Abstract
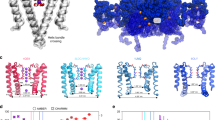
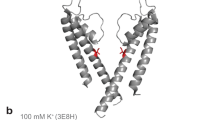
K+ channels conduct and regulate K+ flux across the cell membrane. Several crystal structures and biophysical studies of tetrameric ion channels have revealed many of the structural details of ion selectivity and gating. A narrow pore lined with four arrays of carbonyl groups is responsible for ion selectivity, whereas a conformational change of the four inner transmembrane helices (TM2) is involved in gating. We used NMR to examine full-length KcsA, a prototypical K+ channel, in its open, closed and intermediate states. These studies reveal that at least two conformational states exist both in the selectivity filter and near the C-terminal ends of the TM2 helices. In the ion-conducting open state, we observed rapid structural exchange between two conformations of the filter, presumably of low and high K+ affinity, respectively. Such measurements of millisecond-timescale dynamics reveal the basis for simultaneous ion selection and gating.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hille, B. Ion Channels of Excitable Membranes 3rd edn. (Sinauder, Sunderland, Massachusetts, USA, 2001).
Doyle, D.A. et al. The structure of the potassium channel: molecular basis of K+ conduction and selectivity. Science 280, 69–77 (1998).
Jiang, Y. et al. Crystal structure and mechanism of a calcium-gated potassium channel. Nature 417, 515–522 (2002).
Kuo, A. et al. Crystal structure of the potassium channel KirBac1.1 in the closed state. Science 300, 1922–1926 (2003).
Jiang, Y. et al. X-ray structure of a voltage-dependent K+ channel. Nature 423, 33–41 (2003).
Shi, N., Ye, S., Alam, A., Chen, L. & Jiang, Y. Atomic structure of a Na+ and K+-conducting channel. Nature 440, 570–574 (2006).
Valiyaveetil, F.I., Sekedat, M., MacKinnon, R. & Muir, T.W. Glycine as a d-amino acid surrogate in the K+-selectivity filter. Proc. Natl. Acad. Sci. USA 101, 17045–17049 (2004).
Bezanilla, F. & Armstrong, C.M. Negative conductance caused by entry of sodium and cesium ions into the potassium channels of squid axons. J. Gen. Physiol. 60, 588–608 (1972).
Noskov, S.Y. & Roux, B. Importance of hydration and dynamics on the selectivity of the KcsA and NaK channels. J. Gen. Physiol. 129, 135–143 (2007).
Perozo, E., Cortes, D.M. & Cuello, L.G. Structural rearrangements underlying K+-channel activation gating. Science 285, 73–78 (1999).
Heginbotham, L., LeMasurier, M., Kolmakova-Partensky, L. & Miller, C. Single Streptomyces lividans K+ channels: functional asymmetries and sidedness of proton activation. J. Gen. Physiol. 114, 551–560 (1999).
Liu, Y.S., Sompornpisut, P. & Perozo, E. Structure of the KcsA channel intracellular gate in the open state. Nat. Struct. Biol. 8, 883–887 (2001).
Jiang, Y. et al. The open pore conformation of potassium channels. Nature 417, 523–526 (2002).
Cordero-Morales, J.F. et al. Molecular determinants of gating at the potassium-channel selectivity filter. Nat. Struct. Mol. Biol. 13, 311–318 (2006).
Valiyaveetil, F.I., Sekedat, M., MacKinnon, R. & Muir, T.W. Structural and functional consequences of an amide-to-ester substitution in the selectivity filter of a potassium channel. J. Am. Chem. Soc. 128, 11591–11599 (2006).
Proks, P., Capener, C.E., Jones, P. & Ashcroft, F. Mutations within the P-loop of Kir6.2 modulates the intraburst kinetics of the ATP-sensitive potassium channel. J. Gen. Physiol. 118, 341–353 (2001).
Berneche, S. & Roux, B. Energetics of ion conduction through the K+ channel. Nature 414, 73–77 (2001).
MacKinnon, R., Cohen, S.L., Kuo, A., Lee, A. & Chait, B.T. Structural conversion in prokaryotic and eukaryotic potassium channels. Science 280, 106–109 (1998).
Chill, J.H., Louis, J.M., Miller, C. & Bax, A. NMR study of the tetrameric KcsA potassium channel in detergent micelles. Protein Sci. 15, 684–698 (2006).
Pervushin, K., Riek, R., Wider, G. & Wuthrich, K. Attenuated T2 relaxation by mutual cancellation of dipole-dipole coupling and chemical shift anisotropy indicates an avenue to NMR structures of very large biological macromolecules in solution. Proc. Natl. Acad. Sci. USA 94, 12366–12371 (1997).
