Abstract
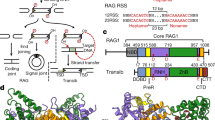
The Rag proteins carry out V(D)J recombination through a process mechanistically similar to cut-and-paste transposition. Specifically, Rag complexes form DNA hairpins through direct transesterification, using a catalytic Asp-Asp-Glu (DDE) triad in Rag1. How is sufficient DNA distortion introduced to allow hairpin formation? We hypothesized that, like certain transposases, the Rag proteins might use aromatic amino acid residues to stabilize a flipped-out base. Through in vivo and in vitro experiments and structural predictions, we identified residues in Rag1 crucial for hairpin formation. One of these, a conserved tryptophan (Trp893), probably participates in base-stacking interactions near the cleavage site, as do Trp298, Trp265 and Trp319 in the Tn5, Tn10 and Hermes transposases, respectively. Other residues surrounding the catalytic glutamate (YKEFRK) may share functional similarities with the YREK motif in IS4 family transposases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Roth, D.B. Restraining the V(D)J recombinase. Nat. Rev. Immunol. 3, 656–666 (2003).
Roth, D.B. & Craig, N.L. VDJ recombination: A transposase goes to work. Cell 94, 411–414 (1998).
Haren, L., Ton-Hoang, B. & Chandler, M. Integrating DNA: transposases and retroviral integrases. Annu. Rev. Microbiol. 53, 245–281 (1999).
Gellert, M. V(D)J recombination: RAG proteins, repair factors, and regulation. Annu. Rev. Biochem. 71, 101–132 (2002).
Roth, D.B., Menetski, J.P., Nakajima, P.B., Bosma, M.J. & Gellert, M. V(D)J recombination: broken DNA molecules with covalently sealed (hairpin) coding ends in scid mouse thymocytes. Cell 70, 983–991 (1992).
McBlane, J.F. et al. Cleavage at a V(D)J recombination signal requires only RAG1 and RAG2 proteins and occurs in two steps. Cell 83, 387–395 (1995).
van Gent, D.C., Mizuuchi, K. & Gellert, M. Similarities between initiation of V(D)J recombination and retroviral integration. Science 271, 1592–1594 (1996).
Agrawal, A., Eastman, Q.M. & Schatz, D.G. Transposition mediated by RAG1 and RAG2 and its implications for the evolution of the immune system. Nature 394, 744–751 (1998).
Hiom, K., Melek, M. & Gellert, M. DNA transposition by the RAG1 and RAG2 proteins: a possible source of oncogenic translocations. Cell 94, 463–470 (1998).
Kennedy, A.K., Guhathakurta, A., Kleckner, N. & Haniford, D.B. Tn10 transposition via a DNA hairpin intermediate. Cell 95, 125–134 (1998).
Bhasin, A., Goryshin, I.Y. & Reznikoff, W.S. Hairpin formation in Tn5 transposition. J. Biol. Chem. 274, 37021–37029 (1999).
Zhou, L. et al. Transposition of hAT elements links transposable elements and V(D)J recombination. Nature 432, 995–1001 (2004).
Bolland, S. & Kleckner, N. The three chemical steps of Tn10/IS10 transposition involve repeated utilization of a single active site. Cell 84, 223–233 (1996).
Davies, D.R., Mahnke Braam, L., Reznikoff, W.S. & Rayment, I. The three-dimensional structure of a Tn5 transposase-related protein determined to 2.9-Å resolution. J. Biol. Chem. 274, 11904–11913 (1999).
Kulkosky, J., Jones, K.S., Katz, R.A., Mack, J.P.G. & Skalka, A.M. Residues critical for retroviral integrative recombination in a region that is highly conserved among retroviral/retrotransposon integrases and bacterial insertion sequence transposases. Mol. Cell. Biol. 12, 2331–2338 (1992).
Grindley, N.D.F. & Leschziner, A.E. DNA transposition: From a black box to a color monitor. Cell 83, 1063–1066 (1995).
Rice, P.A. & Baker, T.A. Comparative architecture of transposase and integrase complexes. Nat. Struct. Biol. 8, 302–307 (2001).
Kim, D.R., Dai, Y., Mundy, C.L., Yang, W. & Oettinger, M.A. Mutations of acidic residues in RAG1 define the active site of the V(D)J recombinase. Genes Dev. 13, 3070–3080 (1999).
Landree, M.A., Wibbenmeyer, J.A. & Roth, D.B. Mutational analysis of RAG-1 and RAG-2 identifies three active site amino acids in RAG-1 critical for both cleavage steps of V(D)J recombination. Genes Dev. 13, 3059–3069 (1999).
Fugmann, S.D., Villey, I.J., Ptaszek, L.M. & Schatz, D.G. Identification of two catalytic residues in RAG1 that define a single active site within the RAG1/RAG2 protein complex. Mol. Cell 5, 97–107 (2000).
Cuomo, C.A., Mundy, C.L. & Oettinger, M.A. DNA sequence and structure requirements for cleavage of V(D)J recombination signal sequences. Mol. Cell. Biol. 16, 5683–5690 (1996).
