Abstract
Damaged mitochondria undergo mitophagy, a specialized form of autophagy that is initiated by the protein kinase PINK1 and the ubiquitin E3 ligase Parkin. Ubiquitin-specific protease USP30 antagonizes Parkin-mediated ubiquitination events on mitochondria and is a key negative regulator of mitophagy. Parkin and USP30 both show a preference for assembly or disassembly, respectively, of Lys6-linked polyubiquitin, a chain type that has not been well studied. Here we report crystal structures of human USP30 bound to monoubiquitin and Lys6-linked diubiquitin, which explain how USP30 achieves Lys6-linkage preference through unique ubiquitin binding interfaces. We assess the interplay between USP30, PINK1 and Parkin and show that distally phosphorylated ubiquitin chains impair USP30 activity. Lys6-linkage-specific affimers identify numerous mitochondrial substrates for this modification, and we show that USP30 regulates Lys6-polyubiquitinated TOM20. Our work provides insights into the architecture, activity and regulation of USP30, which will aid drug design against this and related enzymes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Komander, D. & Rape, M. The ubiquitin code. Annu. Rev. Biochem. 81, 203–229 (2012).
Hershko, A. & Ciechanover, A. The ubiquitin system. Annu. Rev. Biochem. 67, 425–479 (1998).
Yau, R. & Rape, M. The increasing complexity of the ubiquitin code. Nat. Cell Biol. 18, 579–586 (2016).
Swatek, K.N. & Komander, D. Ubiquitin modifications. Cell Res. 26, 399–422 (2016).
Pickrell, A.M. & Youle, R.J. The roles of PINK1, Parkin, and mitochondrial fidelity in Parkinson's disease. Neuron 85, 257–273 (2015).
Nguyen, T.N., Padman, B.S. & Lazarou, M. Deciphering the molecular signals of PINK1/Parkin mitophagy. Trends Cell Biol. 26, 733–744 (2016).
Kazlauskaite, A. & Muqit, M.M.K. PINK1 and Parkin—mitochondrial interplay between phosphorylation and ubiquitylation in Parkinson's disease. FEBS J. 282, 215–223 (2015).
Narendra, D.P. et al. PINK1 is selectively stabilized on impaired mitochondria to activate Parkin. PLoS Biol. 8, e1000298 (2010).
Kazlauskaite, A. et al. Parkin is activated by PINK1-dependent phosphorylation of ubiquitin at Ser65. Biochem. J. 460, 127–139 (2014).
Koyano, F. et al. Ubiquitin is phosphorylated by PINK1 to activate parkin. Nature 510, 162–166 (2014).
Kane, L.A. et al. PINK1 phosphorylates ubiquitin to activate Parkin E3 ubiquitin ligase activity. J. Cell Biol. 205, 143–153 (2014).
Ordureau, A. et al. Quantitative proteomics reveal a feedforward mechanism for mitochondrial PARKIN translocation and ubiquitin chain synthesis. Mol. Cell 56, 360–375 (2014).
Wauer, T. et al. Ubiquitin Ser65 phosphorylation affects ubiquitin structure, chain assembly and hydrolysis. EMBO J. 34, 307–325 (2015).
Wauer, T., Simicek, M., Schubert, A. & Komander, D. Mechanism of phospho-ubiquitin-induced PARKIN activation. Nature 524, 370–374 (2015).
Kazlauskaite, A. et al. Binding to serine 65-phosphorylated ubiquitin primes Parkin for optimal PINK1-dependent phosphorylation and activation. EMBO Rep. 16, 939–954 (2015).
Sauvé, V. et al. A Ubl/ubiquitin switch in the activation of Parkin. EMBO J. 34, 2492–2505 (2015).
Kumar, A. et al. Disruption of the autoinhibited state primes the E3 ligase parkin for activation and catalysis. EMBO J. 34, 2506–2521 (2015).
Kondapalli, C. et al. PINK1 is activated by mitochondrial membrane potential depolarization and stimulates Parkin E3 ligase activity by phosphorylating Serine 65. Open Biol. 2, 120080 (2012).
