Abstract
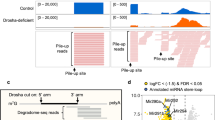
Adenosine deaminases acting on RNA (ADARs) are involved in editing of adenosine residues to inosine in double-stranded RNA (dsRNA). Although this editing recodes and alters functions of several mammalian genes, its most common targets are noncoding repeat sequences, indicating the involvement of this editing system in currently unknown functions other than recoding of protein sequences. Here we show that specific adenosine residues of certain microRNA (miRNA) precursors are edited by ADAR1 and ADAR2. Editing of pri–miR-142, the precursor of miRNA-142, expressed in hematopoietic tissues, resulted in suppression of its processing by Drosha. The edited pri–miR-142 was degraded by Tudor-SN, a component of RISC and also a ribonuclease specific to inosine-containing dsRNAs. Consequently, mature miRNA-142 expression levels increased substantially in ADAR1 null or ADAR2 null mice. Our results demonstrate a new function of RNA editing in the control of miRNA biogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Bass, B.L. & Weintraub, H. An unwinding activity that covalently modifies its double-stranded RNA substrate. Cell 55, 1089–1098 (1988).
Wagner, R.W., Smith, J.E., Cooperman, B.S. & Nishikura, K. A double-stranded RNA unwinding activity introduces structural alterations by means of adenosine to inosine conversions in mammalian cells and Xenopus eggs. Proc. Natl. Acad. Sci. USA 86, 2647–2651 (1989).
Masquida, B. & Westhof, E. On the wobble GoU and related pairs. RNA 6, 9–15 (2000).
Bass, B.L. RNA editing by adenosine deaminases that act on RNA. Annu. Rev. Biochem. 71, 817–846 (2002).
Keegan, L.P., Gallo, A. & O'Connell, M.A. The many roles of an RNA editor. Nat. Rev. Genet. 2, 869–878 (2001).
Patterson, J.B. & Samuel, C.E. Expression and regulation by interferon of a double-stranded-RNA-specific adenosine deaminase from human cells: evidence for two forms of the deaminase. Mol. Cell. Biol. 15, 5376–5388 (1995).
Kim, U., Wang, Y., Sanford, T., Zeng, Y. & Nishikura, K. Molecular cloning of cDNA for double-stranded RNA adenosine deaminase, a candidate enzyme for nuclear RNA editing. Proc. Natl. Acad. Sci. USA 91, 11457–11461 (1994).
Lai, F., Drakas, R. & Nishikura, K. Mutagenic analysis of double-stranded RNA adenosine deaminase, a candidate enzyme for RNA editing of glutamate-gated ion channel transcripts. J. Biol. Chem. 270, 17098–17105 (1995).
Melcher, T. et al. RED2, a brain-specific member of the RNA-specific adenosine deaminase family. J. Biol. Chem. 271, 31795–31798 (1996).
Melcher, T. et al. A mammalian RNA editing enzyme. Nature 379, 460–464 (1996).
Burns, C.M. et al. Regulation of serotonin-2C receptor G-protein coupling by RNA editing. Nature 387, 303–308 (1997).
Higuchi, M. et al. RNA editing of AMPA receptor subunit GluR-B: a base-paired intron-exon structure determines position and efficiency. Cell 75, 1361–1370 (1993).
Hoopengardner, B., Bhalla, T., Staber, C. & Reenan, R. Nervous system targets of RNA editing identified by comparative genomics. Science 301, 832–836 (2003).
Lomeli, H. et al. Control of kinetic properties of AMPA receptor channels by nuclear RNA editing. Science 266, 1709–1713 (1994).
Hartner, J.C. et al. Liver disintegration in the mouse embryo caused by deficiency in the RNA-editing enzyme ADAR1. J. Biol. Chem. 279, 4894–4902 (2004).
Wang, Q. et al. Stress-induced apoptosis associated with null mutation of ADAR1 RNA editing deaminase gene. J. Biol. Chem. 279, 4952–4961 (2004).
Cho, D.S. et al. Requirement of dimerization for RNA editing activity of adenosine deaminases acting on RNA. J. Biol. Chem. 278, 17093–17102 (2003).
Athanasiadis, A., Rich, A. & Maas, S. Widespread A → I RNA editing of Alu-containing mRNAs in the human transcriptome. PLoS Biol. 2, e391 (2004).
Kim, D.D. et al. Widespread RNA editing of embedded alu elements in the human transcriptome. Genome Res. 14, 1719–1725 (2004).
Levanon, E.Y. et al. Systematic identification of abundant A → I editing sites in the human transcriptome. Nat. Biotechnol. 22, 1001–1005 (2004).
Nishikura, K. Editing the message from A to I. Nat. Biotechnol. 22, 962–963 (2004).
Lagos-Quintana, M., Rauhut, R., Meyer, J., Borkhardt, A. & Tuschl, T. New microRNAs from mouse and human. RNA 9, 175–179 (2003).
Lim, L.P., Glasner, M.E., Yekta, S., Burge, C.B. & Bartel, D.P. Vertebrate microRNA genes. Science 299, 1540 (2003).
Pfeffer, S. et al. Identification of microRNAs of the herpesvirus family. Nat. Methods 2, 269–276 (2005).
Lee, Y. et al. The nuclear RNase III Drosha initiates microRNA processing. Nature 425, 415–419 (2003).
Zeng, Y., Yi, R. & Cullen, B.R. Recognition and cleavage of primary microRNA precursors by the nuclear processing enzyme Drosha. EMBO J. 24, 138–148 (2005).
