Abstract
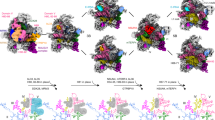
The 40S small ribosomal subunit is cotranscriptionally assembled in the nucleolus as part of a large chaperone complex called the 90S preribosome or small-subunit processome. Here, we present the 3.2-Å-resolution structure of the Chaetomium thermophilum 90S preribosome, which allowed us to build atomic structures for 34 assembly factors, including the Mpp10 complex, Bms1, Utp14 and Utp18, and the complete U3 small nucleolar ribonucleoprotein. Moreover, we visualized the U3 RNA heteroduplexes with a 5′ external transcribed spacer (5′ ETS) and pre-18S RNA, and their stabilization by 90S factors. Overall, the structure explains how a highly intertwined network of assembly factors and pre-rRNA guide the sequential, independent folding of the individual pre-40S domains while the RNA regions forming the 40S active sites are kept immature. Finally, by identifying the unprocessed A1 cleavage site and the nearby Utp24 endonuclease, we suggest a proofreading model for regulated 5′-ETS separation and 90S–pre-40S transition.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Woolford, J.L. Jr. & Baserga, S.J. Ribosome biogenesis in the yeast Saccharomyces cerevisiae. Genetics 195, 643–681 (2013).
Henras, A.K., Plisson-Chastang, C., O'Donohue, M.F., Chakraborty, A. & Gleizes, P.E. An overview of pre-ribosomal RNA processing in eukaryotes. Wiley Interdiscip. Rev. RNA 6, 225–242 (2015).
Turowski, T.W. & Tollervey, D. Cotranscriptional events in eukaryotic ribosome synthesis. Wiley Interdiscip. Rev. RNA 6, 129–139 (2015).
Grandi, P. et al. 90S pre-ribosomes include the 35S pre-rRNA, the U3 snoRNP, and 40S subunit processing factors but predominantly lack 60S synthesis factors. Mol. Cell 10, 105–115 (2002).
Dragon, F. et al. A large nucleolar U3 ribonucleoprotein required for 18S ribosomal RNA biogenesis. Nature 417, 967–970 (2002).
Chaker-Margot, M., Hunziker, M., Barandun, J., Dill, B.D. & Klinge, S. Stage-specific assembly events of the 6-MDa small-subunit processome initiate eukaryotic ribosome biogenesis. Nat. Struct. Mol. Biol. 22, 920–923 (2015).
Pérez-Fernández, J., Martín-Marcos, P. & Dosil, M. Elucidation of the assembly events required for the recruitment of Utp20, Imp4 and Bms1 onto nascent pre-ribosomes. Nucleic Acids Res. 39, 8105–8121 (2011).
Zhang, L., Wu, C., Cai, G., Chen, S. & Ye, K. Stepwise and dynamic assembly of the earliest precursors of small ribosomal subunits in yeast. Genes Dev. 30, 718–732 (2016).
Granneman, S., Petfalski, E., Swiatkowska, A. & Tollervey, D. Cracking pre-40S ribosomal subunit structure by systematic analyses of RNA-protein cross-linking. EMBO J. 29, 2026–2036 (2010).
Lebaron, S. et al. Rrp5 binding at multiple sites coordinates pre-rRNA processing and assembly. Mol. Cell 52, 707–719 (2013).
Lin, J., Lu, J., Feng, Y., Sun, M. & Ye, K. An RNA-binding complex involved in ribosome biogenesis contains a protein with homology to tRNA CCA-adding enzyme. PLoS Biol. 11, e1001669 (2013).
Mougey, E.B. et al. The terminal balls characteristic of eukaryotic rRNA transcription units in chromatin spreads are rRNA processing complexes. Genes Dev. 7, 1609–1619 (1993).
Dutca, L.M., Gallagher, J.E. & Baserga, S.J. The initial U3 snoRNA:pre-rRNA base pairing interaction required for pre-18S rRNA folding revealed by in vivo chemical probing. Nucleic Acids Res. 39, 5164–5180 (2011).
Horn, D.M., Mason, S.L. & Karbstein, K. Rcl1 protein, a novel nuclease for 18 S ribosomal RNA production. J. Biol. Chem. 286, 34082–34087 (2011).
Karbstein, K. & Doudna, J.A. GTP-dependent formation of a ribonucleoprotein subcomplex required for ribosome biogenesis. J. Mol. Biol. 356, 432–443 (2006).
