Abstract
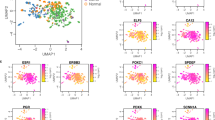
Invasive ductal carcinomas (IDCs) and invasive lobular carcinomas (ILCs) are the two major pathological types of breast cancer. Epidemiological and histoclinical data suggest biological differences, but little is known about the molecular alterations involved in ILCs. We undertook a comparative large-scale study by both array-compared genomic hybridization and cDNA microarray of a set of 50 breast tumors (21 classic ILCs and 29 IDCs) selected on homogeneous histoclinical criteria. Results were validated on independent tumor sets, as well as by quantitative RT–PCR. ILCs and IDCs presented differences at both the genomic and expression levels with ILCs being less rearranged and heterogeneous than IDCs. Supervised analysis defined a 75-BACs signature discriminating accurately ILCs from IDCs. Expression profiles identified two subgroups of ILCs: typical ILCs (∼50%), which were homogeneous and displayed a normal-like molecular pattern, and atypical ILCs, more heterogeneous with features intermediate between ILCs and IDCs. Supervised analysis identified a 75-gene expression signature that discriminated ILCs from IDCs, with many genes involved in cell adhesion, motility, apoptosis, protein folding, extracellular matrix and protein phosphorylation. Although ILCs and IDCs share common alterations, our data show that ILCs and IDCs could be distinguished on the basis of their genomic and expression profiles suggesting that they evolve along distinct genetic pathways.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bertucci F, Borie N, Ginestier C, Groulet A, Charafe-Jauffret E, Adelaide J et al. (2004). Identification and validation of an ERBB2 gene expression signature in breast cancers. Oncogene 23: 2564–2575.
Bertucci F, Finetti P, Cervera N, Maraninchi D, Viens P, Birnbaum D . (2006). Gene expression profiling and clinical outcome in breast cancer. OMICS 10: 429–443.
Berx G, Cleton-Jansen AM, Nollet F, de Leeuw WJ, van de Vijver M, Cornelisse C et al. (1995). E-cadherin is a tumour/invasion suppressor gene mutated in human lobular breast cancers. EMBO J 14: 6107–6115.
Eisen MB, Spellman PT, Brown PO, Botstein D . (1998). Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 95: 14863–14868.
Flagiello D, Gerbault-Seureau M, Sastre-Garau X, Padoy E, Vielh P, Dutrillaux B . (1998). Highly recurrent der(1;16)(q10;p10) and other 16q arm alterations in lobular breast cancer. Genes Chromosomes Cancer 23: 300–306.
Fridlyand J, Snijders AM, Ylstra B, Li H, Olshen A, Segraves R et al. (2006). Breast tumor copy number aberration phenotypes and genomic instability. BMC Cancer 6: 96.
Gunther K, Merkelbach-Bruse S, Amo-Takyi BK, Handt S, Schroder W, Tietze L . (2001). Differences in genetic alterations between primary lobular and ductal breast cancers detected by comparative genomic hybridization. J Pathol 193: 40–47.
Hicks J, Krasnitz A, Lakshmi B, Navin NE, Riggs M, Leibu E et al. (2006). Novel patterns of genome rearrangement and their association with survival in breast cancer. Genome Res 16: 1465–1479.
Jacquemier J, Ginestier C, Rougemont J, Bardou VJ, Charafe-Jauffret E, Geneix J et al. (2005). Protein expression profiling identifies subclasses of breast cancer and predicts prognosis. Cancer Res 65: 767–779.
Katz A, Saad ED, Porter P, Pusztai L . (2007). Primary systemic chemotherapy of invasive lobular carcinoma of the breast. Lancet Oncol 8: 55–62.
Korkola JE, DeVries S, Fridlyand J, Hwang ES, Estep AL, Chen YY et al. (2003). Differentiation of lobular versus ductal breast carcinomas by expression microarray analysis. Cancer Res 63: 7167–7175.
Lamovec J, Bracko M . (1991). Metastatic pattern of infiltrating lobular carcinoma of the breast: an autopsy study. J Surg Oncol 48: 28–33.
Li CI, Anderson BO, Daling JR, Moe RE . (2003). Trends in incidence rates of invasive lobular and ductal breast carcinoma. JAMA 289: 1421–1424.
