohistochemistry, showed ZKSCAN3 protein expression in a majority, and in a manner that paralleled messenger levels in cell lines. ZKSCAN3 overexpression was associated with increased gene copy number or genomic DNA gain/amplification in a subset based on analysis of data from the MMGP, and from fluorescence in situ hybridization studies of cell lines and primary samples. Overexpression of ZKSCAN3 induced cyclin D2 promoter activity in a MAF/MAFB-independent manner, and to an extent that was influenced by the number of consensus ZKSCAN3 binding sites. Moreover, ZKSCAN3 protein expression correlated with cyclin D2 levels in cell lines and primary samples, and its overexpression induced cyclin D2. Conversely, ZKSCAN3 suppression using small hairpin RNAs (shRNAs) reduced cyclin D2 levels, and, importantly, inhibited myeloma cell line proliferation. Finally, ZKSCAN3 was noted to specifically bind to oligonucleotides representing sequences from the cyclin D2 promoter, and to the endogenous promoter itself in myeloma cells. Taken together, the data support the conclusion that ZKSCAN3 induction represents a mechanism by which myeloma cells can induce cyclin D2 dysregulation, and contribute to disease pathogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Arora T, Jelinek DF . (1998). Differential myeloma cell responsiveness to interferon-alpha correlates with differential induction of p19(INK4d) and cyclin D2 expression. J Biol Chem 273: 11799–11805.
Bergsagel PL, Kuehl WM . (2005). Molecular pathogenesis and a consequent classification of multiple myeloma. J Clin Oncol 23: 6333–6338.
Bergsagel PL, Kuehl WM, Zhan F, Sawyer J, Barlogie B, Shaughnessy Jr J . (2005). Cyclin D dysregulation: an early and unifying pathogenic event in multiple myeloma. Blood 106: 296–303.
Carrasco DR, Tonon G, Huang Y, Zhang Y, Sinha R, Feng B et al. (2006). High-resolution genomic profiles define distinct clinico-pathogenetic subgroups of multiple myeloma patients. Cancer Cell 9: 313–325.
Chang H, Qi X, Trieu Y, Xu W, Reader JC, Ning Y et al. (2006). Multiple myeloma patients with CKS1B gene amplification have a shorter progression-free survival post-autologous stem cell transplantation. Br J Haematol 135: 486–491.
Chesi M, Bergsagel PL, Brents LA, Smith CM, Gerhard DS, Kuehl WM . (1996). Dysregulation of cyclin D1 by translocation into an IgH gamma switch region in two multiple myeloma cell lines. Blood 88: 674–681.
Chesi M, Bergsagel PL, Shonukan OO, Martelli ML, Brents LA, Chen T et al. (1998a). Frequent dysregulation of the c-maf proto-oncogene at 16q23 by translocation to an Ig locus in multiple myeloma. Blood 91: 4457–4463.
Chesi M, Nardini E, Brents LA, Schrock E, Ried T, Kuehl WM et al. (1997). Frequent translocation t(4;14)(p16.3;q32.3) in multiple myeloma is associated with increased expression and activating mutations of fibroblast growth factor receptor 3. Nat Genet 16: 260–264.
Chesi M, Nardini E, Lim RS, Smith KD, Kuehl WM, Bergsagel PL . (1998b). The t(4;14) translocation in myeloma dysregulates both FGFR3 and a novel gene, MMSET, resulting in IgH/MMSET hybrid transcripts. Blood 92: 3025–3034.
Ferti A, Panani A, Arapakis G, Raptis S . (1984). Cytogenetic study in multiple myeloma. Cancer Genet Cytogenet 12: 247–253.
Gabrea A, Bergsagel PL, Chesi M, Shou Y, Kuehl WM . (1999). Insertion of excised IgH switch sequences causes overexpression of cyclin D1 in a myeloma tumor cell. Mol Cell 3: 119–123.
Gutierrez NC, Garcia JL, Hernandez JM, Lumbreras E, Castellanos M, Rasillo A et al. (2004). Prognostic and biologic significance of chromosomal imbalances assessed by comparative genomic hybridization in multiple myeloma. Blood 104: 2661–2666.
Hideshima T, Mitsiades C, Tonon G, Richardson PG, Anderson KC . (2007). Understanding multiple myeloma pathogenesis in the bone marrow to identify new therapeutic targets. Nat Rev Cancer 7: 585–598.
Hurt EM, Wiestner A, Rosenwald A, Shaffer AL, Campo E, Grogan T et al. (2004). Overexpression of c-maf is a frequent oncogenic event in multiple myeloma that promotes proliferation and pathological interactions with bone marrow stroma. Cancer Cell 5: 191–199.
Kuhn DJ, Chen Q, Voorhees PM, Strader JS, Shenk KD, Sun CM et al. (2007). Potent activity of carfilzomib, a novel, irreversible inhibitor of the ubiquitin-proteasome pathway, against preclinical models of multiple myeloma. Blood 110: 3281–3290.
Kumar S, Witzig TE, Timm M, Haug J, Wellik L, Fonseca R et al. (2003). Expression of VEGF and its receptors by myeloma cells. Leukemia 17: 2025–2031.
Kyle RA, Rajkumar SV . (2008). Multiple myeloma. Blood 111: 2962–2972.
