Abstract
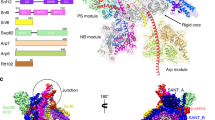
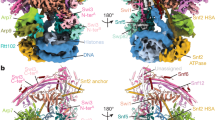
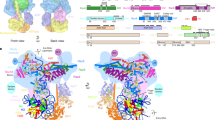
SWI2/SNF2 chromatin-remodeling proteins mediate the mobilization of nucleosomes and other DNA-associated proteins. SWI2/SNF2 proteins contain sequence motifs characteristic of SF2 helicases but do not have helicase activity. Instead, they couple ATP hydrolysis with the generation of superhelical torsion in DNA. The structure of the nucleosome-remodeling domain of zebrafish Rad54, a protein involved in Rad51-mediated homologous recombination, reveals that the core of the SWI2/SNF2 enzymes consist of two α/β-lobes similar to SF2 helicases. The Rad54 helicase lobes contain insertions that form two helical domains, one within each lobe. These insertions contain SWI2/SNF2-specific sequence motifs likely to be central to SWI2/SNF2 function. A broad cleft formed by the two lobes and flanked by the helical insertions contains residues conserved in SWI2/SNF2 proteins and motifs implicated in DNA-binding by SF2 helicases. The Rad54 structure suggests that SWI2/SNF2 proteins use a mechanism analogous to helicases to translocate on dsDNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Flaus, A. & Owen-Hughes, T. Mechanisms for ATP-dependent chromatin remodelling: farewell to the tuna-can octamer? Curr. Opin. Genet. Dev. 14, 165–173 (2004).
Langst, G. & Becker, P.B. Nucleosome remodeling: one mechanism, many phenomena? Biochim. Biophys. Acta 1677, 58–63 (2004).
Flaus, A. & Owen-Hughes, T. Mechanisms for ATP-dependent chromatin remodelling. Curr. Opin. Genet. Dev. 11, 148–154 (2001).
Becker, P.B. & Horz, W. ATP-dependent nucleosome remodeling. Annu. Rev. Biochem. 71, 247–273 (2002).
Davis, J.L., Kunisawa, R. & Thorner, J. A presumptive helicase (MOT1 gene product) affects gene expression and is required for viability in the yeast Saccharomyces cerevisiae. Mol. Cell. Biol. 12, 1879–1892 (1992).
Gorbalenya, A.E. & Koonin, E.V. Helicases: amino acid sequence comparisons and structure-function relationships. Curr. Opin. Struct. Biol. 3, 419–429 (1993).
Singleton, M.R. & Wigley, D.B. Modularity and specialization in superfamily 1 and 2 helicases. J. Bacteriol. 184, 1819–1826 (2002).
Caruthers, J.M. & McKay, D.B. Helicase structure and mechanism. Curr. Opin. Struct. Biol. 12, 123–133 (2002).
Eisen, J.A., Sweder, K.S. & Hanawalt, P.C. Evolution of the SNF2 family of proteins: subfamilies with distinct sequences and functions. Nucleic Acids Res. 23, 2715–2723 (1995).
Richmond, E. & Peterson, C.L. Functional analysis of the DNA-stimulated ATPase domain of yeast SWI2/SNF2. Nucleic Acids Res. 24, 3685–3692 (1996).
Bork, P. & Koonin, E.V. An expanding family of helicases within the 'DEAD/H' superfamily. Nucleic Acids Res. 21, 751–752 (1993).
Cote, J., Quinn, J., Workman, J.L. & Peterson, C.L. Stimulation of GAL4 derivative binding to nucleosomal DNA by the yeast SWI/SNF complex. Science 265, 53–60 (1994).
Havas, K. et al. Generation of superhelical torsion by ATP-dependent chromatin remodeling activities. Cell 103, 1133–1142 (2000).
Krogh, B.O. & Symington, L.S. Recombination proteins in yeast. Annu. Rev. Genet. 38, 233–271 (2004).
Wolner, B., van Komen, S., Sung, P. & Peterson, C.L. Recruitment of the recombinational repair machinery to a DNA double-strand break in yeast. Mol. Cell 12, 221–232 (2003).
Mazin, A.V., Alexeev, A.A. & Kowalczykowski, S.C. A novel function of Rad54 protein. Stabilization of the Rad51 nucleoprotein filament. J. Biol. Chem. 278, 14029–14036 (2003).
