Abstract
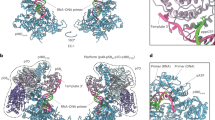
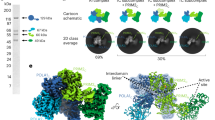
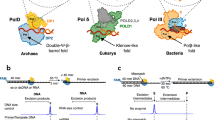
Primases are DNA-dependent RNA polymerases that synthesize the oligoribonucleotide primers essential to DNA replication. In archaeal and eukaryotic organisms, the core primase is a heterodimeric enzyme composed of a small and a large subunit. Here we report a crystallographic and biochemical analysis of the core primase from the archaeon Sulfolobus solfataricus. The structure provides the first three-dimensional description of the large subunit and its interaction with the small subunit. The evolutionary conservation of amino acids at the protein-protein interface implies that the observed mode of subunit association is conserved among archaeal and eukaryotic primases. The orientation of the large subunit in the core primase probably excludes its direct involvement in catalysis. Modeling of a DNA-RNA helix together with structure-based site-directed mutagenesis provides insight into the mechanism of template DNA binding and RNA primer synthesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Kornberg, A. & Baker, T.A. DNA Replication 2nd edn (W.H. Freeman and Company, New York, 1992).
Frick, D.N. & Richardson, C.C. DNA primases. Annu. Rev. Biochem. 70, 39–80 (2001).
Iyer, L.M., Koonin, E.V., Leipe, D.D. & Aravind, L. Origin and evolution of the archaeo-eukaryotic primase superfamily and related palm-domain proteins: structural insights and new members. Nucleic Acids Res. 33, 3875–3896 (2005).
Foiani, M., Lucchini, G. & Plevani, P. The DNA polymerase α-primase complex couples DNA replication, cell-cycle progression and DNA-damage response. Trends Biochem. Sci. 22, 424–427 (1997).
Arezi, B. & Kuchta, R.D. Eukaryotic DNA primase. Trends Biochem. Sci. 25, 572–576 (2000).
Santocanale, C., Foiani, M., Lucchini, G. & Plevani, P. The isolated 48,000-dalton subunit of yeast DNA primase is sufficient for RNA primer synthesis. J. Biol. Chem. 268, 1343–1348 (1993).
Copeland, W.C. & Wang, T.S. Enzymatic characterization of the individual mammalian primase subunits reveals a biphasic mechanism for initiation of DNA replication. J. Biol. Chem. 268, 26179–26189 (1993).
Schneider, A. et al. Primase activity of human DNA polymerase α-primase. Divalent cations stabilize the enzyme activity of the p48 subunit. J. Biol. Chem. 273, 21608–21615 (1998).
Foiani, M., Santocanale, C., Plevani, P. & Lucchini, G. A single essential gene, PRI2, encodes the large subunit of DNA primase in Saccharomyces cerevisiae. Mol. Cell. Biol. 9, 3081–3087 (1989).
Arezi, B., Kirk, B.W., Copeland, W.C. & Kuchta, R.D. Interactions of DNA with human DNA primase monitored with photoactivatable crosslinking agents: implications for the role of the p58 subunit. Biochemistry 38, 12899–12907 (1999).
Zerbe, L.K. & Kuchta, R.D. The p58 subunit of human DNA primase is important for primer initiation, elongation, and counting. Biochemistry 41, 4891–4900 (2002).
Edgell, D.R. & Doolittle, W.F. Archaea and the origin(s) of DNA replication proteins. Cell 89, 995–998 (1997).
Dionne, I. et al. DNA replication in the hyperthermophilic archaeon Sulfolobus solfataricus. Biochem. Soc. Trans. 31, 674–676 (2003).
Grabowski, B. & Kelman, Z. Archeal DNA replication: eukaryal proteins in a bacterial context. Annu. Rev. Microbiol. 57, 487–516 (2003).
Desogus, G., Onesti, S., Brick, P., Rossi, M. & Pisani, F.M. Identification and characterization of a DNA primase from the hyperthermophilic archaeon Methanococcus jannaschii. Nucleic Acids Res. 27, 4444–4450 (1999).
Makarova, K.S. et al. Comparative genomics of the archaea (euryarchaeota): evolution of conserved protein families, the stable core, and the variable shell. Genome Res. 9, 608–628 (1999).
Bocquier, A.A. et al. Archaeal primase: bridging the gap between RNA and DNA polymerases. Curr. Biol. 11, 452–456 (2001).
Liu, L. et al. The archaeal DNA primase: biochemical characterization of the p41-p46 complex from Pyrococcus furiosus. J. Biol. Chem. 276, 45484–45490 (2001).
Lao-Sirieix, S.H. & Bell, S.D. The heterodimeric primase of the hyperthermophilic archaeon Sulfolobus solfataricus possesses DNA and RNA primase, polymerase and 3′-terminal nucleotidyl transferase activities. J. Mol. Biol. 344, 1251–1263 (2004).