Zhou, Y., Morais-Cabral, J.H., Kaufman, A. & MacKinnon, R. Chemistry of ion coordination and hydration revealed by the K+ channel-Fab complex at 2.0 Å resolution. Nature 414, 43–48 (2001).
Cortes, D.M., Cuello, L.G. & Perozo, E. Molecular architecture of full-length KcsA: role of cytoplasmic domains in ion permeation and activation gating. J. Gen. Physiol. 117, 165–180 (2001).
Cuello, L.G., Romero, J.G., Cortes, D.M. & Perozo, E. pH-dependent gating in the Streptomyces lividans K+ channel. Biochemistry 37, 3229–3236 (1998).
Lu, T. et al. Probing ion permeation and gating in a K+ channel with backbone mutations in the selectivity filter. Nat. Neurosci. 4, 239–246 (2001).
Wuthrich, K. NMR of Proteins and Nucleic Acids (Wiley, Indianapolis, 1986).
Takeuchi, K., Takahashi, H., Kawano, S. & Shimada, I. Identification and characterization of the slowly exchanging pH-dependent conformational rearrangement in KcsA. J. Biol. Chem. 282, 15179–15186 (2007).
Noskov, S.Y. & Roux, B. Ion selectivity in potassium channels. Biophys. Chem. 124, 279–291 (2006).
Luzhkov, V.B. & Aqvist, J. A computational study of ion binding and protonation states in the KcsA potassium channel. Biochim. Biophys. Acta 1481, 360–370 (2000).
Bucher, D., Guidoni, L. & Roethlisberger, U. The protonation state of the Glu-71/Asp-80 residues in the KcsA potassium channel. A first-principles QM/MM molecular dynamics study. Biophys. J. 93, 2315–2324 (2007).
Zhou, Y. & MacKinnon, R. The occupancy of ions in the K+ selectivity filter: charge balance and coupling of ion binding to a protein conformational change underlie high conduction rates. J. Mol. Biol. 333, 965–975 (2003).
VanDongen, A.M. K channel gating by an affinity-switching selectivity filter. Proc. Natl. Acad. Sci. USA 101, 3248–3252 (2004).
LeMaster, D.M. & Richards, F.M. NMR sequential assignment of Escherichia coli thioredoxin utilizing random fractional deuteriation. Biochemistry 27, 142–150 (1988).
Marley, J., Lu, M. & Bracken, C. A method for efficient isotopic labeling of recombinant proteins. J. Biomol. NMR 20, 71–75 (2001).
Salzmann, M., Wider, G., Pervushin, K. & Wuthrich, K. Improved sensitivity and coherence selection for [15N,1H]-TROSY elements in triple resonance experiments. J. Biomol. NMR 15, 181–184 (1999).
Guntert, P., Dotsch, V., Wider, G. & Wuthrich, K. Processing of multi-dimensional NMR data with the new software PROSA. J. Biomol. NMR 2, 619–629 (1992).
Acknowledgements
This work was supported in part by the US National Institutes of Health (GM74929 and GM56653). R.R. is a Pew Scholar and the Helen McLoraine Development Chair in Neurobiology. K.A.B. would like to thank the American Heart Association for fellowship support.
Author information
Authors and Affiliations
Contributions
K.A.B. and C.T. prepared KcsA; K.A.B., C.T., W.K. and R.R. collected and analyzed NMR data; K.A.B., C.T., S.C. and R.R. contributed to scientific discussions and prepared the manuscript.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Table 1–3, Supplementary Methods (PDF 4761 kb)
Rights and permissions
About this article
Cite this article
Baker, K., Tzitzilonis, C., Kwiatkowski, W. et al. Conformational dynamics of the KcsA potassium channel governs gating properties. Nat Struct Mol Biol 14, 1089–1095 (2007). https://doi.org/10.1038/nsmb1311
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1311
This article is cited by
-
Central cavity dehydration as a gating mechanism of potassium channels
Nature Communications (2023)
-
Conformational equilibrium shift underlies altered K+ channel gating as revealed by NMR
Nature Communications (2020)
-
Shifts in the selectivity filter dynamics cause modal gating in K+ channels
Nature Communications (2019)
-
Site-specific ion occupation in the selectivity filter causes voltage-dependent gating in a viral K+ channel
Scientific Reports (2018)
-
Decoupled side chain and backbone dynamics for proton translocation – M2 of influenza A
Journal of Molecular Modeling (2017)