Sarraf, S.A. et al. Landscape of the PARKIN-dependent ubiquitylome in response to mitochondrial depolarization. Nature 496, 372–376 (2013).
Durcan, T.M. et al. USP8 regulates mitophagy by removing K6-linked ubiquitin conjugates from parkin. EMBO J. 33, 2473–2491 (2014).
Ordureau, A. et al. Defining roles of PARKIN and ubiquitin phosphorylation by PINK1 in mitochondrial quality control using a ubiquitin replacement strategy. Proc. Natl. Acad. Sci. USA 112, 6637–6642 (2015).
Lazarou, M. et al. The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Nature 524, 309–314 (2015).
Heo, J.-M., Ordureau, A., Paulo, J.A., Rinehart, J. & Harper, J.W. The PINK1-PARKIN mitochondrial ubiquitylation pathway drives a program of OPTN/NDP52 recruitment and TBK1 activation to promote mitophagy. Mol. Cell 60, 7–20 (2015).
Pickrell, A.M. et al. Endogenous parkin preserves dopaminergic substantia nigral neurons following mitochondrial DNA mutagenic stress. Neuron 87, 371–381 (2015).
Corti, O., Lesage, S. & Brice, A. What genetics tells us about the causes and mechanisms of Parkinson's disease. Physiol. Rev. 91, 1161–1218 (2011).
Mevissen, T.E.T. & Komander, D. Mechanisms of deubiquitinase specificity and regulation. Annu. Rev. Biochem. 86, 159–192 (2017).
Clague, M.J. et al. Deubiquitylases from genes to organism. Physiol. Rev. 93, 1289–1315 (2013).
Cornelissen, T. et al. The deubiquitinase USP15 antagonizes Parkin-mediated mitochondrial ubiquitination and mitophagy. Hum. Mol. Genet. 23, 5227–5242 (2014).
Bingol, B. et al. The mitochondrial deubiquitinase USP30 opposes parkin-mediated mitophagy. Nature 510, 370–375 (2014).
Liang, J.-R. et al. USP30 deubiquitylates mitochondrial Parkin substrates and restricts apoptotic cell death. EMBO Rep. 16, 618–627 (2015).
Cunningham, C.N. et al. USP30 and parkin homeostatically regulate atypical ubiquitin chains on mitochondria. Nat. Cell Biol. 17, 160–169 (2015).
Wang, Y. et al. Deubiquitinating enzymes regulate PARK2-mediated mitophagy. Autophagy 11, 595–606 (2015).
Nakamura, N. & Hirose, S. Regulation of mitochondrial morphology by USP30, a deubiquitinating enzyme present in the mitochondrial outer membrane. Mol. Biol. Cell 19, 1903–1911 (2008).
Urbé, S. et al. Systematic survey of deubiquitinase localization identifies USP21 as a regulator of centrosome- and microtubule-associated functions. Mol. Biol. Cell 23, 1095–1103 (2012).
Elia, A.E.H. et al. Quantitative proteomic atlas of ubiquitination and acetylation in the DNA damage response. Mol. Cell 59, 867–881 (2015).
Lin, D.Y.-W., Diao, J., Zhou, D. & Chen, J. Biochemical and structural studies of a HECT-like ubiquitin ligase from Escherichia coli O157:H7. J. Biol. Chem. 286, 441–449 (2011).
Hospenthal, M.K., Freund, S.M.V. & Komander, D. Assembly, analysis and architecture of atypical ubiquitin chains. Nat. Struct. Mol. Biol. 20, 555–565 (2013).
Faesen, A.C. et al. The differential modulation of USP activity by internal regulatory domains, interactors and eight ubiquitin chain types. Chem. Biol. 18, 1550–1561 (2011).
Ritorto, M.S. et al. Screening of DUB activity and specificity by MALDI-TOF mass spectrometry. Nat. Commun. 5, 4763 (2014).
Yue, W. et al. A small natural molecule promotes mitochondrial fusion through inhibition of the deubiquitinase USP30. Cell Res. 24, 482–496 (2014).
