Abstract
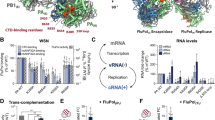
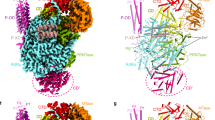
The trimeric influenza virus polymerase, comprising subunits PA, PB1 and PB2, is responsible for transcription and replication of the segmented viral RNA genome. Using a novel library-based screening technique called expression of soluble proteins by random incremental truncation (ESPRIT), we identified an independently folded C-terminal domain from PB2 and determined its solution structure by NMR. Using green fluorescent protein fusions, we show that both the domain and the full-length PB2 subunit are efficiently imported into the nucleus dependent on a previously overlooked bipartite nuclear localization sequence (NLS). The crystal structure of the domain complexed with human importin α5 shows how the last 20 residues unfold to permit binding to the import factor. The domain contains three surface residues implicated in adaptation from avian to mammalian hosts. One of these tethers the NLS-containing peptide to the core of the domain in the unbound state.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Noah, D.L. & Krug, R.M. Influenza virus virulence and its molecular determinants. Adv. Virus Res. 65, 121–145 (2005).
Jung, T.E. & Brownlee, G.G. A new promoter-binding site in the PB1 subunit of the influenza A virus polymerase. J. Gen. Virol. 87, 679–688 (2006).
Li, M.L., Ramirez, B.C. & Krug, R.M. RNA-dependent activation of primer RNA production by influenza virus polymerase: different regions of the same protein subunit constitute the two required RNA-binding sites. EMBO J. 17, 5844–5852 (1998).
Gonzalez, S. & Ortin, J. Characterization of influenza virus PB1 protein binding to viral RNA: two separate regions of the protein contribute to the interaction domain. J. Virol. 73, 631–637 (1999).
Li, M.L., Rao, P. & Krug, R.M. The active sites of the influenza cap-dependent endonuclease are on different polymerase subunits. EMBO J. 20, 2078–2086 (2001).
Honda, A., Mizumoto, K. & Ishihama, A. Two separate sequences of PB2 subunit constitute the RNA cap-binding site of influenza virus RNA polymerase. Genes Cells 4, 475–485 (1999).
Fechter, P. et al. Two aromatic residues in the PB2 subunit of influenza A RNA polymerase are crucial for cap binding. J. Biol. Chem. 278, 20381–20388 (2003).
Nath, S.T. & Nayak, D.P. Function of two discrete regions is required for nuclear localization of polymerase basic protein 1 of A/WSN/33 influenza virus (H1 N1). Mol. Cell. Biol. 10, 4139–4145 (1990).
Mukaigawa, J. & Nayak, D.P. Two signals mediate nuclear localization of influenza virus (A/WSN/33) polymerase basic protein 2. J. Virol. 65, 245–253 (1991).
Nieto, A. et al. Nuclear transport of influenza virus polymerase PA protein. Virus Res. 24, 65–75 (1992).
Fodor, E. & Smith, M. The PA subunit is required for efficient nuclear accumulation of the PB1 subunit of the influenza A virus RNA polymerase complex. J. Virol. 78, 9144–9153 (2004).
Deng, T. et al. Role of ran binding protein 5 in nuclear import and assembly of the influenza virus RNA polymerase complex. J. Virol. 80, 11911–11919 (2006).
Naito, T., Momose, F., Kawaguchi, A. & Nagata, K. Involvement of hsp90 in assembly and nuclear import of influenza virus RNA polymerase subunits. J. Virol. 81, 1339–1349 (2007).
Russell, C.J. & Webster, R.G. The genesis of a pandemic influenza virus. Cell 123, 368–371 (2005).
Li, Z. et al. Molecular basis of replication of duck H5N1 influenza viruses in a mammalian mouse model. J. Virol. 79, 12058–12064 (2005).
Taubenberger, J.K. et al. Characterization of the 1918 influenza virus polymerase genes. Nature 437, 889–893 (2005).
Gabriel, G. et al. The viral polymerase mediates adaptation of an avian influenza virus to a mammalian host. Proc. Natl. Acad. Sci. USA 102, 18590–18595 (2005).
Salomon, R. et al. The polymerase complex genes contribute to the high virulence of the human H5N1 influenza virus isolate A/Vietnam/1203/04. J. Exp. Med. 203, 689–697 (2006).
Martin-Benito, J. et al. Three-dimensional reconstruction of a recombinant influenza virus ribonucleoprotein particle. EMBO Rep. 2, 313–317 (2001).
Area, E. et al. 3D structure of the influenza virus polymerase complex: localization of subunit domains. Proc. Natl. Acad. Sci. USA 101, 308–313 (2004).
Ostermeier, M. & Lutz, S. The creation of ITCHY hybrid protein libraries. Methods Mol. Biol. 231, 129–141 (2003).
Beckett, D., Kovaleva, E. & Schatz, P.J. A minimal peptide substrate in biotin holoenzyme synthetase-catalyzed biotinylation. Protein Sci. 8, 921–929 (1999).
