Abstract
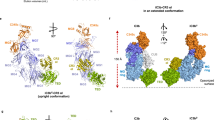
The complement system labels microbes and host debris for clearance. Degradation of surface-bound C3b is pivotal to direct immune responses and protect host cells. How the serine protease factor I (FI), assisted by regulators, cleaves either two or three distant peptide bonds in the CUB domain of C3b remains unclear. We present a crystal structure of C3b in complex with FI and regulator factor H (FH; domains 1–4 with 19–20). FI binds C3b–FH between FH domains 2 and 3 and a reoriented C3b C-terminal domain and docks onto the first scissile bond, while stabilizing its catalytic domain for proteolytic activity. One cleavage in C3b does not affect its overall structure, whereas two cleavages unfold CUB and dislodge the thioester-containing domain (TED), affecting binding of regulators and thereby determining the number of cleavages. These data explain how FI generates late-stage opsonins iC3b or C3dg in a context-dependent manner, to react to foreign, danger or healthy self signals.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Ricklin, D., Hajishengallis, G., Yang, K. & Lambris, J.D. Complement: a key system for immune surveillance and homeostasis. Nat. Immunol. 11, 785–797 (2010).
Stephan, A.H., Barres, B.A. & Stevens, B. The complement system: an unexpected role in synaptic pruning during development and disease. Annu. Rev. Neurosci. 35, 369–389 (2012).
Walport, M.J. Complement. First of two parts. N. Engl. J. Med. 344, 1058–1066 (2001).
Merle, N.S., Church, S.E., Fremeaux-Bacchi, V. & Roumenina, L.T. Complement system part I—molecular mechanisms of activation and regulation. Front. Immunol. 6, 262 (2015).
Merle, N.S., Noe, R., Halbwachs-Mecarelli, L., Fremeaux-Bacchi, V. & Roumenina, L.T. Complement system part II: role in immunity. Front. Immunol. 6, 257 (2015).
Zipfel, P.F., Lauer, N. & Skerka, C. in Inflammation and Retinal Disease: Complement Biology and Pathology Vol. 703 (eds. Lambris, J.D. & Adamis, A.P.) 9–24 (Springer New York, 2010).
Lambert, J.-C. et al. Genome-wide association study identifies variants at CLU and CR1 associated with Alzheimer's disease. Nat. Genet. 41, 1094–1099 (2009).
Brouwers, N. et al. Alzheimer risk associated with a copy number variation in the complement receptor 1 increasing C3b/C4b binding sites. Mol. Psychiatry 17, 223–233 (2012).
Zipfel, P.F. & Skerka, C. Complement regulators and inhibitory proteins. Nat. Rev. Immunol. 9, 729–740 (2009).
Ricklin, D., Reis, E.S. & Lambris, J.D. Complement in disease: a defence system turning offensive. Nat. Rev. Nephrol. 12, 383–401 (2016).
Lin, Z. et al. Complement C3dg-mediated erythrophagocytosis: implications for paroxysmal nocturnal hemoglobinuria. Blood 126, 891–894 (2015).
Ramaglia, V. et al. C3-dependent mechanism of microglial priming relevant to multiple sclerosis. Proc. Natl. Acad. Sci. USA 109, 965–970 (2012).
Carroll, M.C. The complement system in regulation of adaptive immunity. Nat. Immunol. 5, 981–986 (2004).
Kemper, C. & Atkinson, J.P. T-cell regulation: with complements from innate immunity. Nat. Rev. Immunol. 7, 9–18 (2007).
Roversi, P. et al. Structural basis for complement factor I control and its disease-associated sequence polymorphisms. Proc. Natl. Acad. Sci. USA 108, 12839–12844 (2011).
Tsiftsoglou, S.A. et al. The catalytically active serine protease domain of human complement factor I. Biochemistry 44, 6239–6249 (2005).
Lechtenberg, B.C., Johnson, D.J.D., Freund, S.M.V. & Huntington, J.A. NMR resonance assignments of thrombin reveal the conformational and dynamic effects of ligation. Proc. Natl. Acad. Sci. USA 107, 14087–14092 (2010).
