Abstract
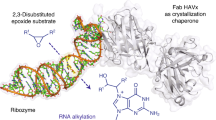
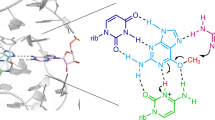
The majority of structural efforts addressing RNA's catalytic function have focused on natural ribozymes, which catalyze phosphodiester transfer reactions. By contrast, little is known about how RNA catalyzes other types of chemical reactions. We report here the crystal structures of a ribozyme that catalyzes enantioselective carbon-carbon bond formation by the Diels-Alder reaction in the unbound state and in complex with a reaction product. The RNA adopts a λ-shaped nested pseudoknot architecture whose preformed hydrophobic pocket is precisely complementary in shape to the reaction product. RNA folding and product binding are dictated by extensive stacking and hydrogen bonding, whereas stereoselection is governed by the shape of the catalytic pocket. Catalysis is apparently achieved by a combination of proximity, complementarity and electronic effects. We observe structural parallels in the independently evolved catalytic pocket architectures for ribozyme- and antibody-catalyzed Diels-Alder carbon-carbon bond-forming reactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Kruger, K. et al. Self-splicing RNA: autoexcision and autocyclization of the ribosomal RNA intervening sequence in Tetrahymena. Cell 31, 145–157 (1982).
Guerrier-Takada, C., Gardiner, K., Marsh, T., Pace, N. & Altman, S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell 35, 849–857 (1983).
Gilbert, W. The RNA world. Nature 319, 618–620 (1986).
Cech, T.R. Ribozymes, the first 20 years. Biochem. Soc. Trans. 30, 1162–1166 (2002).
Doudna, J.A. & Cech, T.R. The chemical repertoire of natural ribozymes. Nature 418, 222–228 (2002).
Steitz, T.A. & Moore, P.B. RNA, the first macromolecular catalyst: the ribosome is a ribozyme. Trends Biochem. Sci. 28, 411–418 (2003).
Wilson, D.S. & Szostak, J.W. In vitro selection of functional nucleic acids. Annu. Rev. Biochem. 68, 611–647 (1999).
Mandal, M. & Breaker, R.R. Gene regulation by riboswitches. Nature Rev. Mol. Cell Biol. 5, 451–463 (2004).
Nudler, E. & Mironov, A.S. The riboswitch control of bacterial metabolism. Trends Biochem. Sci. 29, 11–17 (2004).
Murray, J.M. & Doudna, J.A. Creative catalysis: pieces of the RNA world jigsaw. Trends Biochem. Sci. 26, 699–701 (2001).
Lilley, D.M. Analysis of global conformational transitions in ribozymes. Methods Mol. Biol. 252, 77–108 (2004).
Chapman, K.B. & Szostak, J.W. In vitro selection of catalytic RNAs. Curr. Opin. Struct. Biol. 4, 618–622 (1994).
Famulok, M., Mayer, G. & Blind, M. Nucleic acid aptamers—from selection in vitro to applications in vivo. Acc. Chem. Res. 33, 591–599 (2000).
Jäschke, A. Artificial ribozymes and deoxyribozymes. Curr. Opin. Struct. Biol. 11, 321–326 (2001).
Joyce, G.F. & Orgel, L.E. Prospects for understanding the origin of the RNA world. In The RNA World 2nd edn. (eds. Gesteland, R.F., Cech, T.R. & Atkins, J.F.) 49–77 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 1998).
Seelig, B. & Jäschke, A. A small catalytic RNA motif with Diels-Alderase activity. Chem. Biol. 6, 167–176 (1999).
Tarasow, T.M., Tarasow, S.L. & Eaton, B.E. RNA-catalyzed carbon-carbon bond formation, Nature 389, 54–57 (1997).
Nicolaou, K.C., Snyder, S.A., Montagnon, T. & Vassilikogiannakis, G.E. The Diels-Alder reaction in total synthesis. Angew. Chem. Int. Ed. 41, 1668–1698 (2002).
Seelig, B., Keiper, S., Stuhlmann, F. & Jäschke, A. Enantioselective ribozyme catalysis of a bimolecular cycloaddition reaction. Angew. Chem. Int. Ed. 39, 4576–4579 (2000).
Keiper, S., Bebenroth, D., Seelig, B., Westhof, E. & Jäschke, A. An architecture of a Diels-Alderase ribozyme with a preformed catalytic pocket. Chem. Biol. 11, 1217–1227 (2004).
Stuhlmann, F. & Jäschke, A. Characterization of an RNA active site: interactions between a Diels-Alderase ribozyme and its substrates and products. J. Am. Chem. Soc. 124, 328–344 (2002).
Hermann, T. & Patel, D.J. Adaptive recognition by nucleic acid aptamers. Science 287, 820–825 (2000).
Du, Q. et al. Internal derivatization of oligonucleotides with selenium for X-ray crystallography with MAD. J. Am. Chem. Soc. 124, 2425 (2002).
Höbartner, C. & Micura, R. Chemical synthesis of selenium-modified oligoribonucleotides and their enzymatic ligation leading to an U6 snRNA stem-loop segment. J. Am. Chem. Soc. 126, 1141–1149 (2004).
Teplova, M. et al. Covalent incorporation of selenium into oligonucleotides for X-ray crystal structure determination via MAD: proof of principle. Biochimie 84, 849–858 (2002).
Adams, P.L., Stahley, M.R., Kosek, A.B., Wang, J. & Strobel, S.A. Crystal structure of a self-splicing group I intron with both exons. Nature 430, 45–50 (2004).
