Abstract
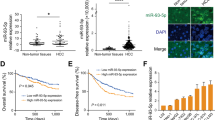
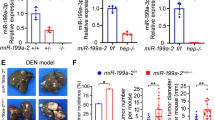
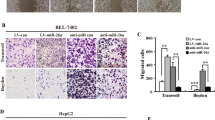
To identify microRNAs (miRNAs) that may have a causal role in hepatocarcinogenesis, we used an animal model in which C57BL/6 mice fed choline-deficient and amino acid defined (CDAA) diet develop preneoplastic lesions at 65 weeks and hepatocellular carcinomas after 84 weeks. miRNA expression profiling showed significant upregulation of miR-181b and miR-181d in the livers of mice as early as 32 weeks that persisted at preneoplastic stage. The expression of tissue inhibitor of metalloprotease 3 (TIMP3), a tumor suppressor and a validated miR-181 target, was markedly suppressed in the livers of mice fed CDAA diet. Upregulation of hepatic transforming growth factor (TGF)β and its downstream mediators Smad 2, 3 and 4 and increase in phospho-Smad2 in the liver nuclear extract correlated with elevated miR-181b/d in mice fed CDAA diet. The levels of the precursor and mature miR-181b were augmented on exposure of hepatic cells to TGFβ and were significantly reduced by small interference RNA-mediated depletion of Smad4, showing the involvement of TGFβ signaling pathway in miR-181b expression. Ectopic expression and depletion of miR-181b showed that miR-181b enhanced matrix metallopeptidases (MMP)2 and MMP9 activity and promoted growth, clonogenic survival, migration and invasion of hepatocellular carcinoma (HCC) cells that could be reversed by modulating TIMP3 level. Further, depletion of miR-181b inhibited tumor growth of HCC cells in nude mice. miR-181b also enhanced resistance of HCC cells to the anticancer drug doxorubicin. On the basis of these results, we conclude that upregulation of miR-181b at early stages of feeding CDAA diet promotes hepatocarcinogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Aravalli RN, Steer CJ, Cressman EN . (2008). Molecular mechanisms of hepatocellular carcinoma. Hepatology 48: 2047–2063.
Bartel DP . (2009). MicroRNAs: target recognition and regulatory functions. Cell 136: 215–233.
Calin GA, Pekarsky Y, Croce CM . (2007). The role of microRNA and other non-coding RNA in the pathogenesis of chronic lymphocytic leukemia. Best Pract Res Clin Haematol 20: 425–437.
Carthew RW, Sontheimer EJ . (2009). Origins and mechanisms of miRNAs and siRNAs. Cell 136: 642–655.
Cheung O, Puri P, Eicken C, Contos MJ, Mirshahi F, Maher JW et al. (2008). Nonalcoholic steatohepatitis is associated with altered hepatic microRNA expression. Hepatology 48: 1810–1820.
Coulouarn C, Factor VM, Thorgeirsson SS . (2008). Transforming growth factor-beta gene expression signature in mouse hepatocytes predicts clinical outcome in human cancer. Hepatology 47: 2059–2067.
Datta J, Kutay H, Nasser MW, Nuovo GJ, Wang B, Majumder S et al. (2008). Methylation mediated silencing of microRNA-1 gene and its role in hepatocellular carcinogenesis. Cancer Res 68: 5049–5058.
Davis BN, Hilyard AC, Lagna G, Hata A . (2008). SMAD proteins control DROSHA-mediated microRNA maturation. Nature 454: 56–61.
Day CP . (2006). Genes or environment to determine alcoholic liver disease and non-alcoholic fatty liver disease. Liver Int 26: 1021–1028.
Denda A, Kitayama W, Kishida H, Murata N, Tsutsumi M, Tsujiuchi T et al. (2002). Development of hepatocellular adenomas and carcinomas associated with fibrosis in C57BL/6J male mice given a choline-deficient, L-amino acid-defined diet. Jpn J Cancer Res 93: 125–132.
El-Serag HB, Rudolph KL . (2007). Hepatocellular carcinoma: epidemiology and molecular carcinogenesis. Gastroenterology 132: 2557–2576.
Elmen J, Lindow M, Schutz S, Lawrence M, Petri A, Obad S et al. (2008). LNA-mediated microRNA silencing in non-human primates. Nature 452: 896–899.
Esau C, Davis S, Murray SF, Yu XX, Pandey SK, Pear M et al. (2006). miR-122 regulation of lipid metabolism revealed by in vivo antisense targeting. Cell Metab 3: 87–98.
Eulalio A, Huntzinger E, Izaurralde E . (2008). Getting to the root of miRNA-mediated gene silencing. Cell 132: 9–14.
Gabriely G, Wurdinger T, Kesari S, Esau CC, Burchard J, Linsley PS et al. (2008). MicroRNA 21 promotes glioma invasion by targeting matrix metalloproteinase regulators. Mol Cell Biol 28: 5369–5380.
Ghoshal K, Li X, Datta J, Bai S, Pogribny I, Pogribny M et al. (2006). A folate- and methyl-deficient diet alters the expression of DNA methyltransferases and methyl CpG binding proteins involved in epigenetic gene silencing in livers of F344 rats. J Nutr 136: 1522–1527.
Ji J, Yamashita T, Budhu A, Forgues M, Jia HL, Li C et al. (2009). Identification of microRNA-181 by genome-wide screening as a critical player in EpCAM-positive hepatic cancer stem cells. Hepatology 50: 472–480.
Kim VN, Han J, Siomi MC . (2009). Biogenesis of small RNAs in animals. Nat Rev Mol Cell Biol 10: 126–139.