Nilsson, S.C. et al. Analysis of binding sites on complement factor I that are required for its activity. J. Biol. Chem. 285, 6235–6245 (2010).
Wu, J. et al. Structure of complement fragment C3b-factor H and implications for host protection by complement regulators. Nat. Immunol. 10, 728–733 (2009).
Forneris, F. et al. Regulators of complement activity mediate inhibitory mechanisms through a common C3b-binding mode. EMBO J. 35, 1133–1149 (2016).
Morgan, H.P. et al. Structural basis for engagement by complement factor H of C3b on a self surface. Nat. Struct. Mol. Biol. 18, 463–470 (2011).
Kajander, T. et al. Dual interaction of factor H with C3d and glycosaminoglycans in host-nonhost discrimination by complement. Proc. Natl. Acad. Sci. USA 108, 2897–2902 (2011).
Lambris, J.D. et al. Dissection of CR1, factor H, membrane cofactor protein, and factor B binding and functional sites in the third complement component. J. Immunol. 156, 4821–4832 (1996).
Sahu, A., Isaacs, S.N., Soulika, A.M. & Lambris, J.D. Interaction of vaccinia virus complement control protein with human complement proteins: factor I-mediated degradation of C3b to iC3b1 inactivates the alternative complement pathway. J. Immunol. 160, 5596–5604 (1998).
Yadav, V.N., Pyaram, K., Mullick, J. & Sahu, A. Identification of hot spots in the variola virus complement inhibitor (SPICE) for human complement regulation. J. Virol. 82, 3283–3294 (2008).
Mortensen, S. et al. Structural basis for the function of complement component C4 within the classical and lectin pathways of complement. J. Immunol. 194, 5488–5496 (2015).
Janssen, B.J.C., Christodoulidou, A., McCarthy, A., Lambris, J.D. & Gros, P. Structure of C3b reveals conformational changes that underlie complement activity. Nature 444, 213–216 (2006).
Ross, G.D. et al. Generation of three different fragments of bound C3 with purified factor I or serum. II. Location of binding sites in the C3 fragments for factors B and H, complement receptors, and bovine conglutinin. J. Exp. Med. 158, 334–352 (1983).
Hocking, H.G. et al. Structure of the N-terminal region of complement factor H and conformational implications of disease-linked sequence variations. J. Biol. Chem. 283, 9475–9487 (2008).
Pechtl, I.C., Kavanagh, D., McIntosh, N., Harris, C.L. & Barlow, P.N. Disease-associated N-terminal complement factor H mutations perturb cofactor and decay-accelerating activities. J. Biol. Chem. 286, 11082–11090 (2011).
Gautam, A.K. et al. Mutational analysis of Kaposica reveals that bridging of MG2 and CUB domains of target protein is crucial for the cofactor activity of RCA proteins. Proc. Natl. Acad. Sci. USA 112, 12794–12799 (2015).
Forneris, F. et al. Structures of C3b in complex with factors B and D give insight into complement convertase formation. Science 330, 1816–1820 (2010).
Nishida, N., Walz, T. & Springer, T.A. Structural transitions of complement component C3 and its activation products. Proc. Natl. Acad. Sci. USA 103, 19737–19742 (2006).
Papanastasiou, M. et al. Structural implications for the formation and function of the complement effector protein iC3b. J. Immunol. 198, 3326–3335 (2017).
Alcorlo, M. et al. Unique structure of iC3b resolved at a resolution of 24 Å by 3D-electron microscopy. Proc. Natl. Acad. Sci. USA 108, 13236–13240 (2011).
Schmidt, C.Q. et al. Rational engineering of a minimized immune inhibitor with unique triple-targeting properties. J. Immunol. 190, 5712–5721 (2013).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Nilsson, S.C. et al. Analysis of binding sites on complement factor I that are required for its activity. J. Biol. Chem. 285, 6235–6245 (2010).
Sartz, L. et al. A novel C3 mutation causing increased formation of the C3 convertase in familial atypical hemolytic uremic syndrome. J. Immunol. 188, 2030–2037 (2012).