Ramsden, D.A., McBlane, J.F., van Gent, D.C. & Gellert, M. Distinct DNA sequence and structure requirements for the two steps of V(D)J recombination signal cleavage. EMBO J. 15, 3197–3206 (1996).
Kale, S.B., Landree, M.A. & Roth, D.B. Conditional RAG-1 mutants block the hairpin formation step of V(D)J recombination. Mol. Cell. Biol. 21, 459–466 (2001).
Davies, D.R., Goryshin, I.Y., Reznikoff, W.S. & Rayment, I. Three-dimensional structure of the Tn5 synaptic complex transposition intermediate. Science 289, 77–85 (2000).
Ason, B. & Reznikoff, W.S. Mutational analysis of the base flipping event found in Tn5 transposition. J. Biol. Chem. 277, 11284–11291 (2002).
Allingham, J.S., Wardle, S.J. & Haniford, D.B. Determinants for hairpin formation in Tn10 transposition. EMBO J. 20, 2931–2942 (2001).
Hickman, A.B. et al. Molecular architecture of a eukaryotic DNA transposase. Nat. Struct. Mol. Biol. 12, 715–721 (2005).
Steen, S.B., Gomelsky, L., Speidel, S.L. & Roth, D.B. Initiation of V(D)J recombination in vivo: role of recombination signal sequences in formation of single and paired double-strand breaks. EMBO J. 16, 2656–2664 (1997).
Huye, L.E., Purugganan, M.M., Jiang, M.M. & Roth, D.B. Mutational analysis of all conserved basic amino acids in RAG-1 reveals catalytic, step arrest, and joining-deficient mutants in the V(D)J recombinase. Mol. Cell. Biol. 22, 3460–3473 (2002).
Naumann, T.A. & Reznikoff, W.S. Tn5 transposase active site mutants. J. Biol. Chem. 2771, 17623–17629 (2002).
Zhang, J. et al. Novel RAG1 mutation in a case of severe combined immunodeficiency. Pediatrics 116, e445–e449 (2005).
Mahillon, J., Seurinck, J., van Rompuy, L., Delcour, J. & Zabeau, M. Nucleotide sequence and structural organization of an insertion sequence element (IS231) from Bacillus thuringiensis strain berliner 1715. EMBO J. 4, 3895–3899 (1985).
Rezsohazy, R., Hallet, B., Delcour, J. & Mahillon, J. The IS4 family of insertion sequences: evidence for a conserved transposase motif. Mol. Microbiol. 9, 1283–1295 (1993).
Spanopoulou, E. et al. The homeodomain region of Rag-1 reveals the parallel mechanisms of bacterial and V(D)J recombination. Cell 87, 263–276 (1996).
Deng, W.P. & Nickoloff, J.A. Site-directed mutagenesis of virtually any plasmid by eliminating a unique site. Anal. Biochem. 200, 81–88 (1992).
Han, J.-O., Steen, S.B. & Roth, D.B. Ku86 is not required for protection of signal ends or for formation of nonstandard V(D)J recombination products. Mol. Cell. Biol. 17, 2226–2234 (1997).
Rost, B., Yachdav, G. & Liu, J. The PredictProtein server. Nucleic Acids Res. 32, W321–W326 (2004).
Acknowledgements
The authors thank members of the Roth laboratory for thoughtful discussions and comments on the manuscript and G. Weller for unpublished data. This work was supported by US National Institutes of Health grant AI36420 and funding from the Irene Diamond Foundation (to D.B.R.).
Author information
Authors and Affiliations
Contributions
H.S. began the mutagenesis project while a student in the Roth lab at Baylor College of Medicine. C.P.L. performed all the experiments shown; P.A.R. performed the structural predictions and analysis; C.P.L., V.L.B. and D.B.R. thought about experiments, analyzed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Nicking and hairpin formation by Rag1/Rag2. (PDF 182 kb)
Supplementary Fig. 2
Time-course analysis and prenicked substrate experiments. (PDF 500 kb)
Supplementary Fig. 3
Comparative alignment of Tn5, Tn10 and Rag1. (PDF 184 kb)
Supplementary Fig. 4
Rough alignment of proposed catalytic domains of Hermes and Rag1. (PDF 79 kb)
Supplementary Table 1
Oligonucleotide sequences. (PDF 29 kb)
Rights and permissions
About this article
Cite this article
Lu, C., Sandoval, H., Brandt, V. et al. Amino acid residues in Rag1 crucial for DNA hairpin formation. Nat Struct Mol Biol 13, 1010–1015 (2006). https://doi.org/10.1038/nsmb1154
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1154
This article is cited by
-
Biochemical Characterization of Kat1: a Domesticated hAT-Transposase that Induces DNA Hairpin Formation and MAT-Switching
Scientific Reports (2016)
-
Crystal structure of the V(D)J recombinase RAG1–RAG2
Nature (2015)
-
Structure of the RAG1 nonamer binding domain with DNA reveals a dimer that mediates DNA synapsis
Nature Structural & Molecular Biology (2009)
-
piggyBac can bypass DNA synthesis during cut and paste transposition
The EMBO Journal (2008)