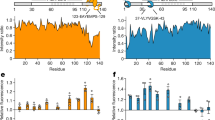
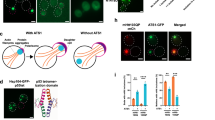
Abstract
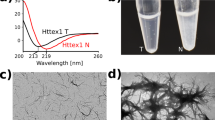
Protein conformational changes that result in misfolding, aggregation and amyloid fibril formation are a common feature of many neurodegenerative disorders. Studies with β-amyloid (Aβ), α-synuclein and other amyloid-forming proteins indicate that the assembly of misfolded protein conformers into fibrils is a complex process that may involve the population of metastable spherical and/or annular oligomeric assemblies. Here, we show by atomic force microscopy that a mutant huntingtin fragment with an expanded polyglutamine repeat forms spherical and annular oligomeric structures reminiscent of those formed by Aβ and α-synuclein. Notably, the molecular chaperones Hsp70 and Hsp40, which are protective in animal models of neurodegeneration, modulate polyglutamine aggregation reactions by partitioning monomeric conformations and disfavoring the accretion of spherical and annular oligomers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Dobson, C.M. Protein folding and misfolding. Nature 426, 884–890 (2003).
Caughey, B. & Lansbury, P.T. Protofibrils, pores, fibrils, and neurodegeneration: separating the responsible protein aggregates from the innocent bystanders. Annu. Rev. Neurosci. 26, 267–298 (2003).
Walsh, D.M., Lomakin, A., Benedek, G.B., Condron, M.M. & Teplow, D.B. Amyloid β-protein fibrillogenesis. Detection of a protofibrillar intermediate. J. Biol. Chem. 272, 22364–22372 (1997).
Harper, J.D., Wong, S.S., Lieber, C.M. & Lansbury, P.T. Observation of metastable Aβ amyloid protofibrils by atomic force microscopy. Chem. Biol. 4, 119–125 (1997).
Lambert, M.P. et al. Diffusible, nonfibrillar ligands derived from Aβ1-42 are potent central nervous system neurotoxins. Proc. Natl. Acad. Sci. USA 95, 6448–6453 (1998).
Conway, K.A., Harper, J.D. & Lansbury, P.T., Jr. Fibrils formed in vitro from αsynuclein and two mutant forms linked to Parkinson's disease are typical amyloid. Biochemistry 39, 2552–2563 (2000).
Conway, K.A. et al. Accelerated oligomerization by Parkinson's disease linked αsynuclein mutants. Ann. NY Acad. Sci. 920, 42–45 (2000).
Rochet, J.C., Conway, K.A. & Lansbury, P.T., Jr. Inhibition of fibrillization and accumulation of prefibrillar oligomers in mixtures of human and mouse α-synuclein. Biochemistry 39, 10619–10626 (2000).
Ding, T.T., Lee, S.J., Rochet, J.C. & Lansbury, P.T. Jr. Annular α-synuclein protofibrils are produced when spherical protofibrils are incubated in solution or bound to brain-derived membranes. Biochemistry 41, 10209–10217 (2002).
The Huntington's Disease Collaborative Research Group. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. Cell 72, 971–983 (1993).
Scherzinger, E. et al. Huntingtin-encoded polyglutamine expansions form amyloidlike protein aggregates in vitro and in vivo. Cell 90, 549–558 (1997).
Hartl, F.U. & Hayer-Hartl, M. Molecular chaperones in the cytosol: from nascent chain to folded protein. Science 295, 1852–1858 (2002).
Muchowski, P.J. Protein misfolding, amyloid formation, and neurodegeneration: a critical role for molecular chaperones? Neuron 35, 9–12 (2002). 24
Muchowski, P.J. et al. Hsp70 and hsp40 chaperones can inhibit self-assembly of polyglutamine proteins into amyloid-like fibrils. Proc. Natl. Acad. Sci. USA 97, 7841–7846 (2000).
Scherzinger, E. et al. Self-assembly of polyglutamine-containing huntingtin fragments into amyloid-like fibrils: implications for Huntington's disease pathology. Proc. Natl. Acad. Sci. USA 96, 4604–4609 (1999).
Fewell, S.W., Day, B.W. & Brodsky, J.L. Identification of an inhibitor of hsc70- mediated protein translocation and ATP hydrolysis. J. Biol. Chem. 276, 910–914 (2001).
Kayed, R. et al. Common structure of soluble amyloid oligomers implies common mechanism of pathogenesis. Science 300, 486–489 (2003).
Chan, H.Y., Warrick, J.M., Gray-Board, G.L., Paulson, H.L. & Bonini, N.M. Mechanisms of chaperone suppression of polyglutamine disease: selectivity, synergy and modulation of protein solubility in Drosophila. Hum. Mol. Genet. 9, 2811–2820 (2000).
Giasson, B.I. et al. Initiation and synergistic fibrillization of τ and α-synuclein. Science 300, 636–640 (2003).
Poirier, M.A. et al. Huntingtin spheroids and protofibrils as precursors in polyglutamine fibrilization. J. Biol. Chem. 277, 41032–41037 (2002).
