Abstract
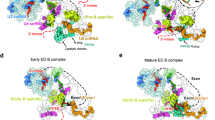
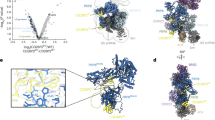
Major structural changes occur in the spliceosome during its transition from the fully assembled complex B to the catalytically activated spliceosome. To understand the rearrangement, it is necessary to know the detailed three-dimensional structures of these complexes. Here, we have immunoaffinity-purified human spliceosomes (designated BΔU1) at a stage after U4/U6·U5 tri-snRNP integration but before activation, and have determined the three-dimensional structure of BΔU1 by single-particle electron cryomicroscopy at a resolution of ∼40 Å. The overall size of the complex is about 370 × 270 × 170 Å. The three-dimensional structure features a roughly triangular body linked to a head domain in variable orientations. The body is very similar in size and shape to the isolated U4/U6·U5 tri-snRNP. This provides initial insight into the structural organization of complex B.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Burge, C., Tuschl, T. & Sharp, P.A. Splicing of precursors to mRNA by the spliceosome. In The RNA World (eds. Cech, T.R. & Atkins, J.F.) 525–560 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 1999).
Staley, J.P. & Guthrie, C. Mechanical devices of the spliceosome: motors, clocks, springs, and things. Cell 92, 315–326 (1998).
Nilsen, T.W. RNA-RNA interactions in the spliceosome: unraveling the ties that bind. Cell 78, 1–4 (1994).
Kambach, C., Walke, S. & Nagai, K. Structure and assembly of the spliceosomal small nuclear ribonucleoprotein particles. Curr. Opin. Struct. Biol. 9, 222–230 (1999).
Behrens, S.E., Tyc, K., Kastner, B., Reichelt, J. & Lührmann, R. Small nuclear ribonucleoprotein (RNP) U2 contains numerous additional proteins and has a bipartite RNP structure under splicing conditions. Mol. Cell. Biol. 13, 307–319 (1993).
Fabrizio, P., Esser, S., Kastner, B. & Lührmann, R. Isolation of S. cerevisiae snRNPs: comparison of U1 and U4/U6·U5 to their human counterparts. Science 264, 261–265 (1994).
Stark, H., Dube, P., Lührmann, R. & Kastner, B. Arrangement of RNA and proteins in the spliceosomal U1 small nuclear ribonucleoprotein particle. Nature 409, 539–542 (2001).
Golas, M.M., Sander, B., Will, C.L., Lührmann, R. & Stark, H. Molecular architecture of the multiprotein splicing factor SF3b. Science 300, 980–984 (2003).
Jurica, M.S. & Moore, M.J. Pre-mRNA splicing: awash in a sea of proteins. Mol. Cell 12, 5–14 (2003).
Nilsen, T.W. The spliceosome: the most complex macromolecular machine in the cell? Bioessays 25, 1147–1149 (2003).
Jurica, M.S., Licklider, L.J., Gygi, S.R., Grigorieff, N. & Moore, M.J. Purification and characterization of native spliceosomes suitable for three-dimensional structural analysis. RNA 8, 426–439 (2002).
Makarov, E.M. et al. Small nuclear ribonucleoprotein remodeling during catalytic activation of the spliceosome. Science 298, 2205–2208 (2002).
Chen, C.H. et al. Functional and physical interactions between components of the Prp19p-associated complex. Nucleic Acids Res. 30, 1029–1037 (2002).
Ohi, M.D. & Gould, K.L. Characterization of interactions among the Cef1p–Prp19p-associated splicing complex. RNA 8, 798–815 (2002).
Ajuh, P. et al. Functional analysis of the human CDC5L complex and identification of its components by mass spectrometry. EMBO J. 19, 6569–6581 (2000).
Makarova, O.V., Makarov, E.M., Liu, S., Vornlocher, H.P. & Lührmann, R. Protein 61K, encoded by a gene (PRPF31) linked to autosomal dominant retinitis pigmentosa, is required for U4/U6·U5 tri-snRNP formation and pre-mRNA splicing. EMBO J. 21, 1148–1157 (2002).
van Heel, M. et al. Single-particle electron cryomicroscopy: towards atomic resolution. Q. Rev. Biophys. 33, 307–369 (2000).
van Heel, M. Angular reconstitution: a posteriori assignment of projection directions for 3D reconstruction. Ultramicroscopy 21, 111–123 (1987).
