Abstract
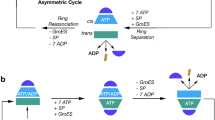
Chaperonins are allosteric double-ring ATPases that mediate cellular protein folding. ATP binding and hydrolysis control opening and closing of the central chaperonin chamber, which transiently provides a protected environment for protein folding. During evolution, two strategies to close the chaperonin chamber have emerged. Archaeal and eukaryotic group II chaperonins contain a built-in lid, whereas bacterial chaperonins use a ring-shaped cofactor as a detachable lid. Here we show that the built-in lid is an allosteric regulator of group II chaperonins, which helps synchronize the subunits within one ring and, to our surprise, also influences inter-ring communication. The lid is dispensable for substrate binding and ATP hydrolysis, but is required for productive substrate folding. These regulatory functions of the lid may serve to allow the symmetrical chaperonins to function as 'two-stroke' motors and may also provide a timer for substrate encapsulation within the closed chamber.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Young, J.C., Agashe, V.R., Siegers, K. & Hartl, F.U. Pathways of chaperone-mediated protein folding in the cytosol. Nat. Rev. Mol. Cell Biol. 5, 781–791 (2004).
Horwich, A.L., Farr, G.W. & Fenton, W.A. GroEL-GroES-mediated protein folding. Chem. Rev. 106, 1917–1930 (2006).
Frydman, J. Folding of newly translated proteins in vivo: the role of molecular chaperones. Annu. Rev. Biochem. 70, 603–647 (2001).
Bukau, B. & Horwich, A.L. The Hsp70 and Hsp60 chaperone machines. Cell 92, 351–366 (1998).
Sigler, P.B. et al. Structure and function in GroEL-mediated protein folding. Annu. Rev. Biochem. 67, 581–608 (1998).
Gutsche, I., Essen, L.O. & Baumeister, W. Group II chaperonins: new TRiC(k)s and turns of a protein folding machine. J. Mol. Biol. 293, 295–312 (1999).
Spiess, C., Meyer, A.S., Reissmann, S. & Frydman, J. Mechanism of the eukaryotic chaperonin: protein folding in the chamber of secrets. Trends Cell Biol. 14, 598–604 (2004).
Ditzel, L. et al. Crystal structure of the thermosome, the archaeal chaperonin and homolog of CCT. Cell 93, 125–138 (1998).
Meyer, A.S. et al. Closing the folding chamber of the eukaryotic chaperonin requires the transition state of ATP hydrolysis. Cell 113, 369–381 (2003).
Shomura, Y. et al. Crystal structures of the group II chaperonin from Thermococcus strain KS-1: steric hindrance by the substituted amino acid, and inter-subunit rearrangement between two crystal forms. J. Mol. Biol. 335, 1265–1278 (2004).
Horovitz, A., Fridmann, Y., Kafri, G. & Yifrach, O. Review: allostery in chaperonins. J. Struct. Biol. 135, 104–114 (2001).
Saibil, H.R., Horwich, A.L. & Fenton, W.A. Allostery and protein substrate conformational change during GroEL/GroES-mediated protein folding. Adv. Protein Chem. 59, 45–72 (2001).
Swain, J.F. & Gierasch, L.M. The changing landscape of protein allostery. Curr. Opin. Struct. Biol. 16, 102–108 (2006).
Kafri, G. & Horovitz, A. Transient kinetic analysis of ATP-induced allosteric transitions in the eukaryotic chaperonin containing TCP-1. J. Mol. Biol. 326, 981–987 (2003).
Yifrach, O. & Horovitz, A. Transient kinetic analysis of adenosine 5′-triphosphate binding-induced conformational changes in the allosteric chaperonin GroEL. Biochemistry 37, 7083–7088 (1998).
Cliff, M.J., Limpkin, C., Cameron, A., Burston, S.G. & Clarke, A.R. Elucidation of steps in the capture of a protein substrate for efficient encapsulation by GroE. J. Biol. Chem. 281, 21266–21275 (2006).
Bigotti, M.G. & Clarke, A.R. Cooperativity in the thermosome. J. Mol. Biol. 348, 13–26 (2005).
Bigotti, M.G., Bellamy, S.R. & Clarke, A.R. The asymmetric ATPase cycle of the thermosome: elucidation of the binding, hydrolysis and product-release steps. J. Mol. Biol. 362, 835–843 (2006).
Yifrach, O. & Horovitz, A. Nested cooperativity in the ATPase activity of the oligomeric chaperonin GroEL. Biochemistry 34, 5303–5308 (1995).
Kafri, G., Willison, K.R. & Horovitz, A. Nested allosteric interactions in the cytoplasmic chaperonin containing TCP-1. Protein Sci. 10, 445–449 (2001).
Ranson, N.A. et al. ATP-bound states of GroEL captured by cryo-electron microscopy. Cell 107, 869–879 (2001).
Ranson, N.A. et al. Allosteric signaling of ATP hydrolysis in GroEL-GroES complexes. Nat. Struct. Mol. Biol. 13, 147–152 (2006).
Roseman, A.M., Chen, S., White, H., Braig, K. & Saibil, H.R. The chaperonin ATPase cycle: mechanism of allosteric switching and movements of substrate-binding domains in GroEL. Cell 87, 241–251 (1996).
Inbar, E. & Horovitz, A. GroES promotes the T to R transition of the GroEL ring distal to GroES in the GroEL-GroES complex. Biochemistry 36, 12276–12281 (1997).
Kusmierczyk, A.R. & Martin, J. Nucleotide-dependent protein folding in the type II chaperonin from the mesophilic archaeon Methanococcus maripaludis. Biochem. J. 371, 669–673 (2003).
