Abstract
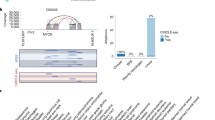
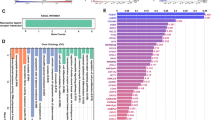
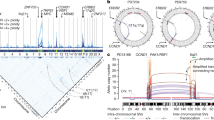
Co-amplification at chromosomes 8p11–8p12 and 11q12–11q14 occurs often in breast tumors, suggesting possible cooperation between genes in these regions in oncogenesis. We used high-resolution array comparative genomic hybridization (array CGH) to map the minimal amplified regions. The 8p and 11q amplicons are complex and consist of at least four amplicon cores at each site. Candidate oncogenes mapping to these regions were identified by combining copy number and RNA and protein expression analyses. These studies also suggested that CCND1 at 11q13 induced expression of ZNF703 mapping at 8p12, which was subsequently shown to be mediated by the Rb/E2F pathway. Nine candidate oncogenes from 8p12 and four from 11q13 were further evaluated for oncogenic function. None of the genes individually promoted colony formation in soft agar or collaborated with each other functionally. On the other hand, FGFR1 and DDHD2 at 8p12 cooperated functionally with MYC, whereas CCND1 and ZNF703 cooperated with a dominant negative form of TP53. These observations highlight the complexity and functional consequences of the genomic rearrangements that occur in these breast cancer amplicons, including transcriptional cross-talk between genes in the 8p and 11q amplicons, as well as their cooperation with major pathways of tumorigenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Adelaide J, Finetti P, Bekhouche I, Repellini L, Geneix J, Sircoulomb F et al. (2007). Integrated profiling of basal and luminal breast cancers. Cancer Res 67: 11565–11575.
Agrawal A, Yang J, Murphy RF, Agrawal DK . (2006). Regulation of the p14ARF-Mdm2-p53 pathway: an overview in breast cancer. Exp Mol Pathol 81: 115–122.
Bautista S, Theillet C . (1998). CCND1 and FGFR1 coamplification results in the colocalization of 11q13 and 8p12 sequences in breast tumor nuclei. Genes Chromosomes Cancer 22: 268–277.
Bekri S, Adelaide J, Merscher S, Grosgeorge J, Caroli-Bosc F, Perucca-Lostanlen D et al. (1997). Detailed map of a region commonly amplified at 11q13→q14 in human breast carcinoma. Cytogenet Cell Genet 79: 125–131.
Bernard-Pierrot I, Gruel N, Stransky N, Vincent-Salomon A, Reyal F, Raynal V et al. (2008). Characterization of the recurrent 8p11–12 amplicon identifies PPAPDC1B, a phosphatase protein, as a new therapeutic target in breast cancer. Cancer Res 68: 7165–7175.
Beroukhim R, Getz G, Nghiemphu L, Barretina J, Hsueh T, Linhart D et al. (2007). Assessing the significance of chromosomal aberrations in cancer: methodology and application to glioma. Proc Natl Acad Sci USA 104: 20007–20012.
Chin K, DeVries S, Fridlyand J, Spellman PT, Roydasgupta R, Kuo WL et al. (2006). Genomic and transcriptional aberrations linked to breast cancer pathophysiologies. Cancer Cell 10: 529–541.
Climent J, Dimitrow P, Fridlyand J, Palacios J, Siebert R, Albertson DG et al. (2007). Deletion of chromosome 11q predicts response to anthracycline-based chemotherapy in early breast cancer. Cancer Res 67: 818–826.
Cuny M, Kramar A, Courjal F, Johannsdottir V, Iacopetta B, Fontaine H et al. (2000). Relating genotype and phenotype in breast cancer: an analysis of the prognostic significance of amplification at eight different genes or loci and of p53 mutations. Cancer Res 60: 1077–1083.
Ethier SP . (1996). Human breast cancer cell lines as models of growth regulation and disease progression. J Mammary Gland Biol Neoplasia 1: 111–121.
Ethier SP, Mahacek ML, Gullick WJ, Frank TS, Weber BL . (1993). Differential isolation of normal luminal mammary epithelial cells and breast cancer cells from primary and metastatic sites using selective media. Cancer Res 53: 627–635.
Forozan F, Karhu R, Kononen J, Kallioniemi A, Kallioniemi OP . (1997). Genome screening by comparative genomic hybridization. Trends Genet 13: 405–409.
Fridlyand J, Snijders AM, Ylstra B, Li H, Olshen A, Segraves R et al. (2006). Breast tumor copy number aberration phenotypes and genomic instability. BMC Cancer 6: 96.
