Abstract
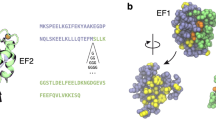
Calbindin-D28K is a Ca2+-binding protein, performing roles as both a calcium buffer and calcium sensor. The NMR solution structure of Ca2+-loaded calbindin-D28K reveals a single, globular fold consisting of six distinct EF-hand subdomains, which coordinate Ca2+ in loops on EF1, EF3, EF4 and EF5. Target peptides from Ran-binding protein M and myo-inositol monophosphatase, along with a new target from procaspase-3, are shown to interact with the protein on a surface comprised of α5 (EF3), α8 (EF4) and the EF2-EF3 and EF4-EF5 loops. Fluorescence experiments reveal that calbindin-D28K adopts discrete hydrophobic states as it binds Ca2+. The structure, binding interface and hydrophobic characteristics of Ca2+-loaded calbindin-D28K provide the first detailed insights into how this essential protein may function. This structure is one of the largest high-resolution NMR structures and the largest monomeric EF-hand protein to be solved to date.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Gross, M. & Kumar, R. Physiology and biochemistry of vitamin D-dependent calcium binding proteins. Am. J. Physiol. 259, F195–F209 (1990).
Oberholtzer, J.C., Buettger, C., Summers, M.C. & Matschinsky, F.M. The 28-kDa calbindin-D is a major calcium-binding protein in the basilar papilla of the chick. Proc. Natl. Acad. Sci. USA 85, 3387–3390 (1988).
Lutz, W. et al. Calbindin D28K interacts with Ran-binding protein M: identification of interacting domains by NMR spectroscopy. Biochem. Biophys. Res. Commun. 303, 1186–1192 (2003).
Berggard, T., Szczepankiewicz, O., Thulin, E. & Linse, S. Myo-inositol monophosphatase is an activated target of calbindin D28K . J. Biol. Chem. 277, 41954–41959 (2002).
Christakos, S. & Liu, Y. Biological actions and mechanism of action of calbindin in the process of apoptosis. J. Steroid Biochem. Mol. Biol. 89–90, 401–404 (2004).
Berggard, T., Thulin, E., Akerfeldt, K.S. & Linse, S. Fragment complementation of calbindin D28K . Protein Sci. 9, 2094–2108 (2000).
Linse, S. et al. Domain organization of calbindin D28K as determined from the association of six synthetic EF-hand fragments. Protein Sci. 6, 2385–2396 (1997).
Bellido, T., Huening, M., Raval-Pandya, M., Manolagas, S.C. & Christakos, S. Calbindin D28K is expressed in osteoblastic cells and suppresses their apoptosis by inhibiting caspase-3 activity. J. Biol. Chem. 275, 26328–26332 (2000).
Rabinovitch, A., Suarez-Pinzon, W.L., Sooy, K., Strynadka, K. & Christakos, S. Expression of calbindin- D28K in a pancreatic islet beta-cell line protects against cytokine-induced apoptosis and necrosis. Endocrinology 142, 3649–3655 (2001).
Liu, Y. et al. Prevention of glucocorticoid-induced apoptosis in osteocytes and osteoblasts by calbindin- D28K . J. Bone Miner. Res. 19, 479–490 (2004).
Christakos, S. et al. Vitamin D target proteins: function and regulation. J. Cell. Biochem. 88, 238–244 (2003).
Shamir, A., Elhadad, N., Belmaker, R.H. & Agam, G. Interaction of calbindin D28K and inositol monophosphatase in human postmortem cortex: possible implications for bipolar disorder. Bipolar Disord. 7, 42–48 (2005).
Schmidt, H., Schwaller, B. & Eilers, J. Calbindin D28K targets myo-inositol monophosphatase in spines and dendrites of cerebellar Purkinje neurons. Proc. Natl. Acad. Sci. USA 102, 5850–5855 (2005).
Venters, R.A. et al. The effects of Ca2+ binding on the conformation of calbindin D28K: a nuclear magnetic resonance and microelectrospray mass spectrometry study. Anal. Biochem. 317, 59–66 (2003).
Morgan, D.W., Welton, A.F., Heick, A.E. & Christakos, S. Specific in vitro activation of Ca,Mg-ATPase by vitamin D-dependent rat renal calcium binding protein (calbindin D28K). Biochem. Biophys. Res. Commun. 138, 547–553 (1986).
Reisner, P.D., Christakos, S. & Vanaman, T.C. In vitro enzyme activation with calbindin- D28K, the vitamin D-dependent 28 kDa calcium binding protein. FEBS Lett. 297, 127–131 (1992).
Ikura, M. Calcium binding and conformational response in EF-hand proteins. Trends Biochem. Sci. 21, 14–17 (1996).
Nakamura, M. et al. When overexpressed, a novel centrosomal protein, RanBPM, causes ectopic microtubule nucleation similar to gamma-tubulin. J. Cell Biol. 143, 1041–1052 (1998).
Nishimoto, T. A new role of ran GTPase. Biochem. Biophys. Res. Commun. 262, 571–574 (1999).
