Abstract
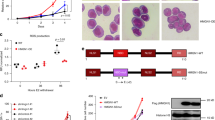
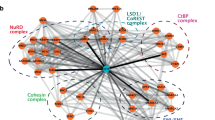
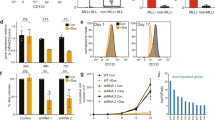
MOZ and MLL, encoding a histone acetyltransferase (HAT) and a histone methyltransferase, respectively, are targets for recurrent chromosomal translocations found in acute myeloblastic or lymphoblastic leukemia. In MOZ (MOnocytic leukemia Zinc-finger protein)/CBP- or mixed lineage leukemia (MLL)-rearranged leukemias, abnormal levels of HOX transcription factors have been found to be critical for leukemogenesis. We show that MOZ and MLL cooperate to regulate these key genes in human cord blood CD34+ cells. These chromatin-modifying enzymes interact, colocalize and functionally cooperate, and both are recruited to multiple HOX promoters. We also found that WDR5, an adaptor protein essential for lysine 4 trimethylation of histone H3 (H3K4me3) by MLL, colocalizes and interacts with MOZ. We detected the binding of the HAT MOZ to H3K4me3, thus linking histone methylation to acetylation. In CD34+ cells, depletion of MLL causes release of MOZ from HOX promoters, which is correlated to defective histone activation marks, leading to repression of HOX gene expression and alteration of commitment of CD34+ cells into myeloid progenitors. Thus, our results unveil the role of the interaction between MOZ and MLL in CD34+ cells in which both proteins have a critical role in hematopoietic cell-fate decision, suggesting a new molecular mechanism by which MOZ or MLL deregulation leads to leukemogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Argiropoulos B, Humphries RK . (2007). Hox genes in hematopoiesis and leukemogenesis. Oncogene 26: 6766–6776.
Ayton PM, Cleary ML . (2003). Transformation of myeloid progenitors by MLL oncoproteins is dependent on Hoxa7 and Hoxa9. Genes Dev 17: 2298–2307.
Borrow J, Shearman AM, Stanton Jr VP, Becher R, Collins T, Williams AJ et al. (1996a). The t(7;11)(p15;p15) translocation in acute myeloid leukaemia fuses the genes for nucleoporin NUP98 and class I homeoprotein HOXA9. Nat Genet 12: 159–167.
Borrow J, Stanton Jr VP, Andresen JM, Becher R, Behm FG, Chaganti RS et al. (1996b). The translocation t(8;16)(p11;p13) of acute myeloid leukaemia fuses a putative acetyltransferase to the CREB-binding protein. Nat Genet 14: 33–41.
Bristow CA, Shore P . (2003). Transcriptional regulation of the human MIP-1alpha promoter by RUNX1 and MOZ. Nucleic Acids Res 31: 2735–2744.
Camos M, Esteve J, Jares P, Colomer D, Rozman M, Villamor N et al. (2006). Gene expression profiling of acute myeloid leukemia with translocation t(8;16)(p11;p13) and MYST3-CREBBP rearrangement reveals a distinctive signature with a specific pattern of HOX gene expression. Cancer Res 66: 6947–6954.
Carrozza MJ, Utley RT, Workman JL, Cote J . (2003). The diverse functions of histone acetyltransferase complexes. Trends Genet 19: 321–329.
Champagne N, Pelletier N, Yang XJ . (2001). The monocytic leukemia zinc finger protein MOZ is a histone acetyltransferase. Oncogene 20: 404–409.
Chan EM, Chan RJ, Comer EM, Goulet III RJ, Crean CD, Brown ZD et al. (2007). MOZ and MOZ-CBP cooperate with NF-kappaB to activate transcription from NF-kappaB-dependent promoters. Exp Hematol 35: 1782–1792.
Crooks GM, Fuller J, Petersen D, Izadi P, Malik P, Pattengale PK et al. (1999). Constitutive HOXA5 expression inhibits erythropoiesis and increases myelopoiesis from human hematopoietic progenitors. Blood 94: 519–528.
Dion MF, Altschuler SJ, Wu LF, Rando OJ . (2005). Genomic characterization reveals a simple histone H4 acetylation code. Proc Natl Acad Sci USA 102: 5501–5506.
Dou Y, Milne TA, Tackett AJ, Smith ER, Fukuda A, Wysocka J et al. (2005). Physical association and coordinate function of the H3 K4 methyltransferase MLL1 and the H4 K16 acetyltransferase MOF. Cell 121: 873–885.