Rodriguez, E., Rallapalli, P.M., Osborne, A.J. & Perkins, S.J. New functional and structural insights from updated mutational databases for complement factor H, Factor I, membrane cofactor protein and C3. Biosci. Rep. 34, 635–649 (2014).
Dragon-Durey, M.-A. et al. Heterozygous and homozygous factor h deficiencies associated with hemolytic uremic syndrome or membranoproliferative glomerulonephritis: report and genetic analysis of 16 cases. J. Am. Soc. Nephrol. 15, 787–795 (2004).
Moore, I. et al. Association of factor H autoantibodies with deletions of CFHR1, CFHR3, CFHR4, and with mutations in CFH, CFI, CD46, and C3 in patients with atypical hemolytic uremic syndrome. Blood 115, 379–387 (2010).
Kavanagh, D., Goodship, T.H. & Richards, A. Atypical hemolytic uremic syndrome. Semin. Nephrol. 33, 508–530 (2013).
Ahmad, M. et al. Domain swapping reveals complement control protein modules critical for imparting cofactor and decay-accelerating activities in vaccinia virus complement control protein. J. Immunol. 185, 6128–6137 (2010).
Ricklin, D., Reis, E.S., Mastellos, D.C., Gros, P. & Lambris, J.D. Complement component C3—The “Swiss Army Knife” of innate immunity and host defense. Immunol. Rev. 274, 33–58 (2016).
Janssen, B.J.C. et al. Insights into complement convertase formation based on the structure of the factor B-cobra venom factor complex. EMBO J. 28, 2469–2478 (2009).
Hart, S.P., Ross, J.A., Ross, K., Haslett, C. & Dransfield, I. Molecular characterization of the surface of apoptotic neutrophils: implications for functional downregulation and recognition by phagocytes. Cell Death Differ. 7, 493–503 (2000).
Gershov, D., Kim, S., Brot, N. & Elkon, K.B. C-Reactive protein binds to apoptotic cells, protects the cells from assembly of the terminal complement components, and sustains an antiinflammatory innate immune response: implications for systemic autoimmunity. J. Exp. Med. 192, 1353–1364 (2000).
Verbovetski, I. et al. Opsonization of apoptotic cells by autologous iC3b facilitates clearance by immature dendritic cells, down-regulates DR and CD86, and up-regulates CC chemokine receptor 7. J. Exp. Med. 196, 1553–1561 (2002).
Trouw, L.A. et al. C4b-binding protein and factor H compensate for the loss of membrane-bound complement inhibitors to protect apoptotic cells against excessive complement attack. J. Biol. Chem. 282, 28540–28548 (2007).
Kang, Y.-H., Urban, B.C., Sim, R.B. & Kishore, U. Human complement Factor H modulates C1q-mediated phagocytosis of apoptotic cells. Immunobiology 217, 455–464 (2012).
Verschoor, A. et al. A platelet-mediated system for shuttling blood-borne bacteria to CD8α+ dendritic cells depends on glycoprotein GPIb and complement C3. Nat. Immunol. 12, 1194–1201 (2011).
Broadley, S.P. et al. Dual-track clearance of circulating bacteria balances rapid restoration of blood sterility with induction of adaptive immunity. Cell Host Microbe 20, 36–48 (2016).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Evans, P.R. & Murshudov, G.N. How good are my data and what is the resolution? Acta Crystallogr. D Biol. Crystallogr. 69, 1204–1214 (2013).
Strong, M. et al. Toward the structural genomics of complexes: crystal structure of a PE/PPE protein complex from Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 103, 8060–8065 (2006).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Joosten, R.P., Long, F., Murshudov, G.N. & Perrakis, A. The PDB_REDO server for macromolecular structure model optimization. IUCrJ 1, 213–220 (2014).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
de la Rosa-Trevín, J.M. et al. Xmipp 3.0: an improved software suite for image processing in electron microscopy. J. Struct. Biol. 184, 321–328 (2013).