Nishitani, H. et al. Full-sized RanBPM cDNA encodes a protein possessing a long stretch of proline and glutamine within the N-terminal region, comprising a large protein complex. Gene 272, 25–33 (2001).
Rao, M.A. et al. RanBPM, a nuclear protein that interacts with and regulates transcriptional activity of androgen receptor and glucocorticoid receptor. J. Biol. Chem. 277, 48020–48027 (2002).
Seki, T., Hayashi, N. & Nishimoto, T. RCC1 in the Ran pathway. J. Biochem. 120, 207–214 (1996).
Moore, M.S. Generation of GTP-Ran for nuclear protein import. Science 272, 47 (1996).
Cotman, C.W., Poon, W.W., Rissman, R.A. & Blurton-Jones, M. The role of caspase cleavage of tau in Alzheimer disease neuropathology. J. Neuropathol. Exp. Neurol. 64, 104–112 (2005).
Hodges, A. et al. Regional and cellular gene expression changes in human Huntington's disease brain. Hum. Mol. Genet. 15, 965–977 (2006).
Wellington, C.L. et al. Inhibiting caspase cleavage of huntingtin reduces toxicity and aggregate formation in neuronal and nonneuronal cells. J. Biol. Chem. 275, 19831–19838 (2000).
Yuan, J. & Yankner, B.A. Apoptosis in the nervous system. Nature 407, 802–809 (2000).
Lu, T. et al. Gene regulation and DNA damage in the ageing human brain. Nature 429, 883–891 (2004).
Colin, E. et al. Akt is altered in an animal model of Huntington's disease and in patients. Eur. J. Neurosci. 21, 1478–1488 (2005).
Berggard, T., Silow, M., Thulin, E. & Linse, S. Ca2+- and H+-dependent conformational changes of calbindin D28K . Biochemistry 39, 6864–6873 (2000).
Cedervall, T. et al. Calbindin D28K EF-hand ligand binding and oligomerization: four high-affinity sites-three modes of action. Biochemistry 44, 13522–13532 (2005).
Berggard, T. et al. Calbindin D28K exhibits properties characteristic of a Ca2+ sensor. J. Biol. Chem. 277, 16662–16672 (2002).
Veenstra, T.D., Johnson, K.L., Tomlinson, A.J., Naylor, S. & Kumar, R. Determination of calcium-binding sites in rat brain calbindin D28K by electrospray ionization mass spectrometry. Biochemistry 36, 3535–3542 (1997).
Vanbelle, C. et al. Deamidation and disulfide bridge formation in human calbindin D28K with effects on calcium binding. Protein Sci. 14, 968–979 (2005).
Cedervall, T. et al. Redox sensitive cysteine residues in calbindin D28K are structurally and functionally important. Biochemistry 44, 684–693 (2005).
Johnson, K.L. et al. On-line sample clean-up and chromatography coupled with electrospray ionization mass spectrometry to characterize the primary sequence and disulfide bond content of recombinant calcium binding proteins. Biomed. Chromatogr. 13, 37–45 (1999).
Tao, L., Murphy, M.E. & English, A.M. S-nitrosation of Ca2+-loaded and Ca2+-free recombinant calbindin D28K from human brain. Biochemistry 41, 6185–6192 (2002).
Nelson, M.R. & Chazin, W.J. Structures of EF-hand Ca2+-binding proteins: diversity in the organization, packing and response to Ca2+ binding. Biometals 11, 297–318 (1998).
Klaus, W. et al. NMR investigation and secondary structure of domains I and II of rat brain calbindin D28K (1–93). Eur. J. Biochem. 262, 933–938 (1999).
Mueller, G.A. et al. Global folds of proteins with low densities of NOEs using residual dipolar couplings: application to the 370-residue maltodextrin-binding protein. J. Mol. Biol. 300, 197–212 (2000).
Atreya, H.S. & Chary, K.V.R. New chemical shift signatures of bound calcium in EF-hand proteins. Curr. Sci. [online] 83, 1240–1245 (2002).
Biekofsky, R.R., Turjanski, A.G., Estrin, D.A., Feeney, J. & Pastore, A. Ab initio study of NMR 15N chemical shift differences induced by Ca2+ binding to EF-hand proteins. Biochemistry 43, 6554–6564 (2004).
Linge, J.P., Williams, M.A., Spronk, C.A., Bonvin, A.M. & Nilges, M. Refinement of protein structures in explicit solvent. Proteins 50, 496–506 (2003).
Rigden, D.J. & Galperin, M.Y. The DxDxDG motif for calcium binding: multiple structural contexts and implications for evolution. J. Mol. Biol. 343, 971–984 (2004).
Akerfeldt, K.S., Coyne, A.N., Wilk, R.R., Thulin, E. & Linse, S. Ca2+-binding stoichiometry of calbindin D28K as assessed by spectroscopic analyses of synthetic peptide fragments. Biochemistry 35, 3662–3669 (1996).
