Abstract
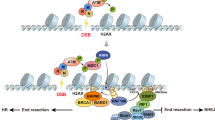
Histone acetylation is important in regulating DNA accessibility. Multifunctional Sin3 proteins bind histone deacetylases (HDACs) to assemble silencing complexes that selectively target chromatin. We show that, in fission yeast, an essential HDAC, Clr6, exists in two distinct Sin3 core complexes. Complex I contains an essential Sin3 homolog, Pst1, and other factors, and predominantly targets gene promoters. Complex II contains a nonessential Sin3 homolog, Pst2, and several conserved proteins. It preferentially targets transcribed chromosomal regions and centromere cores. Defects in complex II abrogate global protective functions of chromatin, causing increased accessibility of DNA to genotoxic agents and widespread antisense transcripts that are processed by the exosome. Notably, the two Clr6 complexes differentially repress forward and reverse centromeric repeat transcripts, suggesting that these complexes regulate transcription in heterochromatin and euchromatin in similar manners, including suppression of spurious transcripts from cryptic start sites.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Jenuwein, T. & Allis, C.D. Translating the histone code. Science 293, 1074–1080 (2001).
Kurdistani, S.K. & Grunstein, M. Histone acetylation and deacetylation in yeast. Nat. Rev. Mol. Cell Biol. 4, 276–284 (2003).
Ekwall, K. Genome-wide analysis of HDAC function. Trends Genet. 21, 608–615 (2005).
Wurtele, H. & Verreault, A. Histone post-translational modifications and the response to DNA double-strand breaks. Curr. Opin. Cell Biol. 18, 137–144 (2006).
Grewal, S.I., Bonaduce, M.J. & Klar, A.J. Histone deacetylase homologs regulate epigenetic inheritance of transcriptional silencing and chromosome segregation in fission yeast. Genetics 150, 563–576 (1998).
Bjerling, P. et al. Functional divergence between histone deacetylases in fission yeast by distinct cellular localization and in vivo specificity. Mol. Cell. Biol. 22, 2170–2181 (2002).
Shankaranarayana, G.D., Motamedi, M.R., Moazed, D. & Grewal, S.I. Sir2 regulates histone H3 lysine 9 methylation and heterochromatin assembly in fission yeast. Curr. Biol. 13, 1240–1246 (2003).
Hansen, K.R. et al. Global effects on gene expression in fission yeast by silencing and RNA interference machineries. Mol. Cell. Biol. 25, 590–601 (2005).
Wiren, M. et al. Genomewide analysis of nucleosome density histone acetylation and HDAC function in fission yeast. EMBO J. 24, 2906–2918 (2005).
Sugiyama, T. et al. SHREC, an effector complex for heterochromatic transcriptional silencing. Cell 128, 491–504 (2007).
Nakayama, J. et al. Alp13, an MRG family protein, is a component of fission yeast Clr6 histone deacetylase required for genomic integrity. EMBO J. 22, 2776–2787 (2003).
Verreault, A., Kaufman, P.D., Kobayashi, R. & Stillman, B. Nucleosomal DNA regulates the core-histone-binding subunit of the human Hat1 acetyltransferase. Curr. Biol. 8, 96–108 (1998).
Bertram, M.J. & Pereira-Smith, O.M. Conservation of the MORF4 related gene family: identification of a new chromo domain subfamily and novel protein motif. Gene 266, 111–121 (2001).
Carrozza, M.J. et al. Histone H3 methylation by Set2 directs deacetylation of coding regions by Rpd3S to suppress spurious intragenic transcription. Cell 123, 581–592 (2005).
Joshi, A.A. & Struhl, K. Eaf3 chromodomain interaction with methylated H3–K36 links histone deacetylation to Pol II elongation. Mol. Cell 20, 971–978 (2005).
Keogh, M.C. et al. Cotranscriptional set2 methylation of histone H3 lysine 36 recruits a repressive Rpd3 complex. Cell 123, 593–605 (2005).
