Abstract
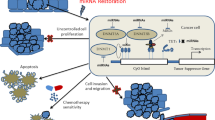
MicroRNAs are small, non-coding RNAs that influence gene regulatory networks by post-transcriptional regulation of specific messenger RNA targets. MicroRNA expression is dysregulated in human malignancies, frequently leading to loss of expression of certain microRNAs. We report that expression of hsa-miR-342, a microRNA encoded in an intron of the gene EVL, is commonly suppressed in human colorectal cancer. The expression of hsa-miR-342 is coordinated with that of EVL and our results indicate that the mechanism of silencing is CpG island methylation upstream of EVL. We found methylation at the EVL/hsa-miR-342 locus in 86% of colorectal adenocarcinomas and in 67% of adenomas, indicating that it is an early event in colorectal carcinogenesis. In addition, we observed a higher frequency of methylation (56%) in histologically normal colorectal mucosa from individuals with concurrent cancer compared to mucosa from individuals without colorectal cancer (12%), suggesting the existence of a ‘field defect’ involving methylated EVL/hsa-miR-342. Furthermore, reconstitution of hsa-miR-342 in the colorectal cancer cell line HT-29 induced apoptosis, suggesting that this microRNA could function as a proapoptotic tumor suppressor. In aggregate, these results support a novel mechanism for silencing intronic microRNAs in cancer by epigenetic alterations of cognate host genes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Alon U, Barkai N, Notterman DA, Gish K, Ybarra S, Mack D et al. (1999). Broad patterns of gene expression revealed by clustering analysis of tumor and normal colon tissues probed by oligonucleotide arrays. Proc Natl Acad Sci USA 96: 6745–6750.
Bandres E, Cubedo E, Agirre X, Malumbres R, Zarate R, Ramirez N et al. (2006). Identification by real-time PCR of 13 mature microRNAs differentially expressed in colorectal cancer and non-tumoral tissues. Mol Cancer 5: 29.
Bartel DP . (2004). MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116: 281–297.
Brueckner B, Stresemann C, Kuner R, Mund C, Musch T, Meister M et al. (2007). The human let-7a-3 locus contains an epigenetically regulated microRNA gene with oncogenic function. Cancer Res 67: 1419–1423.
Cummins JM, He Y, Leary RJ, Pagliarini R, Diaz Jr LA, Sjoblom T et al. (2006). The colorectal microRNAome. Proc Natl Acad Sci USA 103: 3687–3692.
Grady WM, Rajput A, Lutterbaugh J, Markowitz S . (2001). Detection of aberrantly methylated hMLH1 promoter DNA in the serum of patients with microsatellite unstable colon cancer. Cancer Res 61: 900–902.
Griffiths-Jones S, Grocock RJ, van Dongen S, Bateman A, Enright AJ . (2006). miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res 34 (Database issue): D140–D144.
He L, Hannon GJ . (2004). MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet 5: 522–531.
Kim YK, Kim VN . (2007). Processing of intronic microRNAs. EMBO J 26: 775–783.
Kim YH, Petko Z, Dzieciatkowski S, Lin L, Ghiassi M, Stain S et al. (2006). CpG island methylation of genes accumulates during the adenoma progression step of the multistep pathogenesis of colorectal cancer. Genes Chromosomes Cancer 45: 781–789.
Kondo Y, Issa JP . (2004). Epigenetic changes in colorectal cancer. Cancer Metastasis Rev 23: 29–39.
Krause M, Dent EW, Bear JE, Loureiro JJ, Gertler FB . (2003). Ena/VASP proteins: regulators of the actin cytoskeleton and cell migration. Annu Rev Cell Dev Biol 19: 541–564.
Krek A, Grun D, Poy MN, Wolf R, Rosenberg L, Epstein EJ et al. (2005). Combinatorial microRNA target predictions. Nat Genet 37: 495–500.
Lewis BP, Burge CB, Bartel DP . (2005). Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120: 15–20.
Lujambio A, Ropero S, Ballestar E, Fraga MF, Cerrato C, Setien F et al. (2007). Genetic unmasking of an epigenetically silenced microRNA in human cancer cells. Cancer Res 67: 1424–1429.
Michael MZ, O’Connor SM, van Holst Pellekaan NG, Young GP, James RJ . (2003). Reduced accumulation of specific microRNAs in colorectal neoplasia. Mol Cancer Res 1: 882–891.