Harauz, G. & van Heel, M. Exact filters for general geometry three dimensional reconstruction. Optik 73, 146–156 (1986).
Radermacher, M., Wagenknecht, T., Verschoor, A. & Frank, J. Three-dimensional reconstruction from a single-exposure, random conical tilt series applied to the 50S ribosomal subunit of Escherichia coli. J. Microsc. 146, 113–136 (1987).
Frank, J. Three-Dimensional Electron Microscopy of Macromolecular Assemblies (Academic Press, New York, 1996).
Walz, J. et al. 26S proteasome structure revealed by three-dimensional electron microscopy. J. Struct. Biol. 121, 19–29 (1998).
Ares, M. Jr. & Weiser, B. Rearrangement of snRNA structure during assembly and function of the spliceosome. Prog. Nucleic. Acid. Res. Mol. Biol. 50, 131–159 (1995).
Perriman, R., Barta, I., Voeltz, G.K., Abelson, J. & Ares, M. Jr. ATP requirement for Prp5p function is determined by Cus2p and the structure of U2 small nuclear RNA. Proc. Natl. Acad. Sci. USA 100, 13857–13862 (2003).
Reed, R. Mechanisms of fidelity in pre-mRNA splicing. Curr. Opin. Cell Biol. 12, 340–345 (2000).
Zhou, Z., Sim, J., Griffith, J. & Reed, R. Purification and electron microscopic visualization of functional human spliceosomes. Proc. Natl. Acad. Sci. USA 99, 12203–12207 (2002).
Kastner, B. & Lührmann, R. Electron microscopy of U1 small nuclear ribonucleoprotein particles: shape of the particle and position of the 5′ RNA terminus. EMBO J. 8, 277–286 (1989).
Dube, P., Tavares, P., Lurz, R. & van Heel, M. The portal protein of bacteriophage SPP1: a DNA pump with 13-fold symmetry. EMBO J. 12, 1303–1309 (1993).
van Heel, M. & Frank, J. Use of multivariate statistics in analysing the images of biological macromolecules. Ultramicroscopy 6, 187–194 (1981).
van Heel, M., Harauz, G., Orlova, E.V., Schmidt, R. & Schatz, M. A new generation of the IMAGIC image processing system. J. Struct. Biol. 116, 17–24 (1996).
Chalikian, T.V. Volumetric properties of proteins. Annu. Rev. Biophys. Biomol. Struct. 32, 207–235 (2003).
Jurica, M.S., Sousa, D., Moore, M.J., & Grigorieff, N. Three-dimensional structure of C complex spliceosomes by electron microscopy. Nat. Struct. Mol. Biol. 11, 265–269 (2004).
Acknowledgements
We thank P. Kempkes and H. Kohansal for excellent technical assistance and the Bioverfahrenstechnik division of the Gesellschaft für Biotechnologische Forschung (GBF) in Braunschweig for large-scale HeLa cell cultivation. This work was supported by grants from the Deutsche Forschungsgemeinschaft (Lu294/12–1), the Bundesministerium für Bildung und Forschug (BMBF) (031U215B) and the Fonds der Chemischen Industrie to R.L. and a grant from BMBF (031U215B) to H.S.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Boehringer, D., Makarov, E., Sander, B. et al. Three-dimensional structure of a pre-catalytic human spliceosomal complex B. Nat Struct Mol Biol 11, 463–468 (2004). https://doi.org/10.1038/nsmb761
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb761
This article is cited by
-
Structure of a pre-catalytic spliceosome
Nature (2017)
-
Mechanistic insights into precursor messenger RNA splicing by the spliceosome
Nature Reviews Molecular Cell Biology (2017)
-
Cryo-electron microscopy snapshots of the spliceosome: structural insights into a dynamic ribonucleoprotein machine
Nature Structural & Molecular Biology (2017)
-
Cryo-EM structure of the yeast U4/U6.U5 tri-snRNP at 3.7 Å resolution
Nature (2016)
-
A spliceosome intermediate with loosely associated tri-snRNP accumulates in the absence of Prp28 ATPase activity
Nature Communications (2016)