Kornprobst, M. et al. Architecture of the 90S pre-ribosome: a structural view on the birth of the eukaryotic ribosome. Cell 166, 380–393 (2016).
Chaker-Margot, M., Barandun, J., Hunziker, M. & Klinge, S. Architecture of the yeast small subunit processome. Science 355, eaal1880 (2017).
Sun, Q. et al. Molecular architecture of the 90S small subunit pre-ribosome. eLife 6, e22086 (2017).
Zhang, C. et al. Integrative structural analysis of the UTPB complex, an early assembly factor for eukaryotic small ribosomal subunits. Nucleic Acids Res. 44, 7475–7486 (2016).
Hunziker, M. et al. UtpA and UtpB chaperone nascent pre-ribosomal RNA and U3 snoRNA to initiate eukaryotic ribosome assembly. Nat. Commun. 7, 12090 (2016).
Pöll, G. et al. In vitro reconstitution of yeast tUTP/UTP A and UTP B subcomplexes provides new insights into their modular architecture. PLoS One 9, e114898 (2014).
Tafforeau, L. et al. The complexity of human ribosome biogenesis revealed by systematic nucleolar screening of pre-rRNA processing factors. Mol. Cell 51, 539–551 (2013).
Bleichert, F., Granneman, S., Osheim, Y.N., Beyer, A.L. & Baserga, S.J. The PINc domain protein Utp24, a putative nuclease, is required for the early cleavage steps in 18S rRNA maturation. Proc. Natl. Acad. Sci. USA 103, 9464–9469 (2006).
Wells, G.R. et al. The PIN domain endonuclease Utp24 cleaves pre-ribosomal RNA at two coupled sites in yeast and humans. Nucleic Acids Res. 44, 5399–5409 (2016).
Wimberly, B.T. et al. Structure of the 30S ribosomal subunit. Nature 407, 327–339 (2000).
Meyer, B. et al. The Bowen-Conradi syndrome protein Nep1 (Emg1) has a dual role in eukaryotic ribosome biogenesis, as an essential assembly factor and in the methylation of Ψ1191 in yeast 18S rRNA. Nucleic Acids Res. 39, 1526–1537 (2011).
Thomas, S.R., Keller, C.A., Szyk, A., Cannon, J.R. & Laronde-Leblanc, N.A. Structural insight into the functional mechanism of Nep1/Emg1 N1-specific pseudouridine methyltransferase in ribosome biogenesis. Nucleic Acids Res. 39, 2445–2457 (2011).
Ben-Shem, A., Jenner, L., Yusupova, G. & Yusupov, M. Crystal structure of the eukaryotic ribosome. Science 330, 1203–1209 (2010).
Lamanna, A.C. & Karbstein, K. An RNA conformational switch regulates pre-18S rRNA cleavage. J. Mol. Biol. 405, 3–17 (2011).
Granneman, S., Kudla, G., Petfalski, E. & Tollervey, D. Identification of protein binding sites on U3 snoRNA and pre-rRNA by UV cross-linking and high-throughput analysis of cDNAs. Proc. Natl. Acad. Sci. USA 106, 9613–9618 (2009).
Granneman, S. et al. The hU3-55K protein requires 15.5K binding to the box B/C motif as well as flanking RNA elements for its association with the U3 small nucleolar RNA in vitro. J. Biol. Chem. 277, 48490–48500 (2002).
Sharma, K. & Tollervey, D. Base pairing between U3 small nucleolar RNA and the 5′ end of 18S rRNA is required for pre-rRNA processing. Mol. Cell. Biol. 19, 6012–6019 (1999).
Shah, B.N., Liu, X. & Correll, C.C. Imp3 unfolds stem structures in pre-rRNA and U3 snoRNA to form a duplex essential for small subunit processing. RNA 19, 1372–1383 (2013).
Marmier-Gourrier, N., Cléry, A., Schlotter, F., Senty-Ségault, V. & Branlant, C. A second base pair interaction between U3 small nucleolar RNA and the 5′-ETS region is required for early cleavage of the yeast pre-ribosomal RNA. Nucleic Acids Res. 39, 9731–9745 (2011).
Liu, P.C. & Thiele, D.J. Novel stress-responsive genes EMG1 and NOP14 encode conserved, interacting proteins required for 40S ribosome biogenesis. Mol. Biol. Cell 12, 3644–3657 (2001).
Milkereit, P. et al. A Noc complex specifically involved in the formation and nuclear export of ribosomal 40 S subunits. J. Biol. Chem. 278, 4072–4081 (2003).