Augustin, M.A., Huber, R. & Kaiser, J.T. Crystal structure of a DNA-dependent RNA polymerase (DNA primase). Nat. Struct. Biol. 8, 57–61 (2001).
Ito, N., Nureki, O., Shirouzu, M., Yokoyama, S. & Hanaoka, F. Crystal structure of the Pyrococcus horikoshii DNA primase-UTP complex: implications for the mechanism of primer synthesis. Genes Cells 8, 913–923 (2003).
Steitz, T.A., Smerdon, S.J., Jager, J. & Joyce, C.M. A unified polymerase mechanism for nonhomologous DNA and RNA polymerases. Science 266, 2022–2025 (1994).
Lipps, G., Weinzierl, A.O., von Scheven, G., Buchen, C. & Cramer, P. Structure of a bifunctional DNA primase-polymerase. Nat. Struct. Mol. Biol. 11, 157–162 (2004).
Marini, F. et al. A role for DNA primase in coupling DNA replication to DNA damage response. EMBO J. 16, 639–650 (1997).
Copeland, W.C. & Tan, X. Active site mapping of the catalytic mouse primase subunit by alanine scanning mutagenesis. J. Biol. Chem. 270, 3905–3913 (1995).
Holm, L. & Sander, C. Dali: a network tool for protein structure comparison. Trends Biochem. Sci. 20, 478–480 (1995).
Matsui, E. et al. Distinct domain functions regulating de novo DNA synthesis of thermostable DNA primase from hyperthermophile Pyrococcus horikoshii. Biochemistry 42, 14968–14976 (2003).
Glaser, F. et al. ConSurf: identification of functional regions in proteins by surface-mapping of phylogenetic information. Bioinformatics 19, 163–164 (2003).
Pan, H. & Wigley, D.B. Structure of the zinc-binding domain of Bacillus stearothermophilus DNA primase. Structure Fold. Des. 8, 231–239 (2000).
Kato, M., Ito, T., Wagner, G. & Ellenberger, T. A molecular handoff between bacteriophage T7 DNA primase and T7 DNA polymerase initiates DNA synthesis. J. Biol. Chem. 279, 30554–30562 (2004).
Lu, X.J. & Olson, W.K. 3DNA: a software package for the analysis, rebuilding and visualization of three-dimensional nucleic acid structures. Nucleic Acids Res. 31, 5108–5121 (2003).
Horton, N.C. & Finzel, B.C. The structure of an RNA/DNA hybrid: a substrate of the ribonuclease activity of HIV-1 reverse transcriptase. J. Mol. Biol. 264, 521–533 (1996).
Pelletier, H., Sawaya, M.R., Kumar, A., Wilson, S.H. & Kraut, J. Structures of ternary complexes of rat DNA polymerase β, a DNA template-primer, and ddCTP. Science 264, 1891–1903 (1994).
Sheaff, R.J. & Kuchta, R.D. Mechanism of calf thymus DNA primase: slow initiation, rapid polymerization, and intelligent termination. Biochemistry 32, 3027–3037 (1993).
Weeks, C.M. & Miller, R. The design and implementation of SnB version 2.0. J. Appl. Crystallogr. 32, 120–124 (1999).
La Fortelle, E.D. & Bricogne, G. Maximum-likelihood heavy-atom parameter refinement for multiple isomorphous replacement and multiwavelength anomalous diffraction methods. Methods Enzymol. 276, 472–494 (1997).
Emsley, P. & Cowtan, K. Coot: Model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Blanc, E., Roversi, P., Vonrhein, C., Flensburg, S.M. & Bricogne, G. Refinement of severely incomplete structures with maximum likelihood in BUSTER-TNT. Acta Crystallogr. D 60, 2210–2221 (2004).
Acknowledgements
This research was supported by a Wellcome Trust senior research fellowship award to L.P. and by the Medical Research Council in the laboratory of S.D.B. We thank X.-J. Lu for help with X3DNA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Crystallographic analysis of the Sso core primase (PDF 871 kb)
Supplementary Fig. 2
Structure-based multiple PriS sequence alignment. (PDF 1147 kb)
Rights and permissions
About this article
Cite this article
Lao-Sirieix, SH., Nookala, R., Roversi, P. et al. Structure of the heterodimeric core primase. Nat Struct Mol Biol 12, 1137–1144 (2005). https://doi.org/10.1038/nsmb1013
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1013
This article is cited by
-
Primer synthesis by a eukaryotic-like archaeal primase is independent of its Fe-S cluster
Nature Communications (2017)
-
A primase subunit essential for efficient primer synthesis by an archaeal eukaryotic-type primase
Nature Communications (2015)
-
Essential role of the iron-sulfur cluster binding domain of the primase regulatory subunit Pri2 in DNA replication initiation
Protein & Cell (2015)
-
DNA replication and homologous recombination factors: acting together to maintain genome stability
Chromosoma (2013)