Abstract
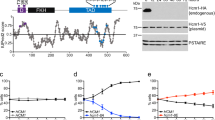
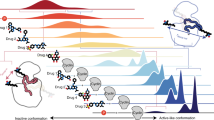
p27 controls cell proliferation by binding and regulating nuclear cyclin-dependent kinases (CDKs). In addition, p27 interacts with other nuclear and cytoplasmic targets and has diverse biological functions. We seek to understand how the structural and dynamic properties of p27 mediate its several functions. We show that, despite showing disorder before binding its targets, p27 has nascent secondary structure that may have a function in molecular recognition. Binding to Cdk2–cyclin A is accompanied by p27 folding, and kinetic data suggest a sequential mechanism that is initiated by binding to cyclin A. p27 regulates CDK–cyclin complexes involved directly in cell cycle control and does not interact with other closely related CDKs. We show that p27-cyclin interactions are an important determinant of this specificity and propose that the homologous cell cycle regulators p21 and p57 function by a similar sequential, folding-on-binding mechanism.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Hengst, L., Dulic, V., Slingerland, J.M., Lees, E. & Reed, S.I. A cell cycle–regulated inhibitor of cyclin-dependent kinases. Proc. Natl. Acad. Sci. USA 91, 5291–5295 (1994).
Polyak, K. et al. Cloning of p27Kip1, a cyclin-dependent kinase inhibitor and a potential mediator of extracellular antimitogenic signals. Cell 78, 59–66 (1994).
Toyoshima, H. & Hunter, T. p27, a novel inhibitor of G1 cyclin–Cdk protein kinase activity, is related to p21. Cell 78, 67–74 (1994).
Gu, Y., Turck, C.W. & Morgan, D.O. Inhibition of CDK2 activity in vivo by an associated 20K regulatory subunit. Nature 366, 707–710 (1993).
El-Deiry, W.S. et al. WAF1, a potential mediator of p53 tumor suppression. Cell 75, 817–825 (1993).
Harper, J.W., Adami, G.R., Wei, N., Keyomarsi, K. & Elledge, S.J. The p21 Cdk-interacting protein Cip1 is a potent inhibitor of G1 cyclin-dependent kinases. Cell 75, 805–816 (1993).
Xiong, Y. et al. p21 is a universal inhibitor of cyclin kinases. Nature 366, 701–704 (1993).
Noda, A., Ning, Y., Venable, S.F., Pereira-Smith, O.M. & Smith, J.R. Cloning of senescent cell–derived inhibitors of DNA synthesis using an expression screen. Exp. Cell Res. 211, 90–98 (1994).
Morgan, D.O. Principles of CDK regulation. Nature 374, 131–134 (1995).
Lee, M.H., Reynisdóttir, I. & Massagué, J. Cloning of p57KIP2, a cyclin-dependent kinase inhibitor with unique domain structure and tissue distribution. Genes Dev. 9, 639–649 (1995).
Matsuoka, S. et al. p57KIP2, a structurally distinct member of the p21CIP1 Cdk inhibitor family, is a candidate tumor suppressor gene. Genes Dev. 9, 650–662 (1995).
Harper, J.W. et al. Inhibition of cyclin-dependent kinases by p21. Mol. Biol. Cell 6, 387–400 (1995).
Baus, F., Gire, V., Fisher, D., Piette, J. & Dulic, V. Permanent cell cycle exit in G2 phase after DNA damage in normal human fibroblasts. EMBO J. 22, 3992–4002 (2003).
van den Heuvel, S. & Harlow, E. Distinct roles for cyclin-dependent kinases in cell cycle control. Science 262, 2050–2054 (1993).
Morgan, D.O. Cyclin-dependent kinases: engines, clocks, and microprocessors. Annu. Rev. Cell Dev. Biol. 13, 261–291 (1997).
Kriwacki, R.W., Hengst, L., Tennant, L., Reed, S.I. & Wright, P.E. Structural studies of p21(waf1/cip1/sdi1) in the free and Cdk2-bound state: conformational disorder mediates binding diversity. Proc. Natl. Acad. Sci. USA 93, 11504–11509 (1996).
Bienkiewicz, E.A., Adkins, J.N. & Lumb, K.J. Functional consequences of preorganized helical structure in the intrinsically disordered cell-cycle inhibitor p27(Kip1). Biochemistry 41, 752–759 (2002).