Alexiadis, V. & Kadonaga, J.T. Strand pairing by Rad54 and Rad51 is enhanced by chromatin. Genes Dev. 16, 2767–2771 (2002).
Alexeev, A., Mazin, A. & Kowalczykowski, S.C. Rad54 protein possesses chromatin-remodeling activity stimulated by the Rad51-ssDNA nucleoprotein filament. Nat. Struct. Biol. 10, 182–186 (2003).
Jaskelioff, M., Van Komen, S., Krebs, J.E., Sung, P. & Peterson, C.L. Rad54p is a chromatin remodeling enzyme required for heteroduplex DNA joint formation with chromatin. J. Biol. Chem. 278, 9212–9218 (2003).
Solinger, J.A., Kiianitsa, K. & Heyer, W.D. Rad54, a Swi2/Snf2-like recombinational repair protein, disassembles Rad51:dsDNA filaments. Mol. Cell 10, 1175–1188 (2002).
Van Komen, S., Petukhova, G., Sigurdsson, S., Stratton, S. & Sung, P. Superhelicity-driven homologous DNA pairing by yeast recombination factors Rad51 and Rad54. Mol. Cell 6, 563–572 (2000).
Tan, T.L., Kanaar, R. & Wyman, C. Rad54, a Jack of all trades in homologous recombination. DNA Repair (Amst.) 2, 787–794 (2003).
Petukhova, G., Stratton, S. & Sung, P. Catalysis of homologous DNA pairing by yeast Rad51 and Rad54 proteins. Nature 393, 91–94 (1998).
Golub, E.I., Kovalenko, O.V., Gupta, R.C., Ward, D.C. & Radding, C.M. Interaction of human recombination proteins Rad51 and Rad54. Nucleic Acids Res. 25, 4106–4110 (1997).
Raschle, M., Van Komen, S., Chi, P., Ellenberger, T. & Sung, P. Multiple interactions with the Rad51 recombinase govern the homologous recombination function of Rad54. J. Biol. Chem. 279, 51973–51980 (2004).
Alexiadis, V., Lusser, A. & Kadonaga, J.T. A conserved N-terminal motif in Rad54 is important for chromatin remodeling and homologous strand pairing. J. Biol. Chem. 279, 27824–27829 (2004).
Holm, L. & Sander, C. Protein structure comparison by alignment of distance matrices. J. Mol. Biol. 233, 123–138 (1993).
Krissinel, E. & Henrick, K. Secondary-structure matching (SSM), a new tool for fast protein structure alignment in three dimensions. Acta Crystallogr. D 60, 2256–2268 (2004).
Singleton, M.R., Scaife, S. & Wigley, D.B. Structural analysis of DNA replication fork reversal by RecG. Cell 107, 79–89 (2001).
Caruthers, J.M., Johnson, E.R. & McKay, D.B. Crystal structure of yeast initiation factor 4A, a DEAD-box RNA helicase. Proc. Natl Acad. Sci. USA 97, 13080–13085 (2000).
Story, R.M., Li, H. & Abelson, J.N. Crystal structure of a DEAD box protein from the hyperthermophile Methanococcus jannaschii. Proc. Natl. Acad. Sci. USA 98, 1465–1470 (2001).
Bernstein, D.A., Zittel, M.C. & Keck, J.L. High-resolution structure of the E. coli RecQ helicase catalytic core. EMBO J. 22, 4910–4921 (2003).
Machius, M., Henry, L., Palnitkar, M. & Deisenhofer, J. Crystal structure of the DNA nucleotide excision repair enzyme UvrB from Thermus thermophilus. Proc. Natl. Acad. Sci. USA 96, 11717–11722 (1999).
Kim, J.L. et al. Hepatitis C virus NS3 RNA helicase domain with a bound oligonucleotide: the crystal structure provides insights into the mode of unwinding. Structure 6, 89–100 (1998).
Bateman, A. et al. The Pfam protein families database. Nucleic Acids Res. 30, 276–280 (2002).
Velankar, S.S., Soultanas, P., Dillingham, M.S., Subramanya, H.S. & Wigley, D.B. Crystal structures of complexes of PcrA DNA helicase with a DNA substrate indicate an inchworm mechanism. Cell 97, 75–84 (1999).