Scheres, S.H.W., Núñez-Ramírez, R., Sorzano, C.O.S., Carazo, J.M. & Marabini, R. Image processing for electron microscopy single-particle analysis using XMIPP. Nat. Protoc. 3, 977–990 (2008).
Hutin, S., Brennich, M., Maillot, B. & Round, A. Online ion-exchange chromatography for small-angle X-ray scattering. Acta Crystallographica D Structural Biology 72, 1090–1099 (2016).
Shkumatov, A.V. & Strelkov, S.V. DATASW, a tool for HPLC-SAXS data analysis. Acta Crystallogr. D Biol. Crystallogr. 71, 1347–1350 (2015).
Petoukhov, M.V. et al. New developments in the ATSAS program package for small-angle scattering data analysis. J. Appl. Crystallogr. 45, 342–350 (2012).
The PyMOL Molecular Graphics System, V 1.8. (Schrödinger, LLC, 2015).
Notredame, C., Higgins, D.G. & Heringa, J. T-Coffee: A novel method for fast and accurate multiple sequence alignment. J. Mol. Biol. 302, 205–217 (2000).
Acknowledgements
We gratefully thank the European Synchrotron Radiation Facility (ESRF) and the Swiss Light Source (SLS) for the provision of synchrotron radiation facilities and beamline scientists of the ESRF, SLS and the European Molecular Biology Laboratory for assistance. We gratefully acknowledge P. Afanasyev and R. Koning for discussion and assistance in EM data collection and analysis. The work was financially supported by a Top grant (700.54.304 to P.G.) by the Council for Chemical Sciences of the Netherlands Organization for Scientific Research (NWO-CW), the European Research Council (grant no. 233229), the European Community′s Seventh Framework Programmes (FP7/2007-2013) under BioStruct-X (grant no. 283570) and FP7/DIREKT (grant no. 602699 to J.D.L.), and grants by the US National Institutes of Health (AI068730, AI030040; to J.D.L.) and National Science Foundation (no. 1423304 to D.R.). P.G. was further supported by the Institute for Chemical Immunology, an NWO Gravitation project funded by the Ministry of Education, Culture and Science of the Netherlands.
Author information
Authors and Affiliations
Contributions
J.W. cloned human factor 1, MCP and CR1. D.R. and C.Q.S. cloned miniFH. X.X. and J.G. cloned DAF. X.X. and F.F. purified C3b. X.X. produced and purified human FI, DAF, MCP and CR1. C.Q.S. produced and purified miniFH. X.X. crystallized C3b-miniFH and C3b-miniFH-FI complexes, collected X-ray diffraction data, determined and refined the structures. X.X. and T.H.S. prepared negative-stain EM samples, collected and analyzed EM data. X.X. collected and analyzed small angle X-ray scattering data. D.R. and P.D.C. performed surface plasmon resonance experiments and analysis. J.D.L. and P.G. supervised the project. X.X. and P.G. performed data analysis and interpretation, and wrote the manuscript. All authors critically revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Model quality of C3b-miniFH-FI and C3b-miniFH crystal structures and overall structure comparisons.
