Abstract
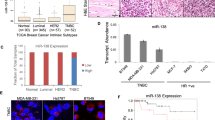
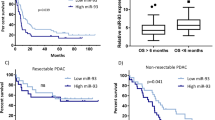
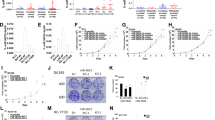
Downregulation of miR-145 in a variety of cancers suggests a possible tumor suppressor function for this microRNA. Here, we show that miR-145 expression is reduced in bladder cancer and urothelial carcinoma in situ, compared with normal urothelium, using transcription profiling and in situ hybridization. Ectopic expression of miR-145 induced extensive apoptosis in urothelial carcinoma cell lines (T24 and SW780) as characterized by caspase activation, nuclear condensation and fragmentation, cellular shrinkage, and detachment. However, cell death also proceeded upon caspase inhibition by the pharmacological inhibitor zVAD-fmk and ectopic expression of anti-apoptotic Bcl-2, indicating the activation of an alternative caspase-independent death pathway. Microarray analysis of transcript levels in T24 cells, before the onset of cell death, showed destabilization of mRNAs enriched for miR-145 7mer target sites. Among these, direct targeting of CBFB, PPP3CA, and CLINT1 was confirmed by a luciferase reporter assay. Notably, a 22-gene signature targeted on enforced miR-145 expression in T24 cells was significantly (P<0.00003) upregulated in 55 Ta bladder tumors with concomitant reduction of miR-145. Our data indicate that reduction in miR-145 expression may provide bladder cancer cells with a selective advantage by inhibition of cell death otherwise triggered in malignant cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Baek D, Villen J, Shin C, Camargo FD, Gygi SP, Bartel DP . (2008). The impact of microRNAs on protein output. Nature 455: 64–71.
Calin GA, Croce CM . (2006). MicroRNA signatures in human cancers. Nat Rev Cancer 6: 857–866.
Calin GA, Sevignani C, Dumitru CD, Hyslop T, Noch E, Yendamuri S et al. (2004). Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci USA 101: 2999–3004.
Cordes KR, Sheehy NT, White MP, Berry EC, Morton SU, Muth AN et al. (2009). miR-145 and miR-143 regulate smooth muscle cell fate and plasticity. Nature 460: 705–710.
Dyrskjot L, Ostenfeld MS, Bramsen JB, Silahtaroglu AN, Lamy P, Ramanathan R et al. (2009). Genomic profiling of microRNAs in bladder cancer: miR-129 is associated with poor outcome and promotes cell death in vitro. Cancer Res 69: 4851–4860.
Dyrskjot L, Zieger K, Kruhoffer M, Thykjaer T, Jensen JL, Primdahl H et al. (2005). A molecular signature in superficial bladder carcinoma predicts clinical outcome. Clin Cancer Res 11: 4029–4036.
Esquela-Kerscher A, Slack FJ . (2006). Oncomirs—microRNAs with a role in cancer. Nat Rev Cancer 6: 259–269.
Hanahan D, Weinberg RA . (2000). The hallmarks of cancer. Cell 100: 57–70.
Hart SM, Foroni L . (2002). Core binding factor genes and human leukemia. Haematologica 87: 1307–1323.
Horrigan SK, Westbrook CA, Kim AH, Banerjee M, Stock W, Larson RA . (1996). Polymerase chain reaction-based diagnosis of del (5q) in acute myeloid leukemia and myelodysplastic syndrome identifies a minimal deletion interval. Blood 88: 2665–2670.
Iorio MV, Ferracin M, Liu CG, Veronese A, Spizzo R, Sabbioni S et al. (2005). MicroRNA gene expression deregulation in human breast cancer. Cancer Res 65: 7065–7070.
Kagoshima H, Nimmo R, Saad N, Tanaka J, Miwa Y, Mitani S et al. (2007). The C. elegans CBFbeta homologue BRO-1 interacts with the Runx factor, RNT-1, to promote stem cell proliferation and self-renewal. Development 134: 3905–3915.