Tuteja, N. & Tuteja, R. Unraveling DNA helicases. Motif, structure, mechanism and function. Eur. J. Biochem. 271, 1849–1863 (2004).
Boerkoel, C.F. et al. Mutant chromatin remodeling protein SMARCAL1 causes Schimke immuno-osseous dysplasia. Nat. Genet. 30, 215–220 (2002).
Hiramoto, T. et al. Mutations of a novel human RAD54 homologue, RAD54B, in primary cancer. Oncogene 18, 3422–3426 (1999).
Smirnova, M., Van Komen, S., Sung, P. & Klein, H.L. Effects of tumor-associated mutations on Rad54 functions. J. Biol. Chem. 279, 24081–24088 (2004).
Matsuda, M. et al. Mutations in the RAD54 recombination gene in primary cancers. Oncogene 18, 3427–3430 (1999).
Wada, T., Kubota, T., Fukushima, Y. & Saitoh, S. Molecular genetic study of Japanese patients with X-linked alpha-thalassemia/mental retardation syndrome (ATR-X). Am. J. Med. Genet. 94, 242–248 (2000).
Ristic, D., Wyman, C., Paulusma, C. & Kanaar, R. The architecture of the human Rad54-DNA complex provides evidence for protein translocation along DNA. Proc. Natl. Acad. Sci. USA 98, 8454–8460 (2001).
Smith, C.L., Horowitz-Scherer, R., Flanagan, J.F., Woodcock, C.L. & Peterson, C.L. Structural analysis of the yeast SWI/SNF chromatin remodeling complex. Nat. Struct. Biol. 10, 141–145 (2003).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Schneider, T.R. & Sheldrick, G.M. Substructure solution with SHELXD. Acta Crystallogr. D 58, 1772–1779 (2002).
Bricogne, G., Vonrhein, C., Flensburg, C., Schiltz, M. & Paciorek, W. Generation, representation and flow of phase information in structure determination: recent developments in and around SHARP 2.0. Acta Crystallogr. D 59, 2023–2030 (2003).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Acknowledgements
The authors thank C. Ogata at APS 8BM, and A. Mulichak and L. Keefe at APS IMCA-CAT ID17, for help during data collection; and M. Lu of the Weill Cornell Medical Center (New York) for help with the analytical ultracentrifugation experiments. N.H.T would like to acknowledge the Human Frontiers Science Program for a long-term fellowship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
The zebrafish Rad54 crystallization construct is active in supercoiling assays. (PDF 501 kb)
Supplementary Fig. 2
Alignment of Rad54 orthologs. (PDF 137 kb)
Supplementary Fig. 3
Section of the electron density map of Rad54. (PDF 883 kb)
Supplementary Fig. 4
DNA binding activity of Rad54. (PDF 486 kb)
Supplementary Fig. 5
Analytical ultra-centrifugation indicated that Rad54 is monomeric in solution. (PDF 116 kb)
Supplementary Table 1
Zebrafish Rad54 has DNA dependent ATPase activity comparable to that of the human enzyme. (PDF 96 kb)
Supplementary Table 2
Conservation of helicase motifs. (PDF 115 kb)
Rights and permissions
About this article
Cite this article
Thomä, N., Czyzewski, B., Alexeev, A. et al. Structure of the SWI2/SNF2 chromatin-remodeling domain of eukaryotic Rad54. Nat Struct Mol Biol 12, 350–356 (2005). https://doi.org/10.1038/nsmb919
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb919
This article is cited by
-
The limits of clinical findings in similar phenotypes, from Carpenter to ATRX syndrome using a whole exome sequencing approach: a case review
Human Genomics (2021)
-
Structural basis for the multi-activity factor Rad5 in replication stress tolerance
Nature Communications (2021)
-
The bromodomain containing protein BRD-9 orchestrates RAD51–RAD54 complex formation and regulates homologous recombination-mediated repair
Nature Communications (2020)
-
RAD54 N-terminal domain is a DNA sensor that couples ATP hydrolysis with branch migration of Holliday junctions
Nature Communications (2018)
-
Actin-related proteins regulate the RSC chromatin remodeler by weakening intramolecular interactions of the Sth1 ATPase
Communications Biology (2018)