Eberharter A, Becker PB . (2002). Histone acetylation: a switch between repressive and permissive chromatin. Second in review series on chromatin dynamics. EMBO Rep 3: 224–229.
Ernst P, Mabon M, Davidson AJ, Zon LI, Korsmeyer SJ . (2004). An Mll-dependent Hox program drives hematopoietic progenitor expansion. Curr Biol 14: 2063–2069.
Ernst P, Wang J, Huang M, Goodman RH, Korsmeyer SJ . (2001). MLL and CREB bind cooperatively to the nuclear coactivator CREB-binding protein. Mol Cell Biol 21: 2249–2258.
Esteyries S, Perot C, Adelaide J, Imbert M, Lagarde A, Pautas C et al. (2008). NCOA3, a new fusion partner for MOZ/MYST3 in M5 acute myeloid leukemia. Leukemia 22: 663–665.
Golub TR, Slonim DK, Tamayo P, Huard C, Gaasenbeek M, Mesirov JP et al. (1999). Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286: 531–537.
Guccione E, Bassi C, Casadio F, Martinato F, Cesaroni M, Schuchlautz H et al. (2007). Methylation of histone H3R2 by PRMT6 and H3K4 by an MLL complex are mutually exclusive. Nature 449: 933–937.
Hake SB, Xiao A, Allis CD . (2004). Linking the epigenetic ‘language’ of covalent histone modifications to cancer. Br J Cancer 90: 761–769.
Hanson RD, Hess JL, Yu BD, Ernst P, van Lohuizen M, Berns A et al. (1999). Mammalian Trithorax and polycomb-group homologues are antagonistic regulators of homeotic development. Proc Natl Acad Sci USA 96: 14372–14377.
Hess JL . (2004). MLL: a histone methyltransferase disrupted in leukemia. Trends Mol Med 10: 500–507.
Hsieh JJ, Cheng EH, Korsmeyer SJ . (2003). Taspase1: a threonine aspartase required for cleavage of MLL and proper HOX gene expression. Cell 115: 293–303.
Huang G, Elf S, Yan X, Wang L, Liu Y, Sashida G et al. (2008). Previously unknown interactions between AML1 and MLL provide epigenetic regulation of gene expression in normal hematopoiesis and in leukemia. Blood 112: 110.
Huang Y, Fang J, Bedford MT, Zhang Y, Xu RM . (2006). Recognition of histone H3 lysine-4 methylation by the double tudor domain of JMJD2A. Science 312: 748–751.
Jenuwein T, Allis CD . (2001). Translating the histone code. Science 293: 1074–1080.
Jude CD, Climer L, Xu D, Artinger E, Fisher JK, Ernst P . (2007). Unique and independent roles for MLL in adult hematopoietic stem cells and progenitors. Cell Stem Cell 1: 324–337.
Katsumoto T, Aikawa Y, Iwama A, Ueda S, Ichikawa H, Ochiya T et al. (2006). MOZ is essential for maintenance of hematopoietic stem cells. Genes Dev 20: 1321–1330.
Kindle KB, Troke PJ, Collins HM, Matsuda S, Bossi D, Bellodi C et al. (2005). MOZ-TIF2 inhibits transcription by nuclear receptors and p53 by impairment of CBP function. Mol Cell Biol 25: 988–1002.
Kitabayashi I, Aikawa Y, Nguyen LA, Yokoyama A, Ohki M . (2001). Activation of AML1-mediated transcription by MOZ and inhibition by the MOZ-CBP fusion protein. EMBO J 20: 7184–7196.
Knittel T, Kessel M, Kim MH, Gruss P . (1995). A conserved enhancer of the human and murine Hoxa-7 gene specifies the anterior boundary of expression during embryonal development. Development 121: 1077–1088.
Kohlmann A, Schoch C, Dugas M, Schnittger S, Hiddemann W, Kern W et al. (2005). New insights into MLL gene rearranged acute leukemias using gene expression profiling: shared pathways, lineage commitment, and partner genes. Leukemia 19: 953–964.
Kouzarides T . (2007). Chromatin modifications and their function. Cell 128: 693–705.
Krivtsov AV, Armstrong SA . (2007). MLL translocations, histone modifications and leukaemia stem-cell development. Nat Rev Cancer 7: 823–833.
McGinnis W, Krumlauf R . (1992). Homeobox genes and axial patterning. Cell 68: 283–302.
McMahon KA, Hiew SY, Hadjur S, Veiga-Fernandes H, Menzel U, Price AJ et al. (2007). Mll has a critical role in fetal and adult hematopoietic stem cell self-renewal. Cell Stem Cell 1: 338–345.