Russo, A.A., Jeffrey, P.D., Patten, A.K., Massague, J. & Pavletich, N.P. Crystal structure of the p27Kip1 cyclin-dependent-kinase inhibitor bound to the cyclin A–Cdk2 complex. Nature 382, 325–331 (1996).
Wright, P.E. & Dyson, H.J. Intrinsically unstructured proteins: re-assessing the protein structure-function paradigm. J. Mol. Biol. 293, 321–331 (1999).
Dunker, A.K. & Obradovic, Z. The protein trinity—linking function and disorder. Nat. Biotechnol. 19, 805–806 (2001).
Adkins, J.N. & Lumb, K.J. Intrinsic structural disorder and sequence features of the cell cycle inhibitor p57Kip2. Proteins 46, 1–7 (2002).
Tompa, P. Intrinsically unstructured proteins. Trends Biochem. Sci. 27, 527–533 (2002).
Schwarzinger, S. et al. Sequence-dependent correction of random coil NMR chemical shifts. J. Am. Chem. Soc. 123, 2970–2978 (2001).
Wishart, D.S. & Sykes, B.D. Chemical shifts as a tool for structure determination. Methods Enzymol. 239, 363–392 (1994).
Kay, L.E. Protein dynamics from NMR. Nat. Struct. Biol. 5 (Suppl.), 513–517 (1998).
Spolar, R.S. & Record, M.T.J. Coupling of local folding to site-specific binding of proteins to DNA. Science 263, 777–784 (1994).
Pavletich, N.P. Mechanisms of cyclin-dependent kinase regulation: Structures of Cdks, their cyclin activators, and Cip and INK4 inhibitors. J. Mol. Biol. 287, 821–828 (1999).
Jeffrey, P.D. et al. Mechanism of CDK activation revealed by the structure of a cyclinA–CDK2 complex. Nature 376, 313–320 (1995).
Koshland, D.E.J. Application of a theory of enzyme specificity to protein synthesis. Proc. Natl. Acad. Sci. USA 44, 98–104 (1958).
Schulman, B.A., Lindstrom, D.L. & Harlow, E. Substrate recruitment to cyclin-dependent kinase 2 by a multipurpose docking site on cyclin A. Proc. Natl. Acad. Sci. USA 95, 10453–10458 (1998).
Brown, N.R., Noble, M.E., Endicott, J.A. & Johnson, L.N. The structural basis for specificity of substrate and recruitment peptides for cyclin-dependent kinases. Nat. Cell Biol. 1, 438–443 (1999).
Kim, K.K., Chamberlin, H.M., Morgan, D.O. & Kim, S.H. Three-dimensional structure of human cyclin H, a positive regulator of the CDK-activating kinase. Nat. Struct. Biol. 3, 849–855 (1996).
Tarricone, C. et al. Structure and regulation of the CDK5–p25nck5a complex. Mol. Cell 8, 657–669 (2001).
Tanaka, H. et al. Cytoplasmic p21(Cip1/WAF1) regulates neurite remodeling by inhibiting Rho-kinase activity. J. Cell Biol. 158, 321–329 (2002).
Huang, S. et al. Sustained activation of the JNK cascade and rapamycin-induced apoptosis are suppressed by p53/p21(Cip1). Mol. Cell 11, 1491–1501 (2003).
Shim, J., Lee, H., Park, J., Kim, H. & Choi, E.J. A non-enzymatic p21 protein inhibitor of stress-activated protein kinases. Nature 381, 804–806 (1996).
Suzuki, A., Tsutomi, Y., Akahane, K., Araki, T. & Miura, M. Resistance to Fas-mediated apoptosis: activation of caspase 3 is regulated by cell cycle regulator p21WAF1 and IAP gene family ILP. Oncogene 17, 931–939 (1998).
Suzuki, A., Tsutomi, Y., Miura, M. & Akahane, K. Caspase 3 inactivation to suppress Fas-mediated apoptosis: identification of binding domain with p21 and ILP and inactivation machinery by p21. Oncogene 18, 1239–1244 (1999).
Suzuki, A. et al. Procaspase 3/p21 complex formation to resist Fas-mediated cell death is initiated as a result of the phosphorylation of p21 by protein kinase A. Cell Death Differ. 7, 721–728 (2000).