Largo C, Saez B, Alvarez S, Suela J, Ferreira B, Blesa D et al. (2007). Multiple myeloma primary cells show a highly rearranged unbalanced genome with amplifications and homozygous deletions irrespective of the presence of immunoglobulin-related chromosome translocations. Haematologica 92: 795–802.
Le Gouill S, Podar K, Amiot M, Hideshima T, Chauhan D, Ishitsuka K et al. (2004). VEGF induces Mcl-1 up-regulation and protects multiple myeloma cells against apoptosis. Blood 104: 2886–2892.
Lu G, Kong Y, Yue C . (2010). Genetic and immunophenotypic profile of IGH@ rearrangement detected by fluorescence in situ hybridization in 149 cases of B-cell chronic lymphocytic leukemia. Cancer Genet Cytogenet 196: 56–63.
Markovic O, Marisavljevic D, Cemerikic V, Vidovic A, Perunicic M, Todorovic M et al. (2008). Expression of VEGF and microvessel density in patients with multiple myeloma: clinical and prognostic significance. Med Oncol 25: 451–457.
Perez-Simon JA, Garcia-Sanz R, Tabernero MD, Almeida J, Gonzalez M, Fernandez-Calvo J et al. (1998). Prognostic value of numerical chromosome aberrations in multiple myeloma: a FISH analysis of 15 different chromosomes. Blood 91: 3366–3371.
Podar K, Tai YT, Davies FE, Lentzsch S, Sattler M, Hideshima T et al. (2001). Vascular endothelial growth factor triggers signaling cascades mediating multiple myeloma cell growth and migration. Blood 98: 428–435.
Ria R, Vacca A, Russo F, Cirulli T, Massaia M, Tosi P et al. (2004). A VEGF-dependent autocrine loop mediates proliferation and capillarogenesis in bone marrow endothelial cells of patients with multiple myeloma. Thromb Haemost 92: 1438–1445.
Santos GC, Zielenska M, Prasad M, Squire JA . (2007). Chromosome 6p amplification and cancer progression. J Clin Pathol 60: 1–7.
Shaughnessy Jr J, Gabrea A, Qi Y, Brents L, Zhan F, Tian E et al. (2001). Cyclin D3 at 6p21 is dysregulated by recurrent chromosomal translocations to immunoglobulin loci in multiple myeloma. Blood 98: 217–223.
Voorhees PM, Chen Q, Kuhn DJ, Small GW, Hunsucker SA, Strader JS et al. (2007). Inhibition of interleukin-6 signaling with CNTO 328 enhances the activity of bortezomib in preclinical models of multiple myeloma. Clin Cancer Res 13: 6469–6478.
Yang L, Hamilton SR, Sood A, Kuwai T, Ellis L, Sanguino A et al. (2008a). The previously undescribed ZKSCAN3 (ZNF306) is a novel ‘driver’ of colorectal cancer progression. Cancer Res 68: 4321–4330.
Yang L, Zhang L, Wu Q, Boyd DD . (2008b). Unbiased Screening for Transcriptional Targets of ZKSCAN3 Identifies Integrin {beta}4 and Vascular Endothelial Growth Factor as Downstream Targets. J Biol Chem 283: 35295–35304.
Acknowledgements
We wish to thank Dr Douglas Boyd for his contributions to the studies of the role of ZKSCAN3 in colon cancer. RZO, a Leukemia & Lymphoma Society Scholar in Clinical Research, would like to acknowledge support from the Leukemia & Lymphoma Society (6096-07) and the National Cancer Institute (R01 CA102278).
Authorship: LY designed and performed the research presented herein, and wrote the paper. HW aided in performing Lentiviral experiments. SK, DAG, JAM, MW, DMW, SKT and JJS were involved in patient sample acquisition, including in the consenting of patients, and later purification and storage of their plasma cells. NZ, LZ, SYY and KAB performed bioinformatics analyses, whereas GL, MZ and RM performed FISH studies. RZO supervised the design and performance of all of the research contained herein, and contributed to the writing and editing of the paper.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Yang, L., Wang, H., Kornblau, S. et al. Evidence of a role for the novel zinc-finger transcription factor ZKSCAN3 in modulating Cyclin D2 expression in multiple myeloma. Oncogene 30, 1329–1340 (2011). https://doi.org/10.1038/onc.2010.515
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2010.515
Keywords
This article is cited by
-
Zkscan3 affects erythroblast development by regulating the transcriptional activity of GATA1 and KLF1 in mice
Journal of Molecular Histology (2022)
-
ZKSCAN3 in severe bacterial lung infection and sepsis-induced immunosuppression
Laboratory Investigation (2021)
-
ZKSCAN3 drives tumor metastasis via integrin β4/FAK/AKT mediated epithelial–mesenchymal transition in hepatocellular carcinoma
Cancer Cell International (2020)
-
Human ZKSCAN3 and Drosophila M1BP are functionally homologous transcription factors in autophagy regulation
Scientific Reports (2020)
-
A30P mutant α-synuclein impairs autophagic flux by inactivating JNK signaling to enhance ZKSCAN3 activity in midbrain dopaminergic neurons
Cell Death & Disease (2019)