Milne TA, Briggs SD, Brock HW, Martin ME, Gibbs D, Allis CD et al. (2002). MLL targets SET domain methyltransferase activity to Hox gene promoters. Mol Cell 10: 1107–1117.
Milne TA, Dou Y, Martin ME, Brock HW, Roeder RG, Hess JL . (2005). MLL associates specifically with a subset of transcriptionally active target genes. Proc Natl Acad Sci USA 102: 14765–14770.
Nabirochkina E, Simonova OB, Mertsalov IB, Kulikova DA, Ladigina NG, Korochkin LI et al. (2002). Expression pattern of dd4, a sole member of the d4 family of transcription factors in drosophila melanogaster. Mech Dev 114: 119–123.
Nakamura T, Largaespada DA, Shaughnessy Jr JD, Jenkins NA, Copeland NG . (1996). Cooperative activation of Hoxa and Pbx1-related genes in murine myeloid leukaemias. Nat Genet 12: 149–153.
Nakamura T, Mori T, Tada S, Krajewski W, Rozovskaia T, Wassell R et al. (2002). ALL-1 is a histone methyltransferase that assembles a supercomplex of proteins involved in transcriptional regulation. Mol Cell 10: 1119–1128.
Ohta K, Ohigashi M, Naganawa A, Ikeda H, Sakai M, Nishikawa J et al. (2007). Histone acetyltransferase MOZ acts as a co-activator of Nrf2-MafK and induces tumour marker gene expression during hepatocarcinogenesis. Biochem J 402: 559–566.
Ohta K, Osada S, Nishikawa JI, Nishihara T . (2005). J Health Science 51: 253–256.
Pelletier N, Champagne N, Stifani S, Yang XJ . (2002). MOZ and MORF histone acetyltransferases interact with the Runt-domain transcription factor Runx2. Oncogene 21: 2729–2740.
Perez-Campo FM, Borrow J, Kouskoff V, Lacaud G . (2009). The histone acetyl transferase activity of monocytic leukemia zinc finger is critical for the proliferation of hematopoietic precursors. Blood 113: 4866–4874.
Rokudai S, Aikawa Y, Tagata Y, Tsuchida N, Taya Y, Kitabayashi I . (2009). Monocytic leukemia zinc finger (MOZ) interacts with p53 to induce p21 expression and cell-cycle arrest. J Biol Chem 284: 237–244.
Rozenblatt-Rosen O, Rozovskaia T, Burakov D, Sedkov Y, Tillib S, Blechman J et al. (1998). The C-terminal SET domains of ALL-1 and TRITHORAX interact with the INI1 and SNR1 proteins, components of the SWI/SNF complex. Proc Natl Acad Sci USA 95: 4152–4157.
Ruthenburg AJ, Allis CD, Wysocka J . (2007). Methylation of lysine 4 on histone H3: intricacy of writing and reading a single epigenetic mark. Mol Cell 25: 15–30.
Santos-Rosa H, Schneider R, Bannister AJ, Sherriff J, Bernstein BE, Emre NC et al. (2002). Active genes are tri-methylated at K4 of histone H3. Nature 419: 407–411.
Sauvageau G, Lansdorp PM, Eaves CJ, Hogge DE, Dragowska WH, Reid DS et al. (1994). Differential expression of homeobox genes in functionally distinct CD34+ subpopulations of human bone marrow cells. Proc Natl Acad Sci USA 91: 12223–12227.
Shen WF, Largman C, Lowney P, Corral JC, Detmer K, Hauser CA et al. (1989). Differential expression of homeobox genes in functionally distinct CD34+ subpopulations of human bone marrow cells. Proc Natl Acad Sci USA 86: 8536–8540.
Sims III RJ, Chen CF, Santos-Rosa H, Kouzarides T, Patel SS, Reinberg D . (2005). Human but not yeast CHD1 binds directly and selectively to histone H3 methylated at lysine 4 via its tandem chromodomains. J Biol Chem 280: 41789–41792.
Terranova R, Agherbi H, Boned A, Meresse S, Djabali M . (2006). Histone and DNA methylation defects at Hox genes in mice expressing a SET domain-truncated form of Mll. Proc Natl Acad Sci USA 103: 6629–6634.
Thomas T, Corcoran LM, Gugasyan R, Dixon MP, Brodnicki T, Nutt SL et al. (2006). Monocytic leukemia zinc finger protein is essential for the development of long-term reconstituting hematopoietic stem cells. Genes Dev 20: 1175–1186.