Loo LW, Grove DI, Williams EM, Neal CL, Cousens LA, Schubert EL et al. (2004). Array comparative genomic hybridization analysis of genomic alterations in breast cancer subtypes. Cancer Res 64: 8541–8549.
Loveday RL, Greenman J, Simcox DL, Speirs V, Drew PJ, Monson JR et al. (2000). Genetic changes in breast cancer detected by comparative genomic hybridization. Int J Cancer 86: 494–500.
Orsetti B, Nugoli M, Cervera N, Lasorsa L, Chuchana P, Rouge C et al. (2006). Genetic profiling of chromosome 1 in breast cancer: mapping of regions of gains and losses and identification of candidate genes on 1q. Br J Cancer 95: 1439–1447.
Pollack JR, Sorlie T, Perou CM, Rees CA, Jeffrey SS, Lonning PE et al. (2002). Microarray analysis reveals a major direct role of DNA copy number alteration in the transcriptional program of human breast tumors. Proc Natl Acad Sci USA 99: 12963–12968.
Sastre-Garau X, Jouve M, Asselain B, Vincent-Salomon A, Beuzeboc P, Dorval T et al. (1996). Infiltrating lobular carcinoma of the breast. Clinicopathologic analysis of 975 cases with reference to data on conservative therapy and metastatic patterns. Cancer 77: 113–120.
Sorlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnsen H et al. (2001). Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA 98: 10869–10874.
Sorlie T, Tibshirani R, Parker J, Hastie T, Marron JS, Nobel A et al. (2003). Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc Natl Acad Sci USA 100: 8418–8423.
Stange DE, Radlwimmer B, Schubert F, Traub F, Pich A, Toedt G et al. (2006). High-resolution genomic profiling reveals association of chromosomal aberrations on 1q and 16p with histologic and genetic subgroups of invasive breast cancer. Clin Cancer Res 12: 345–352.
Toikkanen S, Pylkkanen L, Joensuu H . (1997). Invasive lobular carcinoma of the breast has better short- and long-term survival than invasive ductal carcinoma. Br J Cancer 76: 1234–1240.
Turashvili G, Bouchal J, Baumforth K, Wei W, Dziechciarkova M, Ehrmann J et al. (2007). Novel markers for differentiation of lobular and ductal invasive breast carcinomas by laser microdissection and microarray analysis. BMC Cancer 7: 55.
Zhao H, Langerod A, Ji Y, Nowels KW, Nesland JM, Tibshirani R et al. (2004). Different gene expression patterns in invasive lobular and ductal carcinomas of the breast. Mol Biol Cell 15: 2523–2536.
Acknowledgements
This study was developed as part of a joint program ‘Développement d’outils de diagnostic moléculaire en Cancérologie: Applications aux cancers du sein' Ministère de l'Enseignement Supérieur, de la Recherche et de la Technologie and Fédération Nationale des Centres de Lutte Contre le Cancer and was supported by funds from INSERM, the Association de Recherche sur le Cancer (ARC), grant 5102, Institut National du Cancer, Cancéropoles PACA and Grand Sud Ouest. The help of the Génopole Montpellier Languedoc-Roussillon is gratefully acknowledged. We thank Pr Dominique Maraninchi and Dr Claude Mawas for setting up this work and Mrs Sophie Tourpin for technical help.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc).
Rights and permissions
About this article
Cite this article
Bertucci, F., Orsetti, B., Nègre, V. et al. Lobular and ductal carcinomas of the breast have distinct genomic and expression profiles. Oncogene 27, 5359–5372 (2008). https://doi.org/10.1038/onc.2008.158
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2008.158
Keywords
This article is cited by
-
HER2-positive invasive lobular carcinoma: a rare breast cancer which may not necessarily require anti-HER2 therapy. A population-based study
Breast Cancer (2023)
-
Classification models for Invasive Ductal Carcinoma Progression, based on gene expression data-trained supervised machine learning
Scientific Reports (2020)
-
Gene expression signature of atypical breast hyperplasia and regulation by SFRP1
Breast Cancer Research (2019)
-
Adjuvant chemotherapy in lobular carcinoma of the breast: a clinicopathological score identifies high-risk patient with survival benefit
Breast Cancer Research and Treatment (2019)
-
Invasive lobular and ductal breast carcinoma differ in immune response, protein translation efficiency and metabolism
Scientific Reports (2018)