Fujita, N., Sato, S., Katayama, K., Tsuruo, T. Akt-dependent phosphorylation of p27Kip1 promotes binding to 14-3-3 and cytoplasmic localization. J. Biol. Chem. 277, 28706–28713 (2002).
Studier, F.W., Rosenberg, A.H., Dunn, J.J. & Dubendorff, J.W. Use of T7 RNA polymerase to direct expression of cloned genes. Methods Enzymol. 185, 60–89 (1990).
Edelhoch, H. Spectroscopic determination of tryptophan and tyrosine in proteins. Biochemistry 6, 1948–1954 (1967).
Gill, S.C. & von Hippel, P.H. Calculation of protein extinction coefficients from amino acid sequence data. Anal. Biochem. 182, 319–326 (1989).
Neidhardt, F.C., Bloch, P.L. & Smith, D.F. Culture medium for enterobacteria. J. Bacteriol. 119, 736–747 (1974).
Yamazaki, T. et al. An HNCA pulse scheme for the backbone assignment of 15N, 13C, 2H-labeled proteins: application to a 37-kDa Trp repressor–DNA complex. J. Am. Chem. Soc. 116, 6464–6465 (1994).
Yamazaki, T., Lee, W., Arrowsmith, C.H., Muhandiram, D.R. & Kay, L.E. A suite of triple resonance NMR experiments for the backbone assignment of 15N, 13C, 2H labeled proteins with high sensitivity. J. Am. Chem. Soc. 116, 11655–11666 (1994).
Delaglio, F. et al. NMR Pipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Cavanagh, J., Fairbrother, W.J., Palmer, A.G. III & Skelton, N.J. Protein NMR Spectroscopy (Academic, New York, 1996).
Morton, T.A. & Myszka, D.G. Kinetic analysis of macromolecular interactions using surface plasmon resonance biosensors. Methods Enzymol. 295, 268–294 (1998).
Rost, B. PHD: predicting one-dimensional protein structure by profile-based neural networks. Methods Enzymol. 266, 525–539 (1996).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Kriwacki, R.W., Wu, J., Siuzdak, G. & Wright, P.E. Probing protein/protein interactions with mass spectrometry and isotopic labeling: analysis of the p21/Cdk2 complex. J. Am. Chem. Soc. 118, 5320–5321 (1996).
Kriwacki, R.W., Wu, J., Tennant, L., Wright, P.E. & Siuzdak, G. Probing protein structure using biochemical and biophysical methods. Proteolysis, matrix-assisted laser desorption/ionization mass spectrometry, high-performance liquid chromatography and size-exclusion chromatography of p21Waf1/Cip1/Sdi1. J. Chromatogr. A 777, 23–30 (1997).
Acknowledgements
We thank W. Zhang and M. Stewart for assistance with NMR experiments, Y. Wang for assistance with protein production, C. Ross for computer support, P. Bowman for assistance with p27 structure figures and movie, G. Siuzdak, B. Bothner, C. Galea and T. Record for critically reading the manuscript and N. Pavletich for providing a cyclin A expression plasmid. R. Muhandiram and L. Kay are acknowledged for providing Varian NMR pulse sequences. The authors acknowledge support from the American Lebanese Syrian Associated Charities (ALSAC), the US National Cancer Institute (P30 CA 21765 and RO1 CA 82491), and the US National Center for Research Resources (S10 RR014675).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Lacy, E., Filippov, I., Lewis, W. et al. p27 binds cyclin–CDK complexes through a sequential mechanism involving binding-induced protein folding. Nat Struct Mol Biol 11, 358–364 (2004). https://doi.org/10.1038/nsmb746
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb746
This article is cited by
-
Minimum information guidelines for experiments structurally characterizing intrinsically disordered protein regions
Nature Methods (2023)
-
Quantitative model of eukaryotic Cdk control through the Forkhead CONTROLLER
npj Systems Biology and Applications (2021)
-
Surface charge of Merkel cell polyomavirus small T antigen determines cell transformation through allosteric FBW7 WD40 domain targeting
Oncogenesis (2020)
-
The intracellular domain of BP180/collagen XVII is intrinsically disordered and partially folds in an anionic membrane lipid-mimicking environment
Amino Acids (2020)
-
p27kip1 at the crossroad between actin and microtubule dynamics
Cell Division (2019)