Bingol, B. & Sheng, M. Mechanisms of mitophagy: PINK1, Parkin, USP30 and beyond. Free Radic. Biol. Med. 100, 210–222 (2016).
Michel, M.A., Swatek, K.N., Hospenthal, M.K. & Komander, D. Ubiquitin linkage-specific affimers reveal insights into K6-linked ubiquitin signaling. Mol. Cell (2017).
Ye, Y., Scheel, H., Hofmann, K. & Komander, D. Dissection of USP catalytic domains reveals five common insertion points. Mol. Biosyst. 5, 1797–1808 (2009).
Ekkebus, R. et al. On terminal alkynes that can react with active-site cysteine nucleophiles in proteases. J. Am. Chem. Soc. 135, 2867–2870 (2013).
Hu, M. et al. Crystal structure of a UBP-family deubiquitinating enzyme in isolation and in complex with ubiquitin aldehyde. Cell 111, 1041–1054 (2002).
Ye, Y. et al. Polyubiquitin binding and cross-reactivity in the USP domain deubiquitinase USP21. EMBO Rep. 12, 350–357 (2011).
Renatus, M. et al. Structural basis of ubiquitin recognition by the deubiquitinating protease USP2. Structure 14, 1293–1302 (2006).
Komander, D. et al. Molecular discrimination of structurally equivalent Lys63-linked and linear polyubiquitin chains. EMBO Rep. 10, 466–473 (2009).
Komander, D. et al. The structure of the CYLD USP domain explains its specificity for Lys63-linked polyubiquitin and reveals a B box module. Mol. Cell 29, 451–464 (2008).
Sato, Y. et al. Structures of CYLD USP with Met1- or Lys63-linked diubiquitin reveal mechanisms for dual specificity. Nat. Struct. Mol. Biol. 22, 222–229 (2015).
Okatsu, K. et al. Phosphorylated ubiquitin chain is the genuine Parkin receptor. J. Cell Biol. 209, 111–128 (2015).
Dong, X. et al. Ubiquitin S65 phosphorylation engenders a pH-sensitive conformational switch. Proc. Natl. Acad. Sci. USA 114, 6770–6775 (2017).
Kim, W. et al. Systematic and quantitative assessment of the ubiquitin-modified proteome. Mol. Cell 44, 325–340 (2011).
Wagner, S. A. et al. A proteome-wide, quantitative survey of in vivo ubiquitylation sites reveals widespread regulatory roles. Mol Cell Proteomics 10, M111.013284 (2011).
Newton, K. et al. Ubiquitin chain editing revealed by polyubiquitin linkage-specific antibodies. Cell 134, 668–678 (2008).
Matsumoto, M.L. et al. K11-linked polyubiquitination in cell cycle control revealed by a K11 linkage-specific antibody. Mol. Cell 39, 477–484 (2010).
Matsumoto, M.L. et al. Engineering and structural characterization of a linear polyubiquitin-specific antibody. J. Mol. Biol. 418, 134–144 (2012).
Berrow, N.S. et al. A versatile ligation-independent cloning method suitable for high-throughput expression screening applications. Nucleic Acids Res. 35, e45 (2007).
Higuchi, R., Krummel, B. & Saiki, R.K. A general method of in vitro preparation and specific mutagenesis of DNA fragments: study of protein and DNA interactions. Nucleic Acids Res. 16, 7351–7367 (1988).
Faggiano, S., Alfano, C. & Pastore, A. The missing links to link ubiquitin: methods for the enzymatic production of polyubiquitin chains. Anal. Biochem. 492, 82–90 (2016).
Wilkinson, K.D., Gan-Erdene, T. & Kolli, N. Derivitization of the C-terminus of ubiquitin and ubiquitin-like proteins using intein chemistry: methods and uses. Methods Enzymol. 399, 37–51 (2005).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Waterman, D.G. et al. Diffraction-geometry refinement in the DIALS framework. Acta Crystallogr. D Struct. Biol. 72, 558–575 (2016).
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. D Biol. Crystallogr. 62, 72–82 (2006).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Stein, N. IUCr. CHAINSAW: a program for mutating PDB files used as templates in molecular replacement. J. Appl. Crystallogr. 41, 641–643 (2008).