Garcia MJ, Pole JC, Chin SF, Teschendorff A, Naderi A, Ozdag H et al. (2005). A 1 Mb minimal amplicon at 8p11–12 in breast cancer identifies new candidate oncogenes. Oncogene 24: 5235–5245.
Gelsi-Boyer V, Orsetti B, Cervera N, Finetti P, Sircoulomb F, Rouge C et al. (2005). Comprehensive profiling of 8p11–12 amplification in breast cancer. Mol Cancer Res 3: 655–667.
Haverty PM, Fridlyand J, Li L, Getz G, Beroukhim R, Lohr S et al. (2008). High-resolution genomic and expression analyses of copy number alterations in breast tumors. Genes Chromosomes Cancer 47: 530–542.
Ho GH, Calvano JE, Bisogna M, Abouezzi Z, Borgen PI, Cordon-Cardo C et al. (2001). Genetic alterations of the p14ARF-hdm2-p53 regulatory pathway in breast carcinoma. Breast Cancer Res Treat 65: 225–232.
Hochberg YBY . (1995). Controlling the false discovery rate: a practical and powerful approach to multiple testing. Journal of the Royal Statistical Society Series B 57: 289–300.
Janssen JW, Cuny M, Orsetti B, Rodriguez C, Valles H, Bartram CR et al. (2002). MYEOV: a candidate gene for DNA amplification events occurring centromeric to CCND1 in breast cancer. Int J Cancer 102: 608–614.
Kan CE, Patton JT, Stark GR, Jackson MW . (2007). p53-mediated growth suppression in response to Nutlin-3 in cyclin D1 transformed cells occurs independently of p21. Cancer Res 67: 9862–9868.
Karlseder J, Zeillinger R, Schneeberger C, Czerwenka K, Speiser P, Kubista E et al. (1994). Patterns of DNA amplification at band q13 of chromosome 11 in human breast cancer. Genes Chromosomes Cancer 9: 42–48.
Letessier A, Sircoulomb F, Ginestier C, Cervera N, Monville F, Gelsi-Boyer V et al. (2006). Frequency, prognostic impact, and subtype association of 8p12, 8q24, 11q13, 12p13, 17q12, and 20q13 amplifications in breast cancers. BMC Cancer 6: 245.
Loo LW, Grove DI, Williams EM, Neal CL, Cousens LA, Schubert EL et al. (2004). Array comparative genomic hybridization analysis of genomic alterations in breast cancer subtypes. Cancer Res 64: 8541–8549.
Melchor L, Garcia MJ, Honrado E, Pole JC, Alvarez S, Edwards PA et al. (2007). Genomic analysis of the 8p11–12 amplicon in familial breast cancer. Int J Cancer 120: 714–717.
Mineta H, Borg A, Dictor M, Wahlberg P, Wennerberg J . (1997). Correlation between p53 mutation and cyclin D1 amplification in head and neck squamous cell carcinoma. Oral Oncol 33: 42–46.
Neve RM, Chin K, Fridlyand J, Yeh J, Baehner FL, Fevr T et al. (2006). A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 10: 515–527.
Ossovskaya VS, Mazo IA, Chernov MV, Chernova OB, Strezoska Z, Kondratov R et al. (1996). Use of genetic suppressor elements to dissect distinct biological effects of separate p53 domains. Proc Natl Acad Sci USA 93: 10309–10314.
Paterson AL, Pole JC, Blood KA, Garcia MJ, Cooke SL, Teschendorff AE et al. (2007). Co-amplification of 8p12 and 11q13 in breast cancers is not the result of a single genomic event. Genes Chromosomes Cancer 46: 427–439.
Pole JC, Courtay-Cahen C, Garcia MJ, Blood KA, Cooke SL, Alsop AE et al. (2006). High-resolution analysis of chromosome rearrangements on 8p in breast, colon and pancreatic cancer reveals a complex pattern of loss, gain and translocation. Oncogene 25: 5693–5706.
Prentice LM, Shadeo A, Lestou VS, Miller MA, deLeeuw RJ, Makretsov N et al. (2005). NRG1 gene rearrangements in clinical breast cancer: identification of an adjacent novel amplicon associated with poor prognosis. Oncogene 24: 7281–7289.
Ray ME, Yang ZQ, Albertson D, Kleer CG, Washburn JG, Macoska JA et al. (2004). Genomic and expression analysis of the 8p11–12 amplicon in human breast cancer cell lines. Cancer Res 64: 40–47.
Reis-Filho JS, Simpson PT, Turner NC, Lambros MB, Jones C, Mackay A et al. (2006). FGFR1 emerges as a potential therapeutic target for lobular breast carcinomas. Clin Cancer Res 12: 6652–6662.