Pley, H., Flaherty, K.M. & McKay, D.B. Three-dimensional structure of a hammerhead ribozyme. Nature 372, 68–74 (1994).
Lilley, D.M. The Varkud satellite ribozyme. RNA 10, 151–158 (2004).
Ferre-D'Amare, A.R., Zhou, K. & Doudna, J.A. Crystal structure of a hepatitis delta virus ribozyme. Nature 395, 567–574 (1998).
Schlax, P.J., Xavier, K.A., Gluick, T.C. & Draper, D.E. Translational repression of the Escherichia coli α operon mRNA: importance of an mRNA conformational switch and a ternary entrapment complex. J. Biol. Chem. 276, 38494–38501 (2001).
Majerfeld, I. & Yarus, M. An RNA pocket for an aliphatic hydrophobe. Nat. Struct. Biol. 1, 287–292 (1994).
Williamson, J.R. Induced fit in RNA-protein recognition. Nat. Struct. Biol. 7, 834–837 (2000).
Leulliot, N. & Varani, G. Current topics in RNA-protein recognition: control of specificity and biological function through induced fit and conformational capture. Biochemistry 40, 7947–7956 (2001).
Manoharan, M., De Proft, F. & Geerlings, P. A computational study of aromaticity-controlled Diels-Alder reactions. J. Chem. Soc. Perkin Trans. 2, 1767–1773 (2000).
Wise, K.E. & Wheeler, R.A. Donor-acceptor-assisted Diels-Alder reaction of anthracene and tetracyanoethylene. J. Phys. Chem. 103, 8279–8287 (1999).
Xu, J. et al. Evolution of shape complementarity and catalytic efficiency from a primordial antibody template. Science 286, 23455–2348 (1999).
Chen, J., Deng, Q., Wang, R., Houk, K. & Hilvert, D. Shape complementarity, binding-site dynamics, and transition state stabilization: a theoretical study of Diels-Alder catalysis by antibody 1E9. Chembiochem 1, 255–261 (2000).
Fleming, I. Frontier Orbitals in Organic Chemical Reactions (Wiley, New York, 1976).
Heine, A. et al. An antibody exo Diels-Alderase inhibitor complex at 1.95 Å resolution. Science 279, 1934–1940 (1998).
Romesberg, F.E., Spiller, B., Schultz, P.G. & Stevens, R.C. Immunological origins of binding and catalysis in a Diels-Alderase antibody. Science 279, 1929–1933 (1998).
Hugot, M. et al. A structural basis for the activity of retro-Diels-Alder catalytic antibodies: Evidence for a catalytic aromatic residue. Proc. Natl. Acad. Sci. USA 99, 9674–9678 (2002).
Ose, T. et al. Insights into a natural Diels-Alder reaction from the structure of macrophomate synthase. Nature 422, 185–189 (2003).
Ose, T. et al. Structure of macrophomate synthase. Acta Crystallogr. D 60, 1187–1197 (2004).
Tarasow, T.M. et al. The effect of mutation on RNA Diels-Alderases. J. Am. Chem. Soc. 126, 11843–11851 (2004).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Schneider, T.R. & Sheldrick, G.M. Substructure solution with SHELXD. Acta Crystallogr. D 58, 1772–1779 (2002).
De La Fortelle, E. & Bricogne, G. Maximum-likelihood heavy-atom parameter refinement for multiple isomorphous replacement and multiwavelength anomalous diffraction methods. Methods Enzymol. 276, 472–494 (1997).
Abrahams, J.P. & Leslie, A.G. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D 52, 30–42 (1996).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–245 (1997).
Brunger, A.T. et al. Crystallography and NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Acknowledgements
We gratefully acknowledge support by the US National Institutes of Health, the DeWitt Wallace Foundation and the Abby Rockefeller Mauze Trust (D.J.P.), the Bundesministerium für Bildung und Forschung (BioFuture program), the Deutsche Forschungsgemeinschaft, HFSP and the Fonds der Chemischen Industrie (A.J.), and the Austrian Science Fund FWF (R.M.). We thank V. Kuryavyi for extensive discussions on graphic programs, A. Teplov for help with data collection, M. Becker and the staff of the X25 and X12c beamlines at National Synchrotron Light Source for assistance with data collection, and the personnel of beamlines 14-BM and 19-BM at the Advanced Photon Source for data collection support.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Structural details. (PDF 701 kb)
Supplementary Fig. 2
Folding of the ribozyme monitored by NMR. (PDF 180 kb)
Supplementary Fig. 3
Schematic drawings of RNA pseudoknot topologies. (PDF 104 kb)
Supplementary Fig. 4
Model for the catalytic mechanism. (PDF 615 kb)
Supplementary Fig. 5
Electron density maps. (PDF 552 kb)
Rights and permissions
About this article
Cite this article
Serganov, A., Keiper, S., Malinina, L. et al. Structural basis for Diels-Alder ribozyme-catalyzed carbon-carbon bond formation. Nat Struct Mol Biol 12, 218–224 (2005). https://doi.org/10.1038/nsmb906
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb906
This article is cited by
-
A new RNA performs old chemistry
Nature Chemical Biology (2022)
-
Structure and mechanism of the methyltransferase ribozyme MTR1
Nature Chemical Biology (2022)
-
Structure and mechanism of a methyltransferase ribozyme
Nature Chemical Biology (2022)
-
A Cu(II)–ATP complex efficiently catalyses enantioselective Diels–Alder reactions
Nature Communications (2020)
-
The plasticity of redox cofactors: from metalloenzymes to redox-active DNA
Nature Reviews Chemistry (2018)