Vijay-Kumar, S., Bugg, C.E. & Cook, W.J. Structure of ubiquitin refined at 1.8 A resolution. J. Mol. Biol. 194, 531–544 (1987).
Nicholls, R.A., Fischer, M., McNicholas, S. & Murshudov, G.N. Conformation-independent structural comparison of macromolecules with ProSMART. Acta Crystallogr. D Biol. Crystallogr. 70, 2487–2499 (2014).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P.D. et al. The Phenix software for automated determination of macromolecular structures. Methods 55, 94–106 (2011).
Murshudov, G.N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D Biol. Crystallogr. 67, 355–367 (2011).
Silva, J.C. et al. Quantitative proteomic analysis by accurate mass retention time pairs. Anal. Chem. 77, 2187–2200 (2005).
Acknowledgements
We thank beamline scientists at Diamond Light Source (DLS) for support at beamlines I04-1, I02 and I24. Access to DLS was supported in part by the EU FP7 infrastructure grant BIOSTRUCT-X (contract 283570). We are grateful to F. El Oualid (UbiQ) and Avacta LifeSciences for providing reagents; W. Harper (Harvard University) for providing inducible HeLa Flp-In T-REx cells expressing Parkin; N. Ader, W. Kukulski (MRC LMB), R. Youle (NIH), S. Urbe and M. Clague (Liverpool) for discussions and sharing of unpublished cell biology data; and members of D.K.'s lab for reagents, discussions and advice. This work was supported by the Medical Research Council (U105192732), the European Research Council (309756), the Michael J. Fox Foundation and the Lister Institute for Preventive Medicine. M.G. is supported through a college postdoctoral associateship by Jesus College, Cambridge. M.A.M. was supported by a fellowship of the Boehringer Ingelheim Fonds and a Doc.Mobility fellowship of the Swiss National Science Foundation.
Author information
Authors and Affiliations
Contributions
Conceptualization, M.G. and D.K.; USP30 protein purification, crystallization, model refinement, biophysics, mass spectrometry, biochemical activity assays and cell biology, M.G.; Parkin assembly assays, C.G.; ubiquitin phosphorylation assays, A.F.S.; mass spectrometry, S.M.; reagents and protocols, M.A.M.; writing, M.G. and D.K.; funding acquisition, D.K.
Corresponding author
Ethics declarations
Competing interests
D.K. is part of the DUB Alliance, which includes Cancer Research Technology and FORMA Therapeutics. M.G. is funded by the DUB Alliance.
Integrated supplementary information
Supplementary Figure 1 Linkage preference of USP30 and USP domain architecture.
a, Catalytic efficiencies determined from Coomassie-stained gel-based kinetics for all eight diUb linkages (see Supplementary Data Set 1 for raw data). Error bars represent standard deviations of the mean, obtained from five (Lys6), three (Lys11, Lys48) or two (Lys27, Lys33, Lys63, Met1) time courses. b, Time course of tetraubiquitin cleavage for Lys6, Lys11, Lys48, Lys63 and Met1-linked chains, resolved by SDS-PAGE and visualised by silver staining. The gel of Lys6-tetraubiquitin from Fig. 1c is included for comparison. It shows at the earliest time points that tri- and monoubiquitin are generated, consistent with exo-DUB activity (Hospenthal, M.K et al., Nat Struct Mol Biol 20, 555–565, 2013). c, Annotation of the USP domain of USP7 (PDB 1NBF, Hu, M. et al., Cell 111, 1041–1054, 2002) according to the box system with insertion points indicated (Ye, Y. et al., Mol Biosyst 5, 1797–1808, 2009). d, USP domain nomenclature indicating thumb, finger and palm subdomains (Hu, M. et al., Cell 111, 1041–1054, 2002). e, Plot of average sequence conservation of USP30 (rolling window average of 9 residues. 10: high sequence conservation, 0: low sequence conservation, obtained from ConSurf, http://consurf.tau.ac.il/2016/) and deuterium incorporation obtained from HDX experiments on USP30c2 (compare Fig. 1d), suggesting flexibility of regions with low sequence conservation.