Holm, L. & Sander, C. Protein structure comparison by alignment of distance matrices. J. Mol. Biol. 233, 123–138 (1993).
Perales, B., de la Luna, S., Palacios, I. & Ortin, J. Mutational analysis identifies functional domains in the influenza A virus PB2 polymerase subunit. J. Virol. 70, 1678–1686 (1996).
Fontes, M.R., Teh, T., Jans, D., Brinkworth, R.I. & Kobe, B. Structural basis for the specificity of bipartite nuclear localization sequence binding by importin-alpha. J. Biol. Chem. 278, 27981–27987 (2003).
Zacksenhaus, E., Bremner, R., Phillips, R.A. & Gallie, B.L. A bipartite nuclear localization signal in the retinoblastoma gene product and its importance for biological activity. Mol. Cell. Biol. 13, 4588–4599 (1993).
Conti, E. & Kuriyan, J. Crystallographic analysis of the specific yet versatile recognition of distinct nuclear localization signals by karyopherin alpha. Structure 8, 329–338 (2000).
de Jong, M.D. et al. Fatal outcome of human influenza A (H5N1) is associated with high viral load and hypercytokinemia. Nat. Med. 12, 1203–1207 (2006).
de la Luna, S., Martinez, C. & Ortin, J. Molecular cloning and sequencing of influenza virus A/Victoria/3/75 polymerase genes: sequence evolution and prediction of possible functional domains. Virus Res. 13, 143–155 (1989).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Cryst. 26, 795–800 (1993).
Terwilliger, T.C. Maximum-likelihood density modification. Acta Crystallogr. D Biol. Crystallogr. 56, 965–972 (2000).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Acknowledgements
We thank J. Ortin (Centro Nacional de Biotecnologia, CSIC, Madrid) for the pb2 gene, A. Favier (Institut de Biologie Structurale, Grenoble) for NMR scripts and the EMBL Centre for Molecular and Cellular Imaging for suggestions. Screening for crystals was done by the Partnership for Structural Biology high-throughput crystallization facility. We thank the European Synchrotron Radiation Facility and EMBL Joint Structural Biology group for assistance with the synchrotron beamtime and T. Crepin (EMBL, Grenoble) for help with data collection. Partial funding was provided by the European Commission Framework 5 Integrated Project 'Structural Proteomics in Europe' (SPINE, contract QLG-CT-2002-00988).
Author information
Authors and Affiliations
Contributions
D.J.H. conceived the ESPRIT method. D.J.H., F.T. and P.J.M. implemented ESPRIT. D.G. purified wild-type and mutant DPDE for in vitro binding studies and crystallization and made double-labeled protein for NMR. C.M.B., J.B. and J.-P.S. performed the NMR measurements and structural analysis. S.B. purified importin α5 under the supervision of F.B. and cocrystallized it with DPDE. R.W.H.R. and S.C. initiated the influenza polymerase project, and S.C. determined the crystallographic structure. F.T. and N.D. performed nuclear import assays with instrumentation and methodology established by J.E. S.C. and D.J.H. compiled the text, with contributions from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors have applied for a patent on ESPRIT technology.
Supplementary information
Supplementary Fig. 1
Assigned 1H-15N HSQC spectrum of the C-terminal domain of PB2 at 10 °C (PDF 101 kb)
Supplementary Fig. 2
Plot of r.m.s. deviation and S2 generalized order parameters against primary sequence (PDF 61 kb)
Supplementary Fig. 3
Interaction of the PB2 C-terminal domain with human importin α5 (PDF 176 kb)
Supplementary Fig. 4
Electron density for the bipartite NLS in the complex of human importin α5 with influenza PB2 DPDE domain (PDF 162 kb)
Supplementary Fig. 5
Interactions of the bipartite NLS with human importin α5 (PDF 142 kb)
Supplementary Table 1
NMR and refinement statistics for PB2 C-terminal domain (PDF 34 kb)
Rights and permissions
About this article
Cite this article
Tarendeau, F., Boudet, J., Guilligay, D. et al. Structure and nuclear import function of the C-terminal domain of influenza virus polymerase PB2 subunit. Nat Struct Mol Biol 14, 229–233 (2007). https://doi.org/10.1038/nsmb1212
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1212
This article is cited by
-
The ubiquitination landscape of the influenza A virus polymerase
Nature Communications (2023)
-
The influenza virus RNA polymerase as an innate immune agonist and antagonist
Cellular and Molecular Life Sciences (2021)
-
Comparative study of the interactions between fungal transcription factor nuclear localization sequences with mammalian and fungal importin-alpha
Scientific Reports (2020)
-
Molecular basis of host-adaptation interactions between influenza virus polymerase PB2 subunit and ANP32A
Nature Communications (2020)
-
A prion-like domain in ELF3 functions as a thermosensor in Arabidopsis
Nature (2020)