Wegierski, T., Billy, E., Nasr, F. & Filipowicz, W. Bms1p, a G-domain-containing protein, associates with Rcl1p and is required for 18S rRNA biogenesis in yeast. RNA 7, 1254–1267 (2001).
Karbstein, K., Jonas, S. & Doudna, J.A. An essential GTPase promotes assembly of preribosomal RNA processing complexes. Mol. Cell 20, 633–643 (2005).
McCaughan, U.M. et al. Pre-40S ribosome biogenesis factor Tsr1 is an inactive structural mimic of translational GTPases. Nat. Commun. 7, 11789 (2016).
Madru, C. et al. Chaperoning 5S RNA assembly. Genes Dev. 29, 1432–1446 (2015).
Zheng, S., Lan, P., Liu, X. & Ye, K. Interaction between ribosome assembly factors Krr1 and Faf1 is essential for formation of small ribosomal subunit in yeast. J. Biol. Chem. 289, 22692–22703 (2014).
Strunk, B.S. et al. Ribosome assembly factors prevent premature translation initiation by 40S assembly intermediates. Science 333, 1449–1453 (2011).
Lu, J., Sun, M. & Ye, K. Structural and functional analysis of Utp23, a yeast ribosome synthesis factor with degenerate PIN domain. RNA 19, 1815–1824 (2013).
Thoms, M. et al. The exosome is recruited to RNA substrates through specific adaptor proteins. Cell 162, 1029–1038 (2015).
Wu, S. et al. Diverse roles of assembly factors revealed by structures of late nuclear pre-60S ribosomes. Nature 534, 133–137 (2016).
Barrio-Garcia, C. et al. Architecture of the Rix1-Rea1 checkpoint machinery during pre-60S-ribosome remodeling. Nat. Struct. Mol. Biol. 23, 37–44 (2016).
Sardana, R. et al. The DEAH-box helicase Dhr1 dissociates U3 from the pre-rRNA to promote formation of the central pseudoknot. PLoS Biol. 13, e1002083 (2015).
Segerstolpe, Å. et al. Multiple RNA interactions position Mrd1 at the site of the small subunit pseudoknot within the 90S pre-ribosome. Nucleic Acids Res. 41, 1178–1190 (2013).
Zheng, S.Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Scheres, S.H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Kimanius, D., Forsberg, B.O., Scheres, S.H. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, e18722 (2016).
Scheres, S.H. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 9, 853–854 (2012).
Kucukelbir, A., Sigworth, F.J. & Tagare, H.D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Rabl, J., Leibundgut, M., Ataide, S.F., Haag, A. & Ban, N. Crystal structure of the eukaryotic 40S ribosomal subunit in complex with initiation factor 1. Science 331, 730–736 (2011).
Biasini, M. et al. SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res. 42, W252–W258 (2014).
Goddard, T.D., Huang, C.C. & Ferrin, T.E. Visualizing density maps with UCSF Chimera. J. Struct. Biol. 157, 281–287 (2007).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003).
Popenda, M. et al. Automated 3D structure composition for large RNAs. Nucleic Acids Res. 40, e112 (2012).
Lin, J. et al. Structural basis for site-specific ribose methylation by box C/D RNA protein complexes. Nature 469, 559–563 (2011).
Buchan, D.W., Minneci, F., Nugent, T.C., Bryson, K. & Jones, D.T. Scalable web services for the PSIPRED Protein Analysis Workbench. Nucleic Acids Res. 41, W349–W357 (2013).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Nicholls, R.A., Fischer, M., McNicholas, S. & Murshudov, G.N. Conformation-independent structural comparison of macromolecules with ProSMART. Acta Crystallogr. D Biol. Crystallogr. 70, 2487–2499 (2014).
Brown, A. et al. Tools for macromolecular model building and refinement into electron cryo-microscopy reconstructions. Acta Crystallogr. D Biol. Crystallogr. 71, 136–153 (2015).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Fernández, I.S., Bai, X.C., Murshudov, G., Scheres, S.H. & Ramakrishnan, V. Initiation of translation by cricket paralysis virus IRES requires its translocation in the ribosome. Cell 157, 823–831 (2014).