Supplementary Figure 2 USP30 construct optimization through insertion deletions.
a, Sequence alignment of the C-terminal end of box 4 of USP30 with other USPs that have been crystallised. Of note, the corresponding residues of Phe348, Met350 and Ile353 are typically hydrophilic residues that are solvent exposed on the domain backside in other USP structures, and were mutated in USP30 to increase protein solubility (see Supplementary Table 1). b, Coomassie-stained gel of indicated USP30 proteins used for crystallography. c, Melting temperatures of apo and covalent Ub-PA complexes of indicated USP30 constructs, indicating similar melting temperatures for constructs with or without the box 4-5 insertion and with or without mutation of the three hydrophobic residues shown in a, suggesting that the protein is similarly stable once purified. In contrast, additional deletion of the box 2-3 insertion significantly increased protein stability. Crystallised proteins are indicated with an arrow. Experiments were performed in triplicate and error bars represent standard deviation of the mean. d, Representative anisotropy traces for indicated USP30 proteins in the ubiquitin-KG-TAMRA cleavage assay. e, Catalytic efficiencies of indicated constructs derived from ubiquitin-KG-TAMRA cleavage. Experiments were performed in triplicate and error bars represent standard error. f, Lys6-diubiquitin cleavage assay for indicated constructs.
Supplementary Figure 3 Electron density and SEC-MALS experiment.
Electron density maps in this Figure correspond to weighted 2|FO|–|FC| electron density contoured at 1σ (0.8σ for the 3.6 Å USP30c8~Ub-PA structure). a, Overall view of the asymmetric unit for USP30c13~Ub-PA. b, Overall view of the asymmetric unit for USP30c8~Ub-PA. c, Close-up view of the covalently modified catalytic triad residues of USP30c13~Ub-PA (top) and USP30c8~Ub-PA (bottom). d, Size-exclusion chromatography multi-angle light scattering (SEC-MALS) experiment on complex formation of USP30c13i and Lys6-diubiquitin. While a clear shift is apparent from the elution profile, the observed complex mass is smaller than expected, indicating that the complex is not fully gel-filtration stable at the concentrations tested (80 μM sample concentration, 1.2-fold molar excess of Lys6-diubiquitin). e, Overall view of the asymmetric unit for USP30c13i + Lys6-diUb. f, Close-up view of the isopeptide bond in the USP30 active site from two different perspectives.
Supplementary Figure 4 HDX-MS analysis of ubiquitin binding to USP30.
a, Deuterium incorporation measured by HDX-MS on USP30c8 mapped onto the USP30 molecule from USP30c13i-Lys6-diUb. Data for residues not part of the model are shown as horizontal bars. b, Deuterium incorporation measured by HDX-MS on USP30c8~Ub-PA mapped as in a. c, Subtraction of data from HDX-MS runs on USP30c8 in the presence and absence of Ub-PA mapped onto the model. The deuterium incorporation in the palm and thumb domain is virtually unchanged by ubiquitin binding. There is a significant reduction in deuterium uptake in the fingers domain, suggesting rigidification through ubiquitin binding. The box 2-3 loop insertion (residues 179–217) has regions of high deuterium uptake (>45%) in both the apo and the Ub-PA-bound state, consistent with this part not being required for ubiquitin binding and with no density being observable for this part in the 3.6 Å structure of USP30c8~Ub-PA.
Supplementary Figure 5 Binding of the distal ubiquitin moiety to USP30 and characterization of USP30 active site mutations.
