Abstract
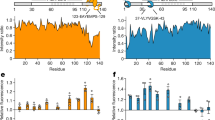
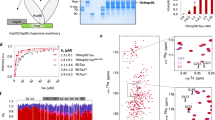
The critical toxic species in over 40 human diseases are misfolded proteins. Their interaction with molecular chaperones such as Hsp90, which preferentially interacts with metastable proteins, is essential for the blocking of disease progression. Here we used nuclear magnetic resonance (NMR) spectroscopy to determine the three-dimensional structure of the misfolded cytotoxic monomer of the amyloidogenic human protein transthyretin, which is characterized by the release of the C-terminal β-strand and perturbations of the A-B loop. The misfolded transthyretin monomer, but not the wild-type protein, binds to human Hsp90. In the bound state, the Hsp90 dimer predominantly populates an open conformation, and transthyretin retains its globular structure. The interaction surface for the transthyretin monomer comprises the N-terminal and middle domains of Hsp90 and overlaps with that of the Alzheimer's-disease-related protein tau. Taken together, the data suggest that Hsp90 uses a mechanism for the recognition of aggregation-prone proteins that is largely distinct from those of other Hsp90 clients.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
13 March 2017
In the version of this article initially published online, there was an error in the y-axis label of Figure 1e. The error has been corrected in the print, PDF and HTML versions of this article.
References
Haass, C. & Selkoe, D.J. Soluble protein oligomers in neurodegeneration: lessons from the Alzheimer's amyloid β-peptide. Nat. Rev. Mol. Cell Biol. 8, 101–112 (2007).
Chiti, F. & Dobson, C.M. Protein misfolding, functional amyloid, and human disease. Annu. Rev. Biochem. 75, 333–366 (2006).
Barral, J.M., Broadley, S.A., Schaffar, G. & Hartl, F.U. Roles of molecular chaperones in protein misfolding diseases. Semin. Cell Dev. Biol. 15, 17–29 (2004).
Bukau, B., Weissman, J. & Horwich, A. Molecular chaperones and protein quality control. Cell 125, 443–451 (2006).
Mayer, M.P. Gymnastics of molecular chaperones. Mol. Cell 39, 321–331 (2010).
Hipp, M.S., Park, S.H. & Hartl, F.U. Proteostasis impairment in protein-misfolding and -aggregation diseases. Trends Cell Biol. 24, 506–514 (2014).
Landry, S.J. & Gierasch, L.M. Polypeptide interactions with molecular chaperones and their relationship to in vivo protein folding. Annu. Rev. Biophys. Biomol. Struct. 23, 645–669 (1994).
Lindberg, I. et al. Chaperones in neurodegeneration. J. Neurosci. 35, 13853–13859 (2015).
Schneider, C. et al. Pharmacologic shifting of a balance between protein refolding and degradation mediated by Hsp90. Proc. Natl. Acad. Sci. USA 93, 14536–14541 (1996).
Jakob, U., Lilie, H., Meyer, I. & Buchner, J. Transient interaction of Hsp90 with early unfolding intermediates of citrate synthase. Implications for heat shock in vivo. J. Biol. Chem. 270, 7288–7294 (1995).
Street, T.O., Lavery, L.A. & Agard, D.A. Substrate binding drives large-scale conformational changes in the Hsp90 molecular chaperone. Mol. Cell 42, 96–105 (2011).
Dickey, C.A. et al. The high-affinity HSP90-CHIP complex recognizes and selectively degrades phosphorylated tau client proteins. J. Clin. Invest. 117, 648–658 (2007).
Echeverria, P.C. & Picard, D. Molecular chaperones, essential partners of steroid hormone receptors for activity and mobility. Biochim. Biophys. Acta 1803, 641–649 (2010).
Falsone, S.F., Kungl, A.J., Rek, A., Cappai, R. & Zangger, K. The molecular chaperone Hsp90 modulates intermediate steps of amyloid assembly of the Parkinson-related protein α-synuclein. J. Biol. Chem. 284, 31190–31199 (2009).
Dekki, N. et al. Transthyretin binds to glucose-regulated proteins and is subjected to endocytosis by the pancreatic β-cell. Cell. Mol. Life Sci. 69, 1733–1743 (2012).
Karagöz, G.E. et al. Hsp90-Tau complex reveals molecular basis for specificity in chaperone action. Cell 156, 963–974 (2014).
Benson, M.D. & Uemichi, T. Transthyretin amyloidosis. Amyloid 3, 44–56 (1996).
Sato, T. et al. Endoplasmic reticulum quality control regulates the fate of transthyretin variants in the cell. EMBO J. 26, 2501–2512 (2007).
Chen, J.J. et al. Endoplasmic reticulum proteostasis influences the oligomeric state of an amyloidogenic protein secreted from mammalian cells. Cell. Chem. Biol. 23, 1282–1293 (2016).
Planté-Bordeneuve, V. & Said, G. Transthyretin related familial amyloid polyneuropathy. Curr. Opin. Neurol. 13, 569–573 (2000).
Connors, L.H., Lim, A., Prokaeva, T., Roskens, V.A. & Costello, C.E. Tabulation of human transthyretin (TTR) variants, 2003. Amyloid 10, 160–184 (2003).
Bekircan-Kurt, C.E., Güneş, N., Yılmaz, A., Erdem-Özdamar, S. & Tan, E. Three Turkish families with different transthyretin mutations. Neuromuscul. Disord. 25, 686–692 (2015).
Jiang, X. et al. An engineered transthyretin monomer that is nonamyloidogenic, unless it is partially denatured. Biochemistry 40, 11442–11452 (2001).
Blake, C.C., Geisow, M.J., Swan, I.D., Rerat, C. & Rerat, B. Strjcture of human plasma prealbumin at 2-5 Å resolution. A preliminary report on the polypeptide chain conformation, quaternary structure and thyroxine binding. J. Mol. Biol. 88, 1–12 (1974).
Johnson, S.M., Connelly, S., Fearns, C., Powers, E.T. & Kelly, J.W. The transthyretin amyloidoses: from delineating the molecular mechanism of aggregation linked to pathology to a regulatory-agency-approved drug. J. Mol. Biol. 421, 185–203 (2012).
Reixach, N., Deechongkit, S., Jiang, X., Kelly, J.W. & Buxbaum, J.N. Tissue damage in the amyloidoses: transthyretin monomers and nonnative oligomers are the major cytotoxic species in tissue culture. Proc. Natl. Acad. Sci. USA 101, 2817–2822 (2004).
Quintas, A., Saraiva, M.J. & Brito, R.M. The tetrameric protein transthyretin dissociates to a non-native monomer in solution. A novel model for amyloidogenesis. J. Biol. Chem. 274, 32943–32949 (1999).
Lim, K.H. et al. Structural changes associated with transthyretin misfolding and amyloid formation revealed by solution and solid-state NMR. Biochemistry 55, 1941–1944 (2016).
Lim, K.H., Dyson, H.J., Kelly, J.W. & Wright, P.E. Localized structural fluctuations promote amyloidogenic conformations in transthyretin. J. Mol. Biol. 425, 977–988 (2013).
Bourgault, S. et al. Mechanisms of transthyretin cardiomyocyte toxicity inhibition by resveratrol analogs. Biochem. Biophys. Res. Commun. 410, 707–713 (2011).