Myohanen S, Baylin S, Herman J . (1998). Hypermethylation can selectively silence individual p16ink4A alleles in neoplasia. Cancer Res 58: 591–593.
Notterman DA, Alon U, Sierk AJ, Levine AJ . (2001). Transcriptional gene expression profiles of colorectal adenoma, adenocarcinoma, and normal tissue examined by oligonucleotide arrays. Cancer Res 61: 3124–3130.
Ohm JE, Baylin SB . (2007). Stem cell chromatin patterns: an instructive mechanism for DNA hypermethylation? Cell Cycle 6: 1040–1043.
Rhodes DR, Kalyana-Sundaram S, Mahavisno V, Varambally R, Yu J, Briggs BB et al. (2007). Oncomine 3.0: genes, pathways, and networks in a collection of 18 000 cancer gene expression profiles. Neoplasia 9: 166–180.
Rodriguez A, Griffiths-Jones S, Ashurst JL, Bradley A . (2004). Identification of mammalian microRNA host genes and transcription units. Genome Res 14 (10A): 1902–1910.
Saini HK, Griffiths-Jones S, Enright AJ . (2007). Genomic analysis of human microRNA transcripts. Proc Natl Acad Sci USA 104: 17719–17724.
Saito Y, Jones PA . (2006). Epigenetic activation of tumor suppressor microRNAs in human cancer cells. Cell Cycle 5: 2220–2222.
Toyota M, Ho C, Ahuja N, Jair KW, Li Q, Ohe-Toyota M et al. (1999). Identification of differentially methylated sequences in colorectal cancer by methylated CpG island amplification. Cancer Res 59: 2307–2312.
Volinia S, Calin GA, Liu CG, Ambs S, Cimmino A, Petrocca F et al. (2006). A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci USA 103: 2257–2261.
Widschwendter M, Fiegl H, Egle D, Mueller-Holzner E, Spizzo G, Marth C et al. (2007). Epigenetic stem cell signature in cancer. Nat Genet 39: 157–158.
Wijnhoven BP, Michael MZ, Watson DI . (2007). MicroRNAs and cancer. Br J Surg 94: 23–30.
Zhang B, Pan X, Cobb GP, Anderson TA . (2007). MicroRNAs as oncogenes and tumor suppressors. Dev Biol 302: 1–12.
Zou TT, Selaru FM, Xu Y, Shustova V, Yin J, Mori Y et al. (2002). Application of cDNA microarrays to generate a molecular taxonomy capable of distinguishing between colon cancer and normal colon. Oncogene 21: 4855–4862.
Acknowledgements
We thank Dr Beatrice Knudsen for advice regarding tissue processing and Dr Julio Vasquez and Dr Dave McDonald for confocal microscopy. We acknowledge support from the Damon Runyon Lilly Cancer Research Fund (to WMG), the Presidential Early Career Award for Scientists and Engineers (to WMG), the Fred Hutchinson Cancer Research Center (FHCRC) Early Detection Initiative Pilot Project Program (to WMG), the V Foundation Scholar Grant (to MT) and the FHCRC Molecular Diagnostics Pilot Project Program (to MT). These studies were carried out with the use of the experimental histopathology, genomics and scientific imaging shared resources at the FHCRC. The Cooperative Human Tissue Network provided some of the primary tissue samples used in the studies.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc)
Rights and permissions
About this article
Cite this article
Grady, W., Parkin, R., Mitchell, P. et al. Epigenetic silencing of the intronic microRNA hsa-miR-342 and its host gene EVL in colorectal cancer. Oncogene 27, 3880–3888 (2008). https://doi.org/10.1038/onc.2008.10
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2008.10
Keywords
This article is cited by
-
Epigenetic insights in the diagnosis, prognosis, and treatment selection in CRC, an updated review
Molecular Biology Reports (2022)
-
Using BioPAX-Parser (BiP) to enrich lists of genes or proteins with pathway data
BMC Bioinformatics (2021)
-
The balance between the intronic miR-342 and its host gene Evl determines hematopoietic cell fate decision
Leukemia (2021)
-
Epigenetic silencing of miR-342-3p in B cell lymphoma and its impact on autophagy
Clinical Epigenetics (2020)
-
UPF1 regulates the malignant biological behaviors of glioblastoma cells via enhancing the stability of Linc-00313
Cell Death & Disease (2019)