(a) Overall 2DFo-mFc electron density contoured at 1.0 σ for C3b-miniFH-FI (left) and C3b-miniFH (right). (b) Average B-factor and real-space correlation coefficient for α and β chain of C3b, miniFH and FI for the two independent copies of the complex in the crystal. FI copy 1 exhibits higher real-space correlation and lower B-factor. (b) Missing electron densities for the poly-glycine linker between CCP4 and CCP19 of miniFH in both copies C3b-miniFH-FI and C3b-miniFH structures. The distances between miniFH CCP4 C-terminal and CCP19 N-terminal in the two copies of C3b-miniFH-FI and C3b-miniFH are 36-37 Å. (d) Left-hand side: structural superposition of C3b-miniFH-FI (grey and orange) with C3b-FH CCP1-4 (pdb: 2WII) (Wu, J. et al. Nature Immunology 10, 728–733, 2009) (blue and purple) and C3b-miniFH (cyan and black) shows similar binding of FH CCP1-4 to C3b. Right-hand side: structural superposition of TED domains in C3b-miniFH-FI (grey and orange) with C3b-miniFH (cyan and black) and C3d-FH CCP19-20 (pdb: 3OXU; yellow and green) (Morgan. H. P. et al. Nature SMB 18, 463–470, 2011) shows similar binding of FH CCP19-20 binding to C3b/C3d TED domains. (e) Cartoon representations of the main chains in the structure of free FI (purple) and of FI as observed in C3b-miniFH-FI (green); left side. Several loops could not be modeled in free FI due to disorder (Roversi, P. et al. PNAS 108, 12839–12844, 2011). In the right panel, electron densities (contoured at 1.0 σ level) for these loops and corresponding parts of the model as observed in C3b-miniFH-FI are shown. (f) Structural superposition of C3b CTC domains shows that the core of the CTC domain behaves as a rigid body. In both C3b-miniFH (cyan) and C3b-miniFH-FI (grey) complex, CTC exhibits a β-turn-β configuration of the ‘neck’ region, whereas in the C3b-FH CCP1-4 complex (blue) CTC exhibits a α-helix in this region. Compared to C3b-miniFH-FI (grey) structure, the CTC domain in C3b-miniFH (cyan) displays a 34° rotation due to a twisting of a hinge loop formed by residues E1515-K1526 (highlighted in purple). A simplified diagram in the right side illustrates this feature.
Supplementary Figure 2 Disease-related mutants at protein interfaces in C3b-FH-FI.
(a) Cartoon representation of C3b-miniFH-FI showing disease-related mutation V1658A in C3b (indicated in sticks) located in the middle of the FI-binding site (with colors as defined in Fig. 1). (b) Structures of C3b-Bb-SCIN (pdb code: 2WIN) (Rooijakkers, S. H. M. et al. Nature Immunology 10, 721–727, 2009), C3b-FB (2XWJ) (Forneris, F. et al. Science 330, 1816–1820, 2010) and C3b-FB-FD (2XWB) (Forneris, F. et al. Science 330, 1816–1820, 2010) indicating that C3b V1658 is outside of FB-binding interface. C3b is shown in grey cartoon with V1658 shown in stick representation; FB and Bb are shown in cyan with either Mg2+ or Ni2+ occupying the metal-ion dependent adhesion site (MIDAS) position, as indicated by a sphere. (c) Disease-related residues variations on interaction interfaces of C3b-FI and miniFH-FI. Red boxes indicate loss-of-function variations and the cyan box indicates a gain-of-function, suggesting that it is perhaps not involved in aHUS (Nilsson, S. C. et al. Eur. J. Immunol. 39, 310–323, 2009). (c) Disease-related mutations on FH at the FH-FI interface. Residues on CCP1 possibly have an indirect effect on FI binding through their interactions with CCP2, since NMR chemical shift of the loop between Cys129 and Cys141 in FH CCP2 (indicated in blue) (Hocking, H. G. et al. JBC 283, 9475–9487, 2008).
Supplementary Figure 3 Stabilization of the FI SP domain
(a) Cartoon representation of free FI (Roversi, P. et al. PNAS 108, 12839–12844, 2011) (cyan and purple for heavy and light chain resp.) superposed on FI in the C3b-miniFH-FI complex (blue and green for heavy and light chain resp.). The relative orientation of heavy and light chains differs by 11° between free FI and FI in the C3b-miniFH-FI complex. Residues at the heavy-light chain interface are shown in stick representation. (b) FH CCP 2-3 contact with loop 358-363, loop 394-408, loop 435-448 and loop 471-485 of FI SP domain. The contact regions in these loops are highlighted in purple. (c) Superposition FI SP with thrombin (Huntington, J. A. et al. Structure 11, 469–479, 2003; Lechtenberg, B. C. et al. PNAS 108, 12839–12844, 2011) (pdb: 1JOU) shows FI loop 358-363 and loop 394-408 correspond to thrombin 37 loop (exosite I) and 70 loop (exosite I) resp. and FI loop 471-485 corresponds to thrombin γ loop.
Supplementary Figure 4 Flexibility of the scissile loops in C3b CUB.