Silverstein, R.A. & Ekwall, K. Sin3: a flexible regulator of global gene expression and genome stability. Curr. Genet. 47, 1–17 (2005).
Hassig, C.A., Fleischer, T.C., Billin, A.N., Schreiber, S.L. & Ayer, D.E. Histone deacetylase activity is required for full transcriptional repression by mSin3A. Cell 89, 341–347 (1997).
Zhang, Y., Iratni, R., Erdjument-Bromage, H., Tempst, P. & Reinberg, D. Histone deacetylases and SAP18, a novel polypeptide, are components of a human Sin3 complex. Cell 89, 357–364 (1997).
Silverstein, R.A., Richardson, W., Levin, H., Allshire, R. & Ekwall, K. A new role for the transcriptional corepressor SIN3; regulation of centromeres. Curr. Biol. 13, 68–72 (2003).
Alland, L. et al. Identification of mammalian Sds3 as an integral component of the Sin3/histone deacetylase corepressor complex. Mol. Cell. Biol. 22, 2743–2750 (2002).
Vannier, D., Balderes, D. & Shore, D. Evidence that the transcriptional regulators SIN3 and RPD3, and a novel gene (SDS3) with similar functions, are involved in transcriptional silencing in S. cerevisiae. Genetics 144, 1343–1353 (1996).
Meehan, W.J. et al. Breast cancer metastasis suppressor 1 (BRMS1) forms complexes with retinoblastoma-binding protein 1 (RBP1) and the mSin3 histone deacetylase complex and represses transcription. J. Biol. Chem. 279, 1562–1569 (2004).
David, G., Turner, G.M., Yao, Y., Protopopov, A. & DePinho, R.A. mSin3-associated protein, mSds3, is essential for pericentric heterochromatin formation and chromosome segregation in mammalian cells. Genes Dev. 17, 2396–2405 (2003).
Loewith, R., Meijer, M., Lees-Miller, S.P., Riabowol, K. & Young, D. Three yeast proteins related to the human candidate tumor suppressor p33(ING1) are associated with histone acetyltransferase activities. Mol. Cell. Biol. 20, 3807–3816 (2000).
Feng, X., Hara, Y. & Riabowol, K. Different HATS of the ING1 gene family. Trends Cell Biol. 12, 532–538 (2002).
Doyon, Y. et al. ING tumor suppressor proteins are critical regulators of chromatin acetylation required for genome expression and perpetuation. Mol. Cell 21, 51–64 (2006).
Kuzmichev, A., Zhang, Y., Erdjument-Bromage, H., Tempst, P. & Reinberg, D. Role of the Sin3-histone deacetylase complex in growth regulation by the candidate tumor suppressor p33(ING1). Mol. Cell. Biol. 22, 835–848 (2002).
Skowyra, D. et al. Differential association of products of alternative transcripts of the candidate tumor suppressor ING1 with the mSin3/HDAC1 transcriptional corepressor complex. J. Biol. Chem. 276, 8734–8739 (2001).
Dang, V.D. et al. A new member of the Sin3 family of corepressors is essential for cell viability and required for retroelement propagation in fission yeast. Mol. Cell. Biol. 19, 2351–2365 (1999).
Cam, H.P. et al. Comprehensive analysis of heterochromatin- and RNAi-mediated epigenetic control of the fission yeast genome. Nat. Genet. 37, 809–819 (2005).
Volpe, T.A. et al. Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297, 1833–1837 (2002).
Chikashige, Y. et al. Composite motifs and repeat symmetry in S. pombe centromeres: direct analysis by integration of NotI restriction sites. Cell 57, 739–751 (1989).
Grewal, S.I. & Klar, A.J. A recombinationally repressed region between mat2 and mat3 loci shares homology to centromeric repeats and regulates directionality of mating-type switching in fission yeast. Genetics 146, 1221–1238 (1997).