Tanaka, M. et al. Expansion of polyglutamine induces the formation of quasiaggregate in the early stage of protein fibrillization. J. Biol. Chem. 278, 34717–34724 (2003).
Chen, S., Ferrone, F.A. & Wetzel, R. Huntington's disease age-of-onset linked to polyglutamine aggregation nucleation. Proc. Natl. Acad. Sci. USA 99, 11884–11889 (2002).
Shorter, J. & Lindquist, S. Hsp104 catalyzes formation and elimination of selfreplicating Sup35 prion conformers. Science 304, 1793–1797 (2004).
Perutz, M.F., Johnson, T., Suzuki, M. & Finch, J.T. Glutamine repeats as polar zippers: their possible role in inherited neurodegenerative diseases. Proc. Natl. Acad. Sci. USA 91, 5355–5358 (1994).
Sugars, K.L. & Rubinsztein, D.C. Transcriptional abnormalities in Huntington disease. Trends. Genet. 19, 233–238 (2003).
Schaffar, G. et al. Cellular toxicity of polyglutamine expansion proteins: mechanism of transcription factor deactivation. Mol. Cell 15, 95–105 (2004).
Stockel, J. & Hartl, F.U. Chaperonin-mediated de novo generation of prion protein aggregates. J. Mol. Biol. 313, 861–872 (2001).
Meriin, A.B. et al. Huntington toxicity in yeast model depends on polyglutamine aggregation mediated by a prion-like protein Rnq1. J. Cell Biol. 157, 997–1004 (2002).
Warrick, J.M. et al. Suppression of polyglutamine-mediated neurodegeneration in Drosophila by the molecular chaperone HSP70. Nat. Genet. 23, 425–428 (1999).
Kazemi-Esfarjani, P. & Benzer, S. Genetic suppression of polyglutamine toxicity in Drosophila. Science 287, 1837–1840 (2000).
Cummings, C.J. et al. Over-expression of inducible HSP70 chaperone suppresses neuropathology and improves motor function in SCA1 mice. Hum. Mol. Genet. 10, 1511–1518 (2001).
Auluck, P.K., Chan, H.Y., Trojanowski, J.Q., Lee, V.M. & Bonini, N.M. Chaperone suppression of α-synuclein toxicity in a Drosophila model for Parkinson's disease. Science 295, 865–868 (2002).
Minami, Y., Hohfeld, J., Ohtsuka, K. & Hartl, F.U. Regulation of the heat-shock protein 70 reaction cycle by the mammalian DnaJ homolog, Hsp40. J. Biol. Chem. 271, 19617–19624 (1996).
Ko, J., Ou, S. & Patterson, P.H. New anti-huntingtin monoclonal antibodies: implications for huntingtin conformation and its binding proteins. Brain Res. Bull. 56, 319–329 (2001).
Collins, S.R., Douglass, A., Vale, R.D. & Weissman, J.S. Mechanism of prion propagation: amyloid growth occurs by monomer addition. PLoS Biol. 2, E321 (2004).
Acknowledgements
P.J.M. is supported by the US National Institute of Neurological Disorders and Stroke (R01NS47237), by a US National Institutes of Health (NIH) construction award (C06 RR 14571), by the Alzheimer's Disease Research Center at the University of Washington and by the Hereditary Disease Foundation under the auspices of the Cure Huntington's Disease Initiative. J.L.W. is funded in part by PHS NRSA T32 GM07270 from the US National Institute of General Medical Sciences. H.F., M.H.Z. and M.S. thank DURINT Program for support. AFM experiments were carried out in the Molecular Biomimetics Facilities at the University of Washington. We thank P. Patterson for the MW1-8 antibodies, S. Finkbeiner for the 3B5H10 antibody and M. Mayer and K. Terada for plasmids.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Three-dimensional topography of a HD53Q annular structure. (PDF 131 kb)
Supplementary Fig. 2
Hsp70 or Hsp40 alone does not destabilize HD53Q oligomers. (PDF 919 kb)
Supplementary Fig. 3
Reactivity of MW7 to HD20Q does not change over time or in the presence of Hsp70 and Hsp40. (PDF 483 kb)
Rights and permissions
About this article
Cite this article
Wacker, J., Zareie, M., Fong, H. et al. Hsp70 and Hsp40 attenuate formation of spherical and annular polyglutamine oligomers by partitioning monomer. Nat Struct Mol Biol 11, 1215–1222 (2004). https://doi.org/10.1038/nsmb860
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb860
This article is cited by
-
Amyloid modifier SERF1a interacts with polyQ-expanded huntingtin-exon 1 via helical interactions and exacerbates polyQ-induced toxicity
Communications Biology (2023)
-
Interplay between recombinant Hsp70 and proteasomes: proteasome activity modulation and ubiquitin-independent cleavage of Hsp70
Cell Stress and Chaperones (2017)
-
Suppression of amyloid fibrils using the GroEL apical domain
Scientific Reports (2016)
-
Polyglutamine Aggregation in Huntington Disease: Does Structure Determine Toxicity?
Molecular Neurobiology (2015)
-
Characterization of prion-like conformational changes of the neuronal isoform of Aplysia CPEB
Nature Structural & Molecular Biology (2013)