a, Sequence alignment of crystallised USP proteins centring on Leu328 and Phe453 of USP30, showing differences in the coordination of the distal ubiquitin’s C-terminal tail compared to other USPs (compare Fig. 2a). b,c, Fluorescence polarisation binding experiment of USP30c8 C77A (one loop deleted) and USP30c2 C77A (no loop deleted) resulting in similarly poor binding of ubiquitin-KG-TAMRA as compared to USP30c13i (Fig. 2b), indicating that loop deletions do not impact on the distal ubiquitin binding site of USP30. Experiments were performed in triplicate and error bars represent standard deviation from the mean. d, Fluorescence polarisation experiment of ubiquitin-KG-TAMRA binding to inactive USP21 C221A, as a positive control for b, c and Fig. 2b. Experiments were performed in triplicate and error bars represent standard deviation from the mean. e, Triplicate-averaged anisotropy traces for indicated USP30 proteins with active site residues mutated in the ubiquitin-KG-TAMRA cleavage assay. f, Observed rate constants were obtained from fitting anisotropy-time courses plotted over enzyme concentrations. The slopes correspond to catalytic efficiencies (kcat/KM) and are plotted in Fig. 2d.
Supplementary Figure 6 Selective recognition and cleavage of Lys6-diUb by USP30.
a, Fluorescence polarisation (FP) binding experiment with FlAsH-tagged Lys6-diubiquitin and indicated USP30 proteins, showing identical affinities compared to USP30c13i (Fig. 3a), indicating that the deleted loops do not affect binding to Lys6-diubiquitin. b, Control FP binding experiment of FlAsH-tagged diubiquitin probes used in Fig. 3a with inactive USP21 C221A, indicating that all probes can bind USP domains in principle. Mean ± standard error from three independent measurements. c, Isothermal titration calorimetry (ITC) data for asymmetric Lys6-diUb* (distal Ub: K6R, K48R; proximal Ub: K48R, ΔGly76) binding to USP30c13i, confirming the affinities obtained from FP binding assays. d, Superposition of free ubiquitin (PDB 1UBQ, Vijay-Kumar, S. et al., J Mol Biol 194, 531–544, 1987), the proximal Ub of Lys6-diubiquitin (PDB 2XK5, Virdee, S. et al., Nat Chem Biol 6, 750–757, 2010) and the proximal Ub in the USP30c13i-Lys6-diUb structure. Close-up on the β1-β2 loops showing how binding to USP30 induces a distinct conformation in the proximal Ub moiety. e, Sequence alignment of structurally characterised USP domains, centred on Glu159. f, Surface representation of USP30c13i-Lys6-diUb and close-up view of the recognition site for the isopeptide linkage. The distal Ub C-terminus and the isopeptide bond are fully engulfed by the USP domain, with Glu159 of the thumb subdomain contacting the palm subdomain and coordinating the Gly76 amide backbone. g, Specificity analysis for USP30 S477D (compare Fig. 1b). h, Diubiquitin cleavage assay with indicated substrates and USP30 Ser477 mutants at identical concentrations.
Supplementary Figure 7 Analysis of Lys6-diUb upon binding to USP30.
a, Comparison of proximal versus distal ubiquitin binding sites in USP30. The USP30c13i-Lys6-diUb complex was characterised by PISA (http://www.ebi.ac.uk/pdbe/pisa/), revealing a significantly better distal as compared to proximal binding interface: larger interface area (1652 Å2 vs. 535 Å2), larger numbers of hydrogen bonds (39 vs. 8) and salt bridges (10 vs. 2) and lower solvation free energy (–6.4 vs. –2.0 kcal/mol), suggesting that the distal ubiquitin may bind USP30 first. b, Cartoon representation of Lys6-diubiquitin (PDB 2XK5). Lys6-diubiquitin adopts a compact conformation, with Ile36, Leu8, Leu71 of the distal ubiquitin contacting the Ile44-patch of the proximal ubiquitin (Hospenthal, M.K et al., Nat Struct Mol Biol 20, 555–565, 2013). c, Superposition of a and b on the distal Ub moiety. USP30 binds the distal ubiquitin’s Ile36 patch via Trp330, which requires Lys6-diubiquitin to ‘open’ for the initial binding event. d,e,g, Superposition centred on the proximal ubiquitin, analogously to a,b,c. The proximal ubiquitin binding site of USP30 recognises the ubiquitin Phe4 patch, which is exposed in Lys6-diubiquitin. This binding site, hence, could recognise compact Lys6-diubiquitin and enable it to open subsequently.