Minoda, A., Saitoh, S., Takahashi, K. & Toda, T. BAF53/Arp4 homolog Alp5 in fission yeast is required for histone H4 acetylation, kinetochore-spindle attachment, and gene silencing at centromere. Mol. Biol. Cell 16, 316–327 (2005).
Grewal, S.I. & Jia, S. Heterochromatin revisited. Nat. Rev. Genet. 8, 35–46 (2007).
Thon, G. & Klar, A.J. The clr1 locus regulates the expression of the cryptic mating-type loci of fission yeast. Genetics 131, 287–296 (1992).
Carrozza, M.J. et al. Stable incorporation of sequence specific repressors Ash1 and Ume6 into the Rpd3L complex. Biochim. Biophys. Acta 1731, 77–87 (2005).
Morris, S.A. et al. Histone H3 K36 methylation is associated with transcription elongation in Schizosaccharomyces pombe. Eukaryot. Cell 4, 1446–1454 (2005).
Huang, Y., Bayfield, M.A., Intine, R.V. & Maraia, R.J. Separate RNA-binding surfaces on the multifunctional La protein mediate distinguishable activities in tRNA maturation. Nat. Struct. Mol. Biol. 13, 611–618 (2006).
Houseley, J., LaCava, J. & Tollervey, D. RNA-quality control by the exosome. Nat. Rev. Mol. Cell Biol. 7, 529–539 (2006).
Bird, A.W. et al. Acetylation of histone H4 by Esa1 is required for DNA double-strand break repair. Nature 419, 411–415 (2002).
Chen, J. & Stubbe, J. Bleomycins: towards better therapeutics. Nat. Rev. Cancer 5, 102–112 (2005).
Lambert, S. & Carr, A.M. Checkpoint responses to replication fork barriers. Biochimie 87, 591–602 (2005).
Vidal, M. & Gaber, R.F. RPD3 encodes a second factor required to achieve maximum positive and negative transcriptional states in Saccharomyces cerevisiae. Mol. Cell. Biol. 11, 6317–6327 (1991).
Reid, J.L., Moqtaderi, Z. & Struhl, K. Eaf3 regulates the global pattern of histone acetylation in Saccharomyces cerevisiae. Mol. Cell. Biol. 24, 757–764 (2004).
Zhang, Y. It takes a PHD to interpret histone methylation. Nat. Struct. Mol. Biol. 13, 572–574 (2006).
Noma, K. et al. RITS acts in cis to promote RNA interference-mediated transcriptional and post-transcriptional silencing. Nat. Genet. 36, 1174–1180 (2004).
Djupedal, I. et al. RNA Pol II subunit Rpb7 promotes centromeric transcription and RNAi-directed chromatin silencing. Genes Dev. 19, 2301–2306 (2005).
Nakagawa, H. et al. Fission yeast CENP-B homologs nucleate centromeric heterochromatin by promoting heterochromatin-specific histone tail modifications. Genes Dev. 16, 1766–1778 (2002).
Katayama, S. et al. Antisense transcription in the mammalian transcriptome. Science 309, 1564–1566 (2005).
Wyers, F. et al. Cryptic pol II transcripts are degraded by a nuclear quality control pathway involving a new poly(A) polymerase. Cell 121, 725–737 (2005).
Grewal, S.I. & Moazed, D. Heterochromatin and epigenetic control of gene expression. Science 301, 798–802 (2003).
Li, X. & Manley, J.L. Cotranscriptional processes and their influence on genome stability. Genes Dev. 20, 1838–1847 (2006).
Huertas, P. & Aguilera, A. Cotranscriptionally formed DNA:RNA hybrids mediate transcription elongation impairment and transcription-associated recombination. Mol. Cell 12, 711–721 (2003).
Jazayeri, A., McAinsh, A.D. & Jackson, S.P. Saccharomyces cerevisiae Sin3p facilitates DNA double-strand break repair. Proc. Natl. Acad. Sci. USA 101, 1644–1649 (2004).