Kong W, Yang H, He L, Zhao JJ, Coppola D, Dalton WS et al. (2008). MicroRNA-155 is regulated by the transforming growth factor beta/Smad pathway and contributes to epithelial cell plasticity by targeting RhoA. Mol Cell Biol 28: 6773–6784.
Krutzfeldt J, Rajewsky N, Braich R, Rajeev KG, Tuschl T, Manoharan M et al. (2005). Silencing of microRNAs in vivo with ‘antagomirs’. Nature 438: 685–689.
Lee EJ, Gusev Y, Jiang J, Nuovo GJ, Lerner MR, Frankel WL et al. (2007). Expression profiling identifies microRNA signature in pancreatic cancer. Int J Cancer 120: 1046–1054.
Livak KJ, Schmittgen TD . (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25: 402–408.
Massague J . (2008). TGFbeta in Cancer. Cell 134: 215–230.
Menghini R, Menini S, Amoruso R, Fiorentino L, Casagrande V, Marzano V et al. (2009). Tissue inhibitor of metalloproteinase 3 deficiency causes hepatic steatosis and adipose tissue inflammation in mice. Gastroenterology 136: 663–672 e4.
Miller TE, Ghoshal K, Ramaswamy B, Roy S, Datta J, Shapiro CL et al. (2008). MicroRNA-221/222 confers tamoxifen resistance in breast cancer by targeting p27Kip1. J Biol Chem 283: 29897–29903.
Mohammed FF, Smookler DS, Taylor SE, Fingleton B, Kassiri Z, Sanchez OH et al. (2004). Abnormal TNF activity in Timp3−/− mice leads to chronic hepatic inflammation and failure of liver regeneration. Nat Genet 36: 969–977.
Nakajima G, Hayashi K, Xi Y, Kudo K, Uchida K, Takasaki K et al. (2006). Non-coding microRNAs hsa-let-7g and hsa-miR-181b are associated with chemoresponse to S-1 in colon cancer. Cancer Genomics Proteomics 3: 317–324.
Nasser MW, Datta J, Nuovo G, Kutay H, Motiwala T, Majumder S et al. (2008). Down-regulation of micro-RNA-1 (miR-1) in lung cancer Suppression of tumorigenic property of lung cancer cells and their sensitization to doxorubicin-induced apoptosis by miR-1. J Biol Chem 283: ##33405.
Schickel R, Boyerinas B, Park SM, Peter ME . (2008). MicroRNAs: key players in the immune system, differentiation, tumorigenesis and cell death. Oncogene 27: 5959–5974.
Selaru FM, Olaru AV, Kan T, David S, Cheng Y, Mori Y et al. (2009). MicroRNA-21 is overexpressed in human cholangiocarcinoma and regulates programmed cell death 4 and tissue inhibitor of metalloproteinase 3. Hepatology 49: 1595–1601.
Shi L, Cheng Z, Zhang J, Li R, Zhao P, Fu Z et al. (2008). hsa-mir-181a and hsa-mir-181b function as tumor suppressors in human glioma cells. Brain Res 1236: 185–193.
Soifer HS, Rossi JJ, Saetrom P . (2007). MicroRNAs in disease and potential therapeutic applications. Mol Ther 15: 2070–2079.
Sun Q, Zhang Y, Yang G, Chen X, Cao G, Wang J et al. (2008). Transforming growth factor-beta-regulated miR-24 promotes skeletal muscle differentiation. Nucleic Acids Res 36: 2690–2699.
Thomas M . (2009). Molecular targeted therapy for hepatocellular carcinoma. J Gastroenterol 44 (Suppl 19): 136–141.
Visone R, Croce CM . (2009). MiRNAs and cancer. Am J Pathol 174: 1131–1138.
Wang B, Majumder S, Nuovo G, Kutay H, Volinia S, Patel T et al. (2009). Role of miR-155 at early stages of hepatocarcinogenesis induced by choline-deficient and amino acid defined diet in C57BL/6 mice. Hepatology 50: 1152–1161 doi 10 1002/hep.23100.
Winter J, Jung S, Keller S, Gregory RI, Diederichs S . (2009). Many roads to maturity: microRNA biogenesis pathways and their regulation. Nat Cell Biol 11: 228–234.
Yan LX, Huang XF, Shao Q, Huang MY, Deng L, Wu QL et al. (2008). MicroRNA miR-21 overexpression in human breast cancer is associated with advanced clinical stage, lymph node metastasis and patient poor prognosis. RNA 14: 2348–2360.
Acknowledgements
We thank Drs David P Bartel and John Taylor for pIS0 vectors and Huh-7 cells, respectively. This work was supported by the grants CA086978 and CA101956 from National Institutes of Health. Grant support: CA086978 and CA101956 from National Institutes of Health.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc)
Rights and permissions
About this article
Cite this article
Wang, B., Hsu, SH., Majumder, S. et al. TGFβ-mediated upregulation of hepatic miR-181b promotes hepatocarcinogenesis by targeting TIMP3. Oncogene 29, 1787–1797 (2010). https://doi.org/10.1038/onc.2009.468
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2009.468
Keywords
This article is cited by
-
Assessing the Potential Apoptotic Effects of Different Hydatid Cyst Fluids on Human Healthy Hepatocytes and Hepatocellular Carcinoma Cells
Acta Parasitologica (2024)
-
Pinostrobin, a fingerroot compound, regulates miR-181b-5p and induces acute leukemic cell apoptosis
Scientific Reports (2023)
-
MicroRNA and their implications in dental pulp inflammation: current trends and future perspectives
Odontology (2023)
-
Formyl peptide receptor 2 determines sex-specific differences in the progression of nonalcoholic fatty liver disease and steatohepatitis
Nature Communications (2022)
-
Mouse models of nonalcoholic steatohepatitis and their application to new drug development
Archives of Pharmacal Research (2022)