Supplementary Figure 8 Characterization USP30 mutations of the proximal ubiquitin binding site.
a, Anisotropy traces for indicated USP30 proteins with proximal binding site residues mutated in the ubiquitin-KG-TAMRA cleavage assay. b, Observed rate constants from a plotted over enzyme concentration to derive catalytic efficiencies. c, Catalytic efficiencies of indicated USP30 proteins derived from the ubiquitin-KG-TAMRA cleavage assay. Replacement of His445 and Trp475 leads to a reduction of catalytic activity on ubiquitin-KG-TAMRA, presumably due to their close proximity to the catalytic residues. Mean ± standard error are shown, derived from three independent experiments. d, Diubiquitin cleavage assay with indicated substrates and USP30 mutants, titrated to similar activity levels. H445N and D479Q retain the preference for Lys6-diUb, whereas H445S and H445Q show a cleavage pattern similar to H445E and W475K (see Fig. 3f).
Supplementary Figure 9 CYLD recognizes Lys63-diubiquitin by a distinctly different interface.
a, Open-book representation of the CYLD(C596A)-Lys63-diubiquitin binding interface, compare Fig. 3d. b, Side-on view of the proximal ubiquitin binding site in CYLD. Specific recognition of the proximal Ub is achieved by extensive contacts made by CYLD’s unique β12-β13 insertion, which is contacted by ubiquitin Glu64, Thr14, Thr12 and Phe4, as well as coordination of Met1, Glu16 and Glu18 by the USP domain. While Phe4 packs against the aliphatic part of the Arg824 side chain, polar contacts and salt bridges overall dominate this interface. Hence, while similar ubiquitin surfaces (Phe4 patch) are involved, the distinct orientations of the proximal ubiquitin in USP30 and CYLD dictate different linkage preferences, facilitated by unique structural elements.
Supplementary Figure 10 Parkin-mediated USP30 ubiquitination.
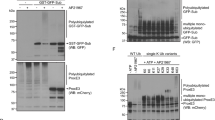
a, Left, ubiquitination of USP30c2 by phosphorylated Parkin, visualised by western blotting with an anti-USP30 antibody. The antibody also detects a prominent proteolytic fragment of USP30 (marked with *) formed by a cleavage in the flexible part of the box 2-3 insertion. As for USP30c13 (Fig. 5a) ubiquitination is enhanced for inactive USP30, suggesting that USP30 can auto-deubiquitinate in trans. Mass spectrometry of these samples identified several (Lys289, Lys235, Lys310) ubiquitination sites. See Supplementary Table 2 for mass spectrometry results. Right, the same samples blotted with an anti-ubiquitin antibody, revealing extensive polyubiquitin assembly by activated Parkin. This contrasts the USP30 blot which does not show evidence for USP30 polyubiquitination. b, Gel shown in Fig. 5c. Anion-exchange chromatography allowed partial separation of F4R-ubiquinated and unmodified USP30 with coeluting E1 and Parkin species. c, Superposition of USP30c13~Ub-PA and USP30c13i-Lys6-diUb, and close-up views on (1) the coordination of Phe4 of the distal ubiquitin moiety by the fingers domain, explaining the rigidification observed in HDX-MS through ubiquitin binding in this region as well as the inability of USP30 to process Ub F4R, (2) the coordination of Lys6 of the distal ubiquitin moiety by the side chain of Glu227 and the carbonyl backbone of Phe273, consistent with the observed exo activity in Lys6-tetraubiquitin cleavage experiments (Fig. 1b).
Supplementary Figure 11 PINK1-mediated monophosphorylation of Lys6-diubiquitin on the distal Ub moiety.