Tamburini, B.A. & Tyler, J.K. Localized histone acetylation and deacetylation triggered by the homologous recombination pathway of double-strand DNA repair. Mol. Cell. Biol. 25, 4903–4913 (2005).
Shogren-Knaak, M. et al. Histone H4–K16 acetylation controls chromatin structure and protein interactions. Science 311, 844–847 (2006).
Ekwall, K., Olsson, T., Turner, B.M., Cranston, G. & Allshire, R.C. Transient inhibition of histone deacetylation alters the structural and functional imprint at fission yeast centromeres. Cell 91, 1021–1032 (1997).
Kelly, T.J. et al. The fission yeast cdc18+ gene product couples S phase to START and mitosis. Cell 74, 371–382 (1993).
Acknowledgements
We thank J. Nakayama and A. Malikzay for helpful contributions, H. Levin (National Institute of Child Health and Human Development, NIH), K. Ekwall (Karolinska Institutet), R. Maraia (National Institute of Child Health and Human Development, NIH) for strains, M. Lichten, O. Sordet and J. Sabl for help in editing the manuscript and K. Noma, G. Mizuguchi, D. Eyre and M. Lichten for helpful discussions and protocols. This research was supported by the Intramural Research Program of the NIH, National Cancer Institute.
Author information
Authors and Affiliations
Contributions
S.I.S.G. and E.N. designed experiments; E.N. performed all biochemical and genetic experiments; E.N and H.P.C. performed the microarray experiments; T.Y. analyzed DNA prepared from bleomycin-treated cells by pulse-field gel electrophoresis; R.K. analyzed purified protein samples by mass spectrometry; P.C.F. helped analyze microarray expression data; S.I.S.G and E.N. wrote the paper; S.I.S.G. and H.P.C. edited the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Genetic and biochemical characterization of Clr6-associated factors. (PDF 3008 kb)
Supplementary Fig. 2
Complexes I and II differentially affect histone acetylation at the promoters and the coding regions of genes. (PDF 222 kb)
Supplementary Fig. 3
Complexes I and II preferentially regulate sense and antisense transcripts. (PDF 63 kb)
Supplementary Fig. 4
Complex II suppresses antisense transcription at the zer1 and hrp1 coding regions. (PDF 558 kb)
Supplementary Fig. 5
Antisense transcription does not correlate with detectable levels of heterochromatic modifications at euchromatic loci. (PDF 2626 kb)
Supplementary Fig. 6
Complex I, but not complex II, is required for silencing of donor mating-type loci. (PDF 553 kb)
Supplementary Fig. 7
Distribution of H3K9 methylation in alp13rrp6 double mutant strain. (PDF 927 kb)
Supplementary Fig. 8
set2 and rrp6 mutant strains are sensitive to bleomycin treatment. (PDF 260 kb)
Supplementary Table 1
Loci showing upregulated antisense transcripts in alp13Δ mutant background. (PDF 39 kb)
Supplementary Table 2
Clr6 complex I and complex II subunits in S. pombe and their homologs in S. cerevisiae and mammals. (PDF 92 kb)
Rights and permissions
About this article
Cite this article
Nicolas, E., Yamada, T., Cam, H. et al. Distinct roles of HDAC complexes in promoter silencing, antisense suppression and DNA damage protection. Nat Struct Mol Biol 14, 372–380 (2007). https://doi.org/10.1038/nsmb1239
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1239
This article is cited by
-
Spurious transcription causing innate immune responses is prevented by 5-hydroxymethylcytosine
Nature Genetics (2023)
-
Structural basis of nucleosome deacetylation and DNA linker tightening by Rpd3S histone deacetylase complex
Cell Research (2023)
-
Rbm10 facilitates heterochromatin assembly via the Clr6 HDAC complex
Epigenetics & Chromatin (2021)
-
Emerging roles of centromeric RNAs in centromere formation and function
Genes & Genomics (2021)
-
Systematic analysis reveals the prevalence and principles of bypassable gene essentiality
Nature Communications (2019)