(a) Superposition of C3b CUB domains of C3b-miniFH-FI (cyan) and other C3b structures (Forneris, F. et al. EMBO J 10, 1133-49, 2016) (colored in grey), C3b (pdb code: 5FO7), C3b-FH CCP1-4 (2WII), C3b-CR1 CCP15-17 (5FOB), C3b-MCP CCP1-4 (5FO8), C3b-DAF CCP2-4 (5FOA) and C3b-SPICE CCP1-4 (5FO9), shows conformational rearrangements in the first scissile loop. The first scissile bond Arg1303 - Ser1304 is indicated by sticks. (b) B-factor putty representations of the CUB domains with scissile bonds Arg1303-Ser1304, Arg1320-Ser1321 and Arg954-Glu955 are colored in yellow, red and blue resp., indicating increased flexibility for the loops containing the first and third scissile bonds in C3b prior to FI binding.
Supplementary Figure 5 Sequence and structure comparisons of DAF with cofactor regulators.
(a) Structure-based sequence alignment of FH (CCP2-3), CR1 (CCP16-17), CR1 (CCP9-10), MCP (CCP2-3) and DAF (CCP3-4). Cysteines in CCP domains are indicated in bold characters. Residues of FH at the FI-binding interface indicated by red background; with, FI-interacting residues conserved among cofactor regulators indicated by red boxes. Red arrows indicate conserved FI-interacting residues not present in DAF. Residues buried in C3b-binding interfaces are indicted by blue background. (b) Overlay of FH CCP2-3 (orange) with DAF CCP3-4 (magenta). Interface residues that differ markedly between FH and DAF are shown in sticks. (c) Structural superposition of the complex of C3b-FH, C3b-CR1, C3b-MCP, C3b-SPICE and C3b-DAF on CCP ii and CCP iii. The orientation of the CUB domain in C3b-DAF crystal structure differs from the CUB position in the C3b complexes with regulators. Zoom-in on the right-hand panel shows that CUB with regulator fragments positions the first scissile loop close to the catalytic center of FI SP, which is not the case for the arrangement observed in C3b-DAF.
Supplementary Figure 6 Charge variations in cofactors at FI interface.
(a) Superposition of C3b-miniFH-FI with C3b-CR1 CCP15-17 (pdb: 5FOB) (Forneris, F. et al. EMBO J 10, 1133-49, 2016), C3b-MCP CCP 1-4 (pdb: 5FO8) (Forneris, F. et al. EMBO J 10, 1133-49, 2016), and C3b-SPICE CCP1-4 (pdb: 5FO9) (Forneris, F. et al. EMBO J 10, 1133-49, 2016) shows FH contains an extra residue, i.e. E243 (highlighted by a purple box in sequence alignment), and a shorter loop between CCP2 Cys129 and Cys141 (stretch of ‘missing’ residues highlighted by a cyan box in structure-based sequence alignment; with FH residue numbers indicated). FH D137 pointed to FI loop 435-448, which is rich in charge residues (indicated by sticks). At the corresponding position of FH D137, CR1, MCP and SPICE have a conserved lysine (indicated by a red box in sequence alignment). (b) Both the loop between CCP2 Cys129 and Cys141 of FH (indicated by red arrows) and its close by stretch loop 435-448 of FI SP domain display high B-factors. (c) Superposition of SPICE CCP2 and CCP3 (pdb: 5FO9; grey) (Forneris, F. et al. EMBO J 10, 1133-49, 2016) on FH CCP2 and CCP3 (orange) of C3b-miniFH-FI. Critical residues for SPICE cofactor activity are shown as sticks (Yadav, V. N. et al. Journal of Virology 82, 3283–3294, 2008). Y98 of SPICE docks into the hydrophobic pocket in FI, formed by W393, P402, L404 and I409. K107 and K120 are close to FI loop 435-448 consisting of a series of positive and negative charge residues (435xKKDxxKKDxExxR448). Y103 of SPICE points away from FI-SP domain.
Supplementary Figure 7 Negative-stain EM and SAXS analysis of C3b, iC3b1 and iC3b.