Kusmierczyk, A.R. & Martin, J. Nested cooperativity and salt dependence of the ATPase activity of the archaeal chaperonin Mm-cpn. FEBS Lett. 547, 201–204 (2003).
Szpikowska, B.K., Swiderek, K.M., Sherman, M.A. & Mas, M.T. MgATP binding to the nucleotide-binding domains of the eukaryotic cytoplasmic chaperonin induces conformational changes in the putative substrate-binding domains. Protein Sci. 7, 1524–1530 (1998).
Iizuka, R. et al. Characterization of archaeal group II chaperonin-ADP-metal fluoride complexes: implications that group II chaperonins operate as a “two-stroke engine”. J. Biol. Chem. 280, 40375–40383 (2005).
Llorca, O. et al. 3D reconstruction of the ATP-bound form of CCT reveals the asymmetric folding conformation of a type II chaperonin. Nat. Struct. Biol. 6, 639–642 (1999).
Schoehn, G., Hayes, M., Cliff, M., Clarke, A.R. & Saibil, H.R. Domain rotations between open, closed and bullet-shaped forms of the thermosome, an archaeal chaperonin. J. Mol. Biol. 301, 323–332 (2000).
Schoehn, G., Quaite-Randall, E., Jimenez, J.L., Joachimiak, A. & Saibil, H.R. Three conformations of an archaeal chaperonin, TF55 from Sulfolobus shibatae. J. Mol. Biol. 296, 813–819 (2000).
Rivenzon-Segal, D., Wolf, S.G., Shimon, L., Willison, K.R. & Horovitz, A. Sequential ATP-induced allosteric transitions of the cytoplasmic chaperonin containing TCP-1 revealed by EM analysis. Nat. Struct. Mol. Biol. 12, 233–237 (2005).
Spiess, C., Miller, E.J., McClellan, A.J. & Frydman, J. Identification of the TRiC/CCT substrate binding sites uncovers the function of subunit diversity in eukaryotic chaperonins. Mol. Cell 24, 25–37 (2006).
Aharoni, A. & Horovitz, A. Inter-ring communication is disrupted in the GroEL mutant Arg13 → Gly; Ala126 → Val with known crystal structure. J. Mol. Biol. 258, 732–735 (1996).
Sewell, B.T. et al. A mutant chaperonin with rearranged inter-ring electrostatic contacts and temperature-sensitive dissociation. Nat. Struct. Mol. Biol. 11, 1128–1133 (2004).
Tian, G., Vainberg, I.E., Tap, W.D., Lewis, S.A. & Cowan, N.J. Specificity in chaperonin-mediated protein folding. Nature 375, 250–253 (1995).
Feldman, D.E., Spiess, C., Howard, D.E. & Frydman, J. Tumorigenic mutations in VHL disrupt folding in vivo by interfering with chaperonin binding. Mol. Cell 12, 1213–1224 (2003).
Weber, F. & Hayer-Hartl, M. Refolding of bovine mitochondrial rhodanese by chaperonins GroEL and GroES. Methods Mol. Biol. 140, 117–126 (2000).
Frydman, J. & Hartl, F.U. Principles of chaperone-assisted protein folding: differences between in vitro and in vivo mechanisms. Science 272, 1497–1502 (1996).
Weissman, J.S., Kashi, Y., Fenton, W.A. & Horwich, A.L. GroEL-mediated protein folding proceeds by multiple rounds of binding and release of nonnative forms. Cell 78, 693–702 (1994).
Schwede, T., Kopp, J., Guex, N. & Peitsch, M.C. SWISS-MODEL: an automated protein homology-modeling server. Nucleic Acids Res. 31, 3381–3385 (2003).
Acknowledgements
We thank M. Thanbichler, R. Andino and members of the Frydman laboratory for discussions and comments on the manuscript. S.R. thanks A. Böck for advice and discussions. J. Leigh (University of Washington) kindly provided M. maripaludis DNA. This research was supported by the US National Institutes of Health, the US National Institutes of Health Roadmap Initiative on Nanomedicine, the Studienstiftung des Deutschen Volkes and the Robert Welch Foundation.
Author information
Authors and Affiliations
Contributions
J.F. and S.R. designed the project and experiments; J.F. was the project leader and S.R. carried out all experiments; C.P. and S.R. carried out the data fitting; C.R.B. and W.C. carried out EM analysis of the complexes; S.R. and J.F. wrote the manuscript. All authors made contributions to the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
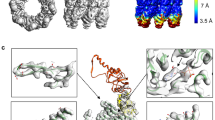
Cryo-EM analysis of cTRiC. (PDF 173 kb)
Supplementary Fig. 2
Direct comparison of first allosteric transition. (PDF 145 kb)
Rights and permissions
About this article
Cite this article
Reissmann, S., Parnot, C., Booth, C. et al. Essential function of the built-in lid in the allosteric regulation of eukaryotic and archaeal chaperonins. Nat Struct Mol Biol 14, 432–440 (2007). https://doi.org/10.1038/nsmb1236
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1236
This article is cited by
-
REP-X: An Evolution-guided Strategy for the Rational Design of Cysteine-less Protein Variants
Scientific Reports (2020)
-
Co-expression of CCT subunits hints at TRiC assembly
Cell Stress and Chaperones (2019)
-
TRiC controls transcription resumption after UV damage by regulating Cockayne syndrome protein A
Nature Communications (2018)
-
Structural and mechanistic characterization of an archaeal-like chaperonin from a thermophilic bacterium
Nature Communications (2017)
-
An information theoretic framework reveals a tunable allosteric network in group II chaperonins
Nature Structural & Molecular Biology (2017)