Denli, A.M., Tops, B.B., Plasterk, R.H., Ketting, R.F. & Hannon, G.J. Processing of primary microRNAs by the Microprocessor complex. Nature 432, 231–235 (2004).
Gregory, R.I. et al. The Microprocessor complex mediates the genesis of microRNAs. Nature 432, 235–240 (2004).
Han, J. et al. The Drosha-DGCR8 complex in primary microRNA processing. Genes Dev. 18, 3016–3027 (2004).
Landthaler, M., Yalcin, A. & Tuschl, T. The human DiGeorge syndrome critical region gene 8 and Its D. melanogaster homolog are required for miRNA biogenesis. Curr. Biol. 14, 2162–2167 (2004).
Lund, E., Guttinger, S., Calado, A., Dahlberg, J.E. & Kutay, U. Nuclear export of microRNA precursors. Science 303, 95–98 (2004).
Chendrimada, T.P. et al. TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature 436, 740–744 (2005).
Förstemann, K. et al. Normal microRNA maturation and germ-line stem cell maintenance requires Loquacious, a double-stranded RNA-binding domain protein. PLoS Biol. 3, e236 (2005).
Ambros, V. The functions of animal microRNAs. Nature 431, 350–355 (2004).
Bartel, D.P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Wienholds, E. & Plasterk, R.H. MicroRNA function in animal development. FEBS Lett. 579, 5911–5922 (2005).
Chen, C.Z., Li, L., Lodish, H.F. & Bartel, D.P. MicroRNAs modulate hematopoietic lineage differentiation. Science 303, 83–86 (2004).
Yang, W. et al. ADAR1 RNA deaminase limits short interfering RNA efficacy in mammalian cells. J. Biol. Chem. 280, 3946–3953 (2005).
Scadden, A.D. & O'Connell, M.A. Cleavage of dsRNAs hyper-edited by ADARs occurs at preferred editing sites. Nucleic Acids Res. 22, 5954–5964 (2005).
Caudy, A.A. et al. A micrococcal nuclease homologue in RNAi effector complexes. Nature 425, 411–414 (2003).
Scadden, A.D. The RISC subunit Tudor-SN binds to hyper-edited double-stranded RNA and promotes its cleavage. Nat. Struct. Mol. Biol. 12, 489–496 (2005).
Higuchi, M. et al. Point mutation in an AMPA receptor gene rescues lethality in mice deficient in the RNA-editing enzyme ADAR2. Nature 406, 78–81 (2000).
Luciano, D.J., Mirsky, H., Vendetti, N.J. & Maas, S. RNA editing of a miRNA precursor. RNA 10, 1174–1177 (2004).
Zeng, Y. & Cullen, B.R. Sequence requirements for micro RNA processing and function in human cells. RNA 9, 112–123 (2003).
Griffiths-Jones, S. The microRNA Registry. Nucleic Acids Res. 32, D109–D111 (2004).
Bass, B.L. Double-stranded RNA as a template for gene silencing. Cell 101, 235–238 (2000).
Scadden, A.D. & Smith, C.W. RNAi is antagonized by A → I hyper-editing. EMBO Rep. 2, 1107–1111 (2001).
Tonkin, L.A. & Bass, B.L. Mutations in RNAi rescue aberrant chemotaxis of ADAR mutants. Science 302, 1725 (2003).
Dabiri, G.A., Lai, F., Drakas, R.A. & Nishikura, K. Editing of the GLuR-B ion channel RNA in vitro by recombinant double-stranded RNA adenosine deaminase. EMBO J. 15, 34–45 (1996).
Rickert, R.C., Rajewsky, K. & Roes, J. Impairment of T-cell-dependent B-cell responses and B-1 cell development in CD19-deficient mice. Nature 376, 352–355 (1995).
Acknowledgements
We thank J.M. Murray for critical reading of the manuscript and J.T. Lee for excellent technical assistance. This work was supported in part by grants from the US National Institutes of Health, the March of Dimes and the Commonwealth Universal Research Enhancement Program of the Pennsylvania Department of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Differential detection of unedited and edited pri-miR-142 RNAs by the primer extension assay. (PDF 341 kb)
Supplementary Fig. 2
Detection of endogenous and recombinant ADAR and miRNA processing enzymes in transfected HEK293 cells. (PDF 465 kb)
Supplementary Fig. 3
Preparation of mammalian TSN proteins. (PDF 130 kb)
Rights and permissions
About this article
Cite this article
Yang, W., Chendrimada, T., Wang, Q. et al. Modulation of microRNA processing and expression through RNA editing by ADAR deaminases. Nat Struct Mol Biol 13, 13–21 (2006). https://doi.org/10.1038/nsmb1041
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1041
This article is cited by
-
ADAR1 polymorphisms contribute to increased susceptibility in pediatric acute lymphoblastic leukemia
Annals of Hematology (2023)
-
RNA Editing Therapeutics: Advances, Challenges and Perspectives on Combating Heart Disease
Cardiovascular Drugs and Therapy (2023)
-
ADAR1p110 promotes Enterovirus D68 replication through its deaminase domain and inhibition of PKR pathway
Virology Journal (2022)
-
Change of Heart: the Epitranscriptome of Small Non-coding RNAs in Heart Failure
Current Heart Failure Reports (2022)
-
Direct identification of A-to-I editing sites with nanopore native RNA sequencing
Nature Methods (2022)