Donepudi, M. & Grutter, M.G. Structure and zymogen activation of caspases. Biophys. Chem. 101–102, 145–153 (2002).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Johnson, B.A. & Blevins, R.A. NMRView—a computer program for the visualization and analysis of NMR data. J. Biomol. NMR 4, 603–614 (1994).
Helgstrand, M., Vanbelle, C., Thulin, E., Linse, S. & Akke, M. Sequential 1H, 15N and 13C NMR assignment of human calbindin D28K . J. Biomol. NMR 28, 305–306 (2004).
Cornilescu, G., Delaglio, F. & Bax, A. Protein backbone angle restraints from searching a database for chemical shift and sequence homology. J. Biomol. NMR 13, 289–302 (1999).
Yang, D., Venters, R.A., Mueller, G.A., Choy, W.Y. & Kay, L.E. TROSY-based HNCO pulse sequences for the measurement of 1HN-15N, 15N-13CO, 1HN-13CO, 13CO-13Cα and 1HN-13Cα dipolar couplings in 15N, 13C, 2H-labeled proteins. J. Biomol. NMR 14, 333–343 (1999).
Wang, Y.X. et al. Measurement of 3hJNC' connectivities across hydrogen bonds in a 30 kDa protein. J. Biomol. NMR 14, 181–184 (1999).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Stein, E.G., Rice, L.M. & Brunger, A.T. Torsion-angle molecular dynamics as a new efficient tool for NMR structure calculation. J. Magn. Reson. 124, 154–164 (1997).
Choy, W.Y., Tollinger, M., Mueller, G.A. & Kay, L.E. Direct structure refinement of high molecular weight proteins against residual dipolar couplings and carbonyl chemical shift changes upon alignment: an application to maltose binding protein. J. Biomol. NMR 21, 31–40 (2001).
Nabuurs, S.B. et al. Quantitative evaluation of experimental NMR restraints. J. Am. Chem. Soc. 125, 12026–12034 (2003).
Laskowski, R.A., Rullmannn, J.A., MacArthur, M.W., Kaptein, R. & Thornton, J.M. AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J. Biomol. NMR 8, 477–486 (1996).
Davis, I.W., Murray, L.W., Richardson, J.S. & Richardson, D.C. MOLPROBITY: structure validation and all-atom contact analysis for nucleic acids and their complexes. Nucleic Acids Res. 32, W615–W619 (2004).
Gouet, P., Robert, X. & Courcelle, E. ESPript/ENDscript: extracting and rendering sequence and 3D information from atomic structures of proteins. Nucleic Acids Res. 31, 3320–3323 (2003).
Acknowledgements
The Duke University NMR Center and the North Carolina State University NMR Facility were established with grants from the US National Institutes of Health, US National Science Foundation and North Carolina Biotechnology Center. We acknowledge support of this work by grants from the US National Institutes of Health (to R.K. and J.C.), the Kenan Institute for Engineering, Technology & Science (to J.C.) and the American Foundation for Aging Research (to D.J.K and D.R.K.).
Author information
Authors and Affiliations
Contributions
D.J.K. contributed to assignments, structure calculations and manuscript preparation; R.A.V. to assignments, structure calculations, NMR data collection and analysis, and manuscript preparation; D.R.K. to sample preparations, fluorescence experiments and peptide-binding titrations; R.J.T. to sample preparations and NMR data collection; R.K. to RanBPM studies; and J.C. to NMR data collection and analysis, peptide-binding titrations and manuscript preparation.
Note: Supplementary information is available on the Nature Structural & Molecular Biology website.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Secondary structure and NOE statistics form Ca2+-loaded calbindin-D28K (PDF 5832 kb)
Supplementary Fig. 2
Measuring hydrophobic surface content via ANS fluorescence as calbindin-D28K binds Ca2+ (PDF 3759 kb)
Supplementary Table 1
NMR experiments used in the analysis of Ca2+-loaded calbindin-D28K (PDF 86 kb)
Rights and permissions
About this article
Cite this article
Kojetin, D., Venters, R., Kordys, D. et al. Structure, binding interface and hydrophobic transitions of Ca2+-loaded calbindin-D28K. Nat Struct Mol Biol 13, 641–647 (2006). https://doi.org/10.1038/nsmb1112
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1112
This article is cited by
-
The root transcriptome analyses of peanut wild species Arachis correntina (Burkart) Krapov. & W.C. Gregory and cultivated variety Xiaobaisha in response to benzoic acid and p-cumaric acid stress
Genetic Resources and Crop Evolution (2020)
-
Expression and localization of the polarity protein CRB2 in adult mouse brain: a comparison with the CRB1rd8 mutant mouse model
Scientific Reports (2018)
-
Solving the crystal structure of human calcium-free S100Z: the siege and conquer of one of the last S100 family strongholds
JBIC Journal of Biological Inorganic Chemistry (2017)
-
Solution structure and dynamics of human S100A14
JBIC Journal of Biological Inorganic Chemistry (2013)
-
Location, location, location: Genetic regulation of neural sex differences
Reviews in Endocrine and Metabolic Disorders (2012)