Abstract
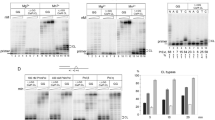
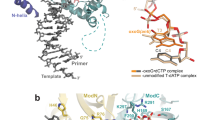
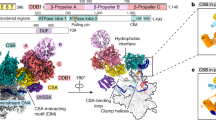
The anticancer drug cisplatin forms 1,2-d(GpG) DNA intrastrand cross-links (cisplatin lesions) that stall RNA polymerase II (Pol II) and trigger transcription-coupled DNA repair. Here we present a structure-function analysis of Pol II stalling at a cisplatin lesion in the DNA template. Pol II stalling results from a translocation barrier that prevents delivery of the lesion to the active site. AMP misincorporation occurs at the barrier and also at an abasic site, suggesting that it arises from nontemplated synthesis according to an 'A-rule' known for DNA polymerases. Pol II can bypass a cisplatin lesion that is artificially placed beyond the translocation barrier, even in the presence of a G·A mismatch. Thus, the barrier prevents transcriptional mutagenesis. The stalling mechanism differs from that of Pol II stalling at a photolesion, which involves delivery of the lesion to the active site and lesion-templated misincorporation that blocks transcription.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Wang, D. & Lippard, S.J. Cellular processing of platinum anticancer drugs. Nat. Rev. Drug Discov. 4, 307–320 (2005).
Kartalou, M. & Essigmann, J.M. Recognition of cisplatin adducts by cellular proteins. Mutat. Res. 478, 1–21 (2001).
Jung, Y. & Lippard, S.J. Multiple states of stalled T7 RNA polymerase at DNA lesions generated by platinum anticancer agents. J. Biol. Chem. 278, 52084–52092 (2003).
Corda, Y., Job, C., Anin, M.F., Leng, M. & Job, D. Transcription by eucaryotic and procaryotic RNA polymerases of DNA modified at a d(GG) or a d(AG) site by the antitumor drug cis-diamminedichloroplatinum(II). Biochemistry 30, 222–230 (1991).
Corda, Y., Job, C., Anin, M.F., Leng, M. & Job, D. Spectrum of DNA-platinum adduct recognition by prokaryotic and eukaryotic DNA-dependent RNA polymerases. Biochemistry 32, 8582–8588 (1993).
Tornaletti, S., Patrick, S.M., Turchi, J.J. & Hanawalt, P.C. Behavior of T7 RNA polymerase and mammalian RNA polymerase II at site-specific cisplatin adducts in the template DNA. J. Biol. Chem. 278, 35791–35797 (2003).
Jung, Y. & Lippard, S.J. RNA polymerase II blockage by cisplatin-damaged DNA. Stability and polyubiquitylation of stalled polymerase. J. Biol. Chem. 281, 1361–1370 (2006).
Brueckner, F., Hennecke, U., Carell, T. & Cramer, P. CPD damage recognition by transcribing RNA polymerase II. Science 315, 859–862 (2007).
Kettenberger, H., Armache, K.-J. & Cramer, P. Complete RNA polymerase II elongation complex structure and its interactions with NTP and TFIIS. Mol. Cell 16, 955–965 (2004).
Gelasco, A. & Lippard, S.J. NMR solution structure of a DNA dodecamer duplex containing a cis-diammineplatinum(II) d(GpG) intrastrand cross-link, the major adduct of the anticancer drug cisplatin. Biochemistry 37, 9230–9239 (1998).
Takahara, P.M., Rosenzweig, A.C., Frederick, C.A. & Lippard, S.J. Crystal structure of double-stranded DNA containing the major adduct of the anticancer drug cisplatin. Nature 377, 649–652 (1995).
Kashkina, E. et al. Template misalignment in multisubunit RNA polymerases and transcription fidelity. Mol. Cell 24, 257–266 (2006).
Gnatt, A.L., Cramer, P., Fu, J., Bushnell, D.A. & Kornberg, R.D. Structural basis of transcription: an RNA polymerase II elongation complex at 3.3 Å resolution. Science 292, 1876–1882 (2001).
Strauss, B.S. The 'A rule' of mutagen specificity: a consequence of DNA polymerase bypass of non-instructional lesions? Bioessays 13, 79–84 (1991).
Taylor, J.S. New structural and mechanistic insight into the A-rule and the instructional and non-instructional behavior of DNA photoproducts and other lesions. Mutat. Res. 510, 55–70 (2002).
Wind, M. & Reines, D. Transcription elongation factor SII. Bioessays 22, 327–336 (2000).
Thomas, M.J., Platas, A.A. & Hawley, D.K. Transcriptional fidelity and proofreading by RNA polymerase II. Cell 93, 627–637 (1998).
Tremeau-Bravard, A., Riedl, T., Egly, J.M. & Dahmus, M.E. Fate of RNA polymerase II stalled at a cisplatin lesion. J. Biol. Chem. 279, 7751–7759 (2004).
Laine, J.P. & Egly, J.M. Initiation of DNA repair mediated by a stalled RNA polymerase IIO. EMBO J. 25, 387–397 (2006).
Armache, K.-J., Kettenberger, H. & Cramer, P. Architecture of the initiation-competent 12-subunit RNA polymerase II. Proc. Natl. Acad. Sci. USA 100, 6964–6968 (2003).
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Cryst. 26, 795–800 (1993).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1996).
McCoy, A.J., Grosse-Kunstleve, R.W., Storoni, L.C. & Read, R.J. Likelihood-enhanced fast translation functions. Acta Crystallogr. D Biol. Crystallogr. 61, 458–464 (2005).
Armache, K.-J., Mitterweger, S., Meinhart, A. & Cramer, P. Structures of complete RNA polymerase II and its subcomplex Rpb4/7. J. Biol. Chem. 280, 7131–7134 (2005).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Acknowledgements
We thank members of the Cramer laboratory for help. P.C. and T.C. were supported by the Deutsche Forschungsgemeinschaft, the Sonderforschungsbereich SFB646 and the Fonds der chemischen Industrie. P.C. was supported by the EU grant 3D repertoire, contract no. LSHG-CT-2005-512028. G.E.D. and F.B. were supported by the Elite Netzwerk Bayern. A.A. was supported by the Marie Curie training and mobility network CLUSTOX DNA.
Author information
Authors and Affiliations
Contributions
G.E.D. performed and analyzed biochemical and crystallographic experiments. A.A. synthesized cisplatin-containing DNA strands and performed MALDI experiments. F.B. assisted with experiments and crystallography. T.C. supervised the project. P.C. supervised the project and wrote the manuscript.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Table 1 (PDF 1161 kb)
Rights and permissions
About this article
Cite this article
Damsma, G., Alt, A., Brueckner, F. et al. Mechanism of transcriptional stalling at cisplatin-damaged DNA. Nat Struct Mol Biol 14, 1127–1133 (2007). https://doi.org/10.1038/nsmb1314
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1314
This article is cited by
-
Organelle-targeted therapies: a comprehensive review on system design for enabling precision oncology
Signal Transduction and Targeted Therapy (2022)
-
Transcriptional processing of an unnatural base pair by eukaryotic RNA polymerase II
Nature Chemical Biology (2021)
-
Human HMGN1 and HMGN2 are not required for transcription-coupled DNA repair
Scientific Reports (2020)
-
APE1 and NPM1 protect cancer cells from platinum compounds cytotoxicity and their expression pattern has a prognostic value in TNBC
Journal of Experimental & Clinical Cancer Research (2019)
-
The DNA damage response to transcription stress
Nature Reviews Molecular Cell Biology (2019)