Acknowledgements
We thank C. Ungewickell, S. Rieder, S. Griesel and B. Jannack for excellent technical assistance. This work was supported by Deutsche Forschungsgemeinschaft (DFG) grants (HU363/10-5 and HU363/12-1 to E.H.; SFB646, GRK1721 and FOR1805 to R.B.). R.B. acknowledges support from the Center for Integrated Protein Science Munich (CiPS-M) and the European Research Council (Advanced Grant CRYOTRANSLATION). We thank the Leibniz-Rechenzentrum Munich (LRZ) for providing computational services and support.
Author information
Authors and Affiliations
Contributions
J.C., N.K., E.H. and R.B. designed the study. N.K. prepared and characterized the sample, and O.B. prepared the EM samples and performed the cryo-EM data collection. J.C. processed the cryo-EM data and built and refined the model. J.C., E.H. and R.B. analyzed and interpreted the structures. J.C., N.K., O.B., E.H. and R.B. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Cryo-EM structure and model of the C. thermophilum 90S preribosome.
a, Gold standard FSC curve showing the average resolution of the 90S preribosome masked and unmasked (left). FSC curves of the refined model against the overall 3.2 Å map (red), the final model refined against one of the two half maps versus the same half map (FSCwork; green), and the model refined against one of the two half map versus the second half map (FSCfree; cyan) (middle). The orientation distribution of the projections is shown on right. b, Side view of the cryo-EM map of the 90S preribosome colored according to local resolution as calculated by ResMap. c, Two side views of the 90S preribosome model. The WD40 domain of Rrp9(PDB: 4J0X) was used as placeholder for the missing WD40 domains of UTP-A complex, UTP-B complex and Enp2. Utp20 could not be modeled and is shown as original electron density. d, Detail view of protein α-helix (left), β-sheet (middle) and RNA A-helix (right)
Supplementary Figure 2 Structures of nonribosomal biogenesis factors (part 1).
The components of UTP-A, UTP-B, U3 snoRNP, Mpp10 complex and Bms1 complex are shown as ribbons (upper row). The isolated complete density maps are shown in the same view colored according to the local resolution (middle row), and density snapshots from each biogenesis factor with molecular models are also shown (lower row).
Supplementary Figure 3 Structures of nonribosomal biogenesis factors (part 2).
Continued from Supplementary Figure 2, the detailed structures of the remaining individual biogenesis factors. Isolated electron density map and molecular models of the 35S rRNA and the U3 snoRNA are also shown. Boxes show the zoom on three RNA regions displaying h43 from the pre-18S region, H8 from the 5’ETS region and a k-turn from the U3 snoRNA.
Supplementary Figure 4 Interactions in UTP-A and UTP-B.
a, Overall structure of the UTP-A complex. Here, the WD40 domain of Rrp9 (PDB: 4J0X) represents the corresponding part of Utp4, Utp17 and the entire Utp8. The UTP-A oligomerization domain is represented by poly-alanine helices. b, Two views highlighting the location of the UTP-A complex in context of the 90S preribosome. c, Close-up view on the detailed interaction between the C-terminus of Utp17 and the N-terminus of Utp10. In agreement with the previously observed resistance of the complex to high salt, the interaction is mediated by hydrophobic residues from both proteins forming a hydrophobic interaction surface. d, Overall structure of the UTP-B complex. Here, four WD40 domains could not be unambiguously assigned, but according to their location, they likely belong to Utp12 and Utp13. e-f, The overall (e) and the close-up view (f) demonstrating the location of UTP-B complex in the context of 90S preribosome. To indicate the location of Utp12 and Utp13, also h44 of the pre-40S is highlighted. g, Close-up view on the interaction of Utp1 (ribbon) with Utp21 (surface).
Supplementary Figure 5 The structure of the pre-40S domains.
a-c, The overall structure of the pre-40S subunit domains within the 90S particle. Close-up are shown for the 5’ domain (a), the platform (b) and part of the head (c) of the pre-40S. All sub-domains of the pre-18S rRNA were colored according to Fig. 4. Note that ribosomal proteins (eS4, uS4, eS6, eS8, uS12 and eS24 for 5 ’Domain; eS1, eS7, uS8, uS11, uS15 for the platform; uS7, uS9 and eS28 for part of the head (h28-30 and h41-h43)) are already in their mature position and interact with the correct rRNA region. They were all fitted into the density. d, Close-up view on part of the 3’ Major domain (h31-h34) of 18S rRNA as represented by a low resolved electron density (shown in purple). In the proposed model, h31 would be located close to the unambiguously fit h30, and its location would be consistent with the closeby position of Emg1. e, View focusing on the location of the other part of the 3’ Major domain (h35-h40). Extra electron density for h35-h40 (purple) is located on top of the Emg1 dimer. Electron density for the 90S preribosome was low-pass filtered to 6 Å and filtered to 12 Å for the 3’ Major domain. f, The immature rRNA helix h16 is shown in color with mature state in grey. The position of h16 was based on superposition with the 5’Domain. g, h18 (left), h28 and h44 (right) are shown as model with density.