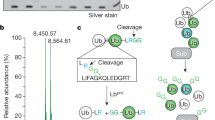
a, ‘Asymmetric’ Lys6-diubiquitin was monophosphorylated by PhPINK1, and following PhPINK1 heat-inactivation cleaved by USP21. Deconvoluted mass spectra are shown. The major phosphorylated species corresponds to the distal Ub moiety. The small peak at 8,618 Da may contain both unphosphorylated distal Ub as well as phosphorylated proximal Ub, which could not be separated due to limited resolution of the mass spectrometer used. b, Close-up view on the distal Ub Ser65 environment of USP30 (see Fig. 6d for superposition in full). c, Deconvoluted mass spectra of the ubiquitin-KG-TAMRA and the Ser65-phosphorylated phosphoUb-KG-TAMRA reagents. Oxidation products (+ 1 oxygen, +16 Da), present also in the untreated sample, are indicated by + [O]. d,e Triplicate averaged anisotropy time traces and observed rate constants for USP30-mediated cleavage of ubiquitin- and phosphoUb-KG-TAMRA (see Fig. 6e for catalytic efficiencies).
Supplementary Figure 12 Characterization data on differently phosphorylated Lys6-diUb substrates.
a, Sample characterisation data for assay shown in Fig. 6f. USP30c13i was ubiquitinated by Parkin, the sample then split and one half treated with PINK1. Analysis of the ubiquitin smear indicates the presence of Ser65-phosphoubiquitin only in the PINK1-treated sample. USP30c13i~Ub is not recognised by either the ubiquitin or the pSer65-ubiquitin antibody. b, Asymmetric Lys6-diUb chains with different phosphorylation states were obtained through assembly from phosphorylated monoUb species. Deconvoluted intact protein mass spectra for reagent characterisation are shown for the chains and the respective monoUb species obtained after treatment with USP21 (compare Supplementary Fig. 11a).
Supplementary Figure 13 Regulation through USP30 overexpression of Lys6-linked polyubiquitin chains on MOM proteins unaffected by USP30 knockdown.
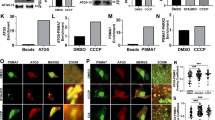
a, Immunoblotting for CISD1 in eluates of Lys6-linked polyubiquitin chain pull-downs from HeLa Flp-In T-REx cells, doxycycline-inducibly expressing wild-type Parkin. Cells treated as indicated, all samples were treated with USP21 to deplete other polyubiquitin linkages. CISD1 forms a constitutive dimer, explaining the pull-down of monoubiquitinated CISD1 protein forms. b, Experiment as in a with USP30 knock down. Immunoblotting for VDAC1, MIRO1 and CISD1. *, non-specific bands. c, Quantification from b with n = 3 independent experiments. Mean ± s.e.m. Wilcoxon-Mann-Whitney test. n.s., non-significant. d, Experiment as in a with overexpression of indicated USP30 variants via transient transfection. e, Quantification from d. Mean ± s.e.m. One-Way ANOVA (F = 48) with Dunnett’s correction, n = 3 independent experiments. ***, P < 0.001.
Supplementary Figure 14 Model of localized mitophagy.
USP30 displays normal levels of activity in the healthy part of the mitochondrial network. In the zone around mitochondrial damage, however, USP30 activity would be reduced due to the activity of PINK1, the presence of phosphoubiquitin and the impaired catalytic of USP30 for phosphoubiquitin substrates.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14, Supplementary Table 1 and Supplementary Note 1. (PDF 5678 kb)
Supplementary Table 2
USP30 peptides identified by mass spectrometry following Parkin-mediated ubiquitination. (XLSX 78 kb)
Supplementary Data Set 1
Images of uncropped gels and western blots. (PDF 3667 kb)
Rights and permissions
About this article
Cite this article
Gersch, M., Gladkova, C., Schubert, A. et al. Mechanism and regulation of the Lys6-selective deubiquitinase USP30. Nat Struct Mol Biol 24, 920–930 (2017). https://doi.org/10.1038/nsmb.3475
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3475
This article is cited by
-
USP30 promotes the progression of breast cancer by stabilising Snail
Cancer Gene Therapy (2024)
-
Parkin inhibits proliferation and migration of bladder cancer via ubiquitinating Catalase
Communications Biology (2024)
-
An expanded lexicon for the ubiquitin code
Nature Reviews Molecular Cell Biology (2023)
-
Molecular basis for ubiquitin/Fubi cross-reactivity in USP16 and USP36
Nature Chemical Biology (2023)
-
Structural and functional characterization of USP47 reveals a hot spot for inhibitor design
Communications Biology (2023)