Rodriguez C, Hughes-Davies L, Valles H, Orsetti B, Cuny M, Ursule L et al. (2004). Amplification of the BRCA2 pathway gene EMSY in sporadic breast cancer is related to negative outcome. Clin Cancer Res 10: 5785–5791.
Rots MG, Pieters R, Peters GJ, Noordhuis P, van Zantwijk CH, Kaspers GJ et al. (1999). Role of folylpolyglutamate synthetase and folylpolyglutamate hydrolase in methotrexate accumulation and polyglutamylation in childhood leukemia. Blood 93: 1677–1683.
Smyth GK . (2004). Statistical Applications in Genetics and Molecular Biology, Vol. 3, pp Vol. 3: Iss. 1, Article 3. Available at: http://www.bepress.com/sagmb/vol3/iss1/art3.
Snijders AM, Fridlyand J, Mans DA, Segraves R, Jain AN, Pinkel D et al. (2003). Shaping of tumor and drug-resistant genomes by instability and selection. Oncogene 22: 4370–4379.
Snijders AM, Nowak N, Segraves R, Blackwood S, Brown N, Conroy J et al. (2001). Assembly of microarrays for genome-wide measurement of DNA copy number. Nat Genet 29: 263–264.
Snijders AM, Schmidt BL, Fridlyand J, Dekker N, Pinkel D, Jordan RC et al. (2005). Rare amplicons implicate frequent deregulation of cell fate specification pathways in oral squamous cell carcinoma. Oncogene 24: 4232–4242.
Tashiro E, Maruki H, Minato Y, Doki Y, Weinstein IB, Imoto M . (2003). Overexpression of cyclin D1 contributes to malignancy by up-regulation of fibroblast growth factor receptor 1 via the pRB/E2F pathway. Cancer Res 63: 424–431.
Wang B, Soule HD, Miller FR . (1997). Transforming and oncogenic potential of activated c-Ha-ras in three immortalized human breast epithelial cell lines. Anticancer Res 17: 4387–4394.
Willmarth NE, Albertson DG, Ethier SP . (2004). Chromosomal instability and lack of cyclin E regulation in hCdc4 mutant human breast cancer cells. Breast Cancer Res 6: R531–R539.
Xian W, Schwertfeger KL, Vargo-Gogola T, Rosen JM . (2005). Pleiotropic effects of FGFR1 on cell proliferation, survival, and migration in a 3D mammary epithelial cell model. J Cell Biol 171: 663–673.
Yang ZQ, Albertson D, Ethier SP . (2004). Genomic organization of the 8p11–p12 amplicon in three breast cancer cell lines. Cancer Genet Cytogenet 155: 57–62.
Yang ZQ, Moffa AB, Haddad R, Streicher KL, Ethier SP . (2007). Transforming properties of TC-1 in human breast cancer: interaction with FGFR2 and beta-catenin signaling pathways. Int J Cancer 121: 1265–1273.
Yang ZQ, Streicher KL, Ray ME, Abrams J, Ethier SP . (2006). Multiple interacting oncogenes on the 8p11–p12 amplicon in human breast cancer. Cancer Res 66: 11632–11643.
Acknowledgements
We thank members of the UCSF Helen Diller Family Comprehensive Cancer Center Genome Analysis, Informatics, and Microarray Shared Resources for performing the TaqMan assays and printing the custom 8p11q array. This work was supported by NIH Grants CA90421 and CA101359. Serena S Kwek was the recipient of a DOD BCRP fellowship, Grant nos. BC021074, DAMD17-03-1-0483.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc)
Rights and permissions
About this article
Cite this article
Kwek, S., Roy, R., Zhou, H. et al. Co-amplified genes at 8p12 and 11q13 in breast tumors cooperate with two major pathways in oncogenesis. Oncogene 28, 1892–1903 (2009). https://doi.org/10.1038/onc.2009.34
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2009.34
Keywords
This article is cited by
-
CCND1 Amplification in Breast Cancer -associations With Proliferation, Histopathological Grade, Molecular Subtype and Prognosis
Journal of Mammary Gland Biology and Neoplasia (2022)
-
Genomic landscape of extraordinary responses in metastatic breast cancer
Communications Biology (2021)
-
Circular RNA circRUNX1 promotes papillary thyroid cancer progression and metastasis by sponging MiR-296-3p and regulating DDHD2 expression
Cell Death & Disease (2021)
-
Eukaryotic initiation factor 4E-binding protein as an oncogene in breast cancer
BMC Cancer (2019)
-
miR-503 represses human cell proliferation and directly targets the oncogene DDHD2 by non-canonical target pairing
BMC Genomics (2015)