Tyagi S, Chabes AL, Wysocka J, Herr W . (2007). E2F activation of S phase promoters via association with HCF-1 and the MLL family of histone H3K4 methyltransferases. Mol Cell 27: 107–119.
Ullah M, Pelletier N, Xiao L, Zhao SP, Wang K, Degerny C et al. (2008). Molecular architecture of quartet MOZ/MORF histone acetyltransferase complexes. Mol Cell Biol 28: 6828–6843.
van Oostveen J, Bijl J, Raaphorst F, Walboomers J, Meijer C . (1999). The role of homeobox genes in normal hematopoiesis and hematological malignancies. Leukemia 13: 1675–1690.
Voss AK, Collin C, Dixon MP, Thomas T . (2009). Moz and retinoic acid coordinately regulate H3K9 acetylation, Hox gene expression, and segment identity. Dev Cell 17: 674–686.
Wysocka J, Swigut T, Milne TA, Dou Y, Zhang X, Burlingame AL et al. (2005). WDR5 associates with histone H3 methylated at K4 and is essential for H3 K4 methylation and vertebrate development. Cell 121: 859–872.
Wysocka J, Swigut T, Xiao H, Milne TA, Kwon SY, Landry J et al. (2006). A PHD finger of NURF couples histone H3 lysine 4 trimethylation with chromatin remodelling. Nature 442: 86–90.
Yang XJ . (2004). The diverse superfamily of lysine acetyltransferases and their roles in leukemia and other diseases. Nucleic Acids Res 32: 959–976.
Yang XJ, Ullah M . (2007). MOZ and MORF, two large MYSTic HATs in normal and cancer stem cells. Oncogene 26: 5408–5419.
Yu BD, Hess JL, Horning SE, Brown GA, Korsmeyer SJ . (1995). Altered Hox expression and segmental identity in Mll-mutant mice. Nature 378: 505–508.
Acknowledgements
We gratefully acknowledge Amandine Bataille, Amandine Chlemaire and Franck Ménétrier for immunocytofluorescence assays; André Bouchot for epifluorescence microscopy; and Christine Arnould for confocal analysis (SERCOBIO, Université de Bourgogne), and the Etablissement Français du Sang (EFS) of Bourgogne Franche-Comté, which kindly supplied the cord blood buffy coats. We also thank Mustapha Oulad-Abdelghani for producing the anti-MOZ antibody. We thank Robert Slany, Issai Kitabayashi, Paul Shore, Michael Cleary and Edward Chan for providing plasmids. We appreciate the fine work performed by Magali Belt in correction of English text. This work was supported by funds from the Fondation de France (Leukemia Committee to LD), the Ligue contre le Cancer (Côte d’Or committee to LD), the Ligue contre le Cancer (Rhône committee to LD), the Conseil Régional de Bourgogne (FABER to LD), the Faculty of Medicine (to J-NB) in Dijon, the National Institute of Cancer (to ES), the Ligue Nationale contre le Cancer (Label to ES), the Agence Nationale de la Recherche of France (to ES and LD) and the Canadian Cancer Society (to XJY). JP was supported by fellowships from the Ministère de l’Enseignement Supérieur et de la Recherche of France and the Association pour la Recherche sur le Cancer (ARC). AL was supported by fellowships from the Inserm and the Région Bourgogne. RA and AJ were supported by fellowships from the Ligue contre le Cancer, Saône-et-Loire committee and national committee, respectively. BL was supported by fellowships from the Ministère de l’Enseignement Supérieur et de la Recherche of France.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Paggetti, J., Largeot, A., Aucagne, R. et al. Crosstalk between leukemia-associated proteins MOZ and MLL regulates HOX gene expression in human cord blood CD34+ cells. Oncogene 29, 5019–5031 (2010). https://doi.org/10.1038/onc.2010.254
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2010.254
Keywords
This article is cited by
-
The many lives of KATs — detectors, integrators and modulators of the cellular environment
Nature Reviews Genetics (2019)
-
Why are so many MLL lysine methyltransferases required for normal mammalian development?
Cellular and Molecular Life Sciences (2019)
-
The basic helix-loop-helix transcription factor SHARP1 is an oncogenic driver in MLL-AF6 acute myelogenous leukemia
Nature Communications (2018)
-
C-terminal BRE overexpression in 11q23-rearranged and t(8;16) acute myeloid leukemia is caused by intragenic transcription initiation
Leukemia (2018)
-
Complementary activities of DOT1L and Menin inhibitors in MLL-rearranged leukemia
Leukemia (2017)