Supplementary Figure 6 Overall structure of the U3 snoRNP.
a, Ribbon model of the U3 snoRNP structure shown in electron density (transparent). In addition, four regions from 35S rRNA (Box A and BoxA’ of pre-18S rRNA, 5’- and 3’ hinge of 5’ETS) which hybridize with U3 snoRNA are shown. b, Close-up views on BoxA’, BoxA, BoxB/C, 5’Hinge, BoxC’/D and 3’Hinge duplex of the U3 snoRNA shown as sticks models and colored according to a. c, Model of the U3 snoRNA. d-e, The proteins of the U3 snoRNP coordinate both UTP-A and UTP-B complex (d) as well as other biogenesis factors like Utp3 and Fcf2 (e).
Supplementary Figure 7 Structures of Utp2 and Mpp10 complexes.
a, Extra density of Utp2-Noc4 cluster (pink) is located on the top surface of Utp15 and close to the Emg1 dimer. The density shows features characteristic for an α-helical HEAT repeat structure and thus is likely representing Noc4. The remaining triangular shaped density connecting with C-terminal helix of Utp2 and the Emg1 dimer likely represents Utp2. b, Close-up view on the interaction between Imp3 and Mpp10. Two helices of Imp3 form a hydrophobic surface and interact with Mpp10 (residues 608-639). Residues in this interface crucial for interaction are labeled. c, Close-up view on the interaction between Imp4 and Mpp10. The β-sheet of Imp4 forms a wide surface for Mpp10 residues 528-552, representing a mixture of hydrophobic and hydrogen bond interactions.
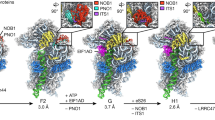
Supplementary Figure 8 The sequential binding model for the 90S preribosome assembly.
Structural analysis suggests dividing in 7 steps illustrated from left to right: Start with early 5’ETS Helix1-2 and UTP-A complex, via complete 5’ETS, 5’Domain, Central, 3’Major, 3’minor and ITS1.
Supplementary Figure 9 Structures of RNA-interacting WD40-domain proteins.
All assigned WD40 domain proteins in the 90S preribosome that interact with RNAs shown as ribbons. Basically, they can be divided into three clusters: Utp1-WD1, Utp1-WD2, Utp7, Utp17-WD1, Sof1 and Utp21-WD2 interact with RNA using their side surfaces; Utp4-WD1, Utp18 and Rrp9 interact with RNA using their top surfaces; Utp21-WD1 interact with RNA using its bottom surface. Based on this analysis, the WD40 domain represents an important RNA binder that can interact with RNA using multiple binding surfaces.
Supplementary Figure 10 Secondary-structure diagram of mature and immature 18S rRNA.
The immature 18S rRNA as observed in the 90S particle is shown in full scale and only the region around the pseudoknot is shown for mature 18S rRNA. Different sub-domains are color coded. The connecting sequences of 5’ETS, U3 snoRNA and ITS1 are shown by a dashed line.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Table 1 (PDF 3167 kb)
Rights and permissions
About this article
Cite this article
Cheng, J., Kellner, N., Berninghausen, O. et al. 3.2-Å-resolution structure of the 90S preribosome before A1 pre-rRNA cleavage. Nat Struct Mol Biol 24, 954–964 (2017). https://doi.org/10.1038/nsmb.3476
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3476
This article is cited by
-
Identification and characterization of sugar-regulated promoters in Chaetomium thermophilum
BMC Biotechnology (2023)
-
Artificial intelligence-assisted cryoEM structure of Bfr2-Lcp5 complex observed in the yeast small subunit processome
Communications Biology (2022)
-
A distinct assembly pathway of the human 39S late pre-mitoribosome
Nature Communications (2021)
-
In situ cryo-electron tomography reveals gradient organization of ribosome biogenesis in intact nucleoli
Nature Communications (2021)
-
The RNA helicase Dbp7 promotes domain V/VI compaction and stabilization of inter-domain interactions during early 60S assembly
Nature Communications (2021)