(a) C3b was incubated with VCP and FI for 2 hours at 37°C. Shown is chromatogram after separation on a monoQ 15/50GL column. (b) SDS-PAGE of corresponding fractions. The red box indicates the sample fractions of iC3b1 used for EM and SAXS analyses. (c) Raw electron micrographs of purified C3b, iC3b1 and iC3b samples. (d) SAXS modeling. Ab-initio models of C3b (red), iC3b1 (green) and iC3b (blue). iC3b1 resembles C3b, whereas iC3b shows an elongated CUB-TED.
Supplementary Figure 8 Cofactor activities and relative binding affinities of regulator fragments.
(a) SDS gel showing cleavage products from long-term cofactor assay (8-hours incubation at 37 °C) showing FI with either miniFH or FH CCP1-3 yield C3dg/C3d and C3c fragments corresponding to three cleavages, whereas FI with MCP only yields iC3b corresponding to two cleavages. (b) The left three columns show the processed SPR sensorgrams of the recombinant regulator segments (marked on the far left) to C3b (green), iC3b (blue), and C3c (red). C3 fragments were covalently coupled to individual sensor chip surfaces using amine coupling in a non-oriented manner. Regulators were injected for 60 s in a concentration range of 10 nM to 20 μM. Steady state responses were plotted against the regulator concentration, and all analyses were globally fit across all surfaces to a single binding site model (right column; shown as percentage of maximum binding capacity) to determine an apparent affinity value (KD app); R2 values of the fits were all better than 0.985. Please note that detailed quantitative interaction analyses of regulator molecules binding to C3b have previously been conducted by our group; these were based on oriented surface coupling of C3b via its thioester moiety in a close-to-physiological manner (Forneris, F. et al. EMBO J 10, 1133-49, 2016). For the present study, and due to the absence of a thioester domain in C3c, all C3 fragments were immobilized in random orientation, leading to generally weaker affinity values that cannot be directly compared with previous reports regarding absolute numbers (hence the classification as “apparent KD”). However, the immobilization of the three C3 fragments at closely matched surface density and using the same coupling method, and the simultaneous injection of regulators across all three surfaces ensures the qualitative/semi-quantitative assessment of relative binding activities, which was the main purpose of this experiment. Abbreviations: Rmax, maximum binding response; RU, resonance units.
Supplementary Figure 9 Disease-related mutants in FI.
Classification of disease-related variations in FI (as listed in Supplement Table 1). Residues located in C3b-FI and FH-FI binding interfaces are indicated by purple spheres; mutations causing lower FI expression level in vivo indicated by blue spheres, mutations causing FI misfolding cyan, and mutations on the surface, but not at the binding interfaces, grey.
Supplementary Figure 10 Purification of FI.
(a)Western blot by anti-His antibody shows that purified recombinant human FI (with N-terminal His6 tag) stored at 4 ºC is degraded slowly. (b) Chromatogram of gel filtration by Superdex 200 10/300 GL column. Sample fractions for crystallization are highlighted by red background.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10, Supplementary Table 1 and Supplementary Note 1. (PDF 1829 kb)
Rights and permissions
About this article
Cite this article
Xue, X., Wu, J., Ricklin, D. et al. Regulator-dependent mechanisms of C3b processing by factor I allow differentiation of immune responses. Nat Struct Mol Biol 24, 643–651 (2017). https://doi.org/10.1038/nsmb.3427
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3427
This article is cited by
-
Purified complement C3b triggers phagocytosis and activation of human neutrophils via complement receptor 1
Scientific Reports (2023)
-
Insight into mode-of-action and structural determinants of the compstatin family of clinical complement inhibitors
Nature Communications (2022)
-
Invariant surface glycoprotein 65 of Trypanosoma brucei is a complement C3 receptor
Nature Communications (2022)
-
Deconstruction of Neurotrypsin Reveals a Multi-factorially Regulated Activity Affecting Myotube Formation and Neuronal Excitability
Molecular Neurobiology (2022)
-
A receptor for the complement regulator factor H increases transmission of trypanosomes to tsetse flies
Nature Communications (2020)