Kedde M, Strasser MJ, Boldajipour B, Oude Vrielink JA, Slanchev K, le Sage C et al. (2007). RNA-binding protein Dnd1 inhibits microRNA access to target mRNA. Cell 131: 1273–1286.
Kim M, Park SY, Pai HS, Kim TH, Billiar TR, Seol DW . (2004). Hypoxia inhibits tumor necrosis factor-related apoptosis-inducing ligand-induced apoptosis by blocking Bax translocation. Cancer Res 64: 4078–4081.
Kim R, Emi M, Tanabe K, Murakami S, Uchida Y, Arihiro K . (2006). Regulation and interplay of apoptotic and non-apoptotic cell death. J Pathol 208: 319–326.
Kroemer G, Galluzzi L, Vandenabeele P, Abrams J, Alnemri ES, Baehrecke EH et al. (2008). Classification of cell death: recommendations of the Nomenclature Committee on Cell Death 2009. Cell Death Differ 16: 3–11.
Lee Y, Ahn C, Han J, Choi H, Kim J, Yim J et al. (2003). The nuclear RNase III Drosha initiates microRNA processing. Nature 425: 415–419.
Levine B, Yuan J . (2005). Autophagy in cell death: an innocent convict? J Clin Invest 115: 2679–2688.
Litynska A, Przybylo M, Ksiazek D, Laidler P . (2000). Differences of alpha3beta1 integrin glycans from different human bladder cell lines. Acta Biochim Pol 47: 427–434.
Michael MZ, SM OC, van Holst Pellekaan NG, Young GP, James RJ . (2003). Reduced accumulation of specific microRNAs in colorectal neoplasia. Mol Cancer Res 1: 882–891.
Mitra AP, Birkhahn M, Cote RJ . (2007). p53 and retinoblastoma pathways in bladder cancer. World J Urol 25: 563–571.
Nakajima G, Hayashi K, Xi Y, Kudo K, Uchida K, Takasaki K et al. (2006). Non-coding MicroRNAs hsa-let-7g and hsa-miR-181b are associated with chemoresponse to S-1 in colon cancer. Cancer Genomics Proteomics 3: 317–324.
Nuovo GJ . (2008). In situ detection of precursor and mature microRNAs in paraffin embedded, formalin fixed tissues and cell preparations. Methods 44: 39–46.
Orom UA, Nielsen FC, Lund AH . (2008). MicroRNA-10a binds the 5’UTR of ribosomal protein mRNAs and enhances their translation. Mol Cell 30: 460–471.
Ostenfeld MS, Fehrenbacher N, Hoyer-Hansen M, Thomsen C, Farkas T, Jaattela M . (2005). Effective tumor cell death by sigma-2 receptor ligand siramesine involves lysosomal leakage and oxidative stress. Cancer Res 65: 8975–8983.
Ow YL, Green DR, Hao Z, Mak TW . (2008). Cytochrome c: functions beyond respiration. Nat Rev Mol Cell Biol 9: 532–542.
Porkka KP, Pfeiffer MJ, Waltering KK, Vessella RL, Tammela TL, Visakorpi T . (2007). MicroRNA expression profiling in prostate cancer. Cancer Res 67: 6130–6135.
Rajewsky N . (2006). microRNA target predictions in animals. Nat Genet 38 (Suppl): S8–S13.
Sachdeva M, Zhu S, Wu F, Wu H, Walia V, Kumar S et al. (2009). p53 represses c-Myc through induction of the tumor suppressor miR-145. Proc Natl Acad Sci USA 106: 3207–3212.
Schepeler T, Reinert JT, Ostenfeld MS, Christensen LL, Silahtaroglu AN, Dyrskjot L et al. (2008). Diagnostic and prognostic microRNAs in stage II colon cancer. Cancer Res 68: 6416–6424.
Selbach M, Schwanhausser B, Thierfelder N, Fang Z, Khanin R, Rajewsky N . (2008). Widespread changes in protein synthesis induced by microRNAs. Nature 455: 58–63.
Sempere LF, Christensen M, Silahtaroglu A, Bak M, Heath CV, Schwartz G et al. (2007). Altered MicroRNA expression confined to specific epithelial cell subpopulations in breast cancer. Cancer Res 67: 11612–11620.
Shi B, Sepp-Lorenzino L, Prisco M, Linsley P, deAngelis T, Baserga R . (2007). Micro RNA 145 targets the insulin receptor substrate-1 and inhibits the growth of colon cancer cells. J Biol Chem 282: 32582–32590.
Tavazoie SF, Alarcon C, Oskarsson T, Padua D, Wang Q, Bos PD et al. (2008). Endogenous human microRNAs that suppress breast cancer metastasis. Nature 451: 147–152.
Thorsen K, Sorensen KD, Brems-Eskildsen AS, Modin C, Gaustadnes M, Hein AM et al. (2008). Alternative splicing in colon, bladder, and prostate cancer identified by exon array analysis. Mol Cell Proteomics 7: 1214–1224.
Vasudevan S, Tong Y, Steitz JA . (2007). Switching from repression to activation: microRNAs can up-regulate translation. Science 318: 1931–1934.
Wang X, Tang S, Le SY, Lu R, Rader JS, Meyers C et al. (2008). Aberrant expression of oncogenic and tumor-suppressive microRNAs in cervical cancer is required for cancer cell growth. PLoS ONE 3: e2557.
Yanaihara N, Caplen N, Bowman E, Seike M, Kumamoto K, Yi M et al. (2006). Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell 9: 189–198.
Yang H, Kong W, He L, Zhao JJ, O’Donnell JD, Wang J et al. (2008). MicroRNA expression profiling in human ovarian cancer: miR-214 induces cell survival and cisplatin resistance by targeting PTEN. Cancer Res 68: 425–433.
Yu L, Alva A, Su H, Dutt P, Freundt E, Welsh S et al. (2004). Regulation of an ATG7-beclin 1 program of autophagic cell death by caspase-8. Science 304: 1500–1502.
Acknowledgements
We thank Gitte Høj, Pamela Celis, Hanne Steen, Inge-Lis Thorsen, Gitte Stougård, and Conni Sørensen for technical assistance. We are grateful to M Jäättelä for providing the pCEP4 Bcl-2 vector construct and to Thomas B Hansen for methylation analysis software. We thank the staff at the Departments of Urology, Clinical Biochemistry, and Pathology at Aarhus University Hospital. This work was supported by the Ministry of Technology and Science, The John and Birthe Meyer Foundation, the Lundbeck Foundation, and the Danish Cancer Society, and the European Community's Seventh framework program (FP7/2007–2013) under grant agreement no 201663.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc)
Rights and permissions
About this article
Cite this article
Ostenfeld, M., Bramsen, J., Lamy, P. et al. miR-145 induces caspase-dependent and -independent cell death in urothelial cancer cell lines with targeting of an expression signature present in Ta bladder tumors. Oncogene 29, 1073–1084 (2010). https://doi.org/10.1038/onc.2009.395
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2009.395
Keywords
This article is cited by
-
Expression of miR-145 and miR-18b in Peripheral Blood Samples of Head and Neck Cancer Patients
Indian Journal of Clinical Biochemistry (2023)
-
MicroRNAs and target molecules in bladder cancer
Medical Oncology (2020)
-
A Genome-Wide Analysis of the Penumbral Volume in Inbred Mice following Middle Cerebral Artery Occlusion
Scientific Reports (2019)
-
Whole‐exome sequencing and microRNA profiling reveal PI3K/AKT pathway’s involvement in juvenile myelomonocytic leukemia
Quantitative Biology (2018)
-
Profiling of long non-coding RNAs identifies LINC00958 and LINC01296 as candidate oncogenes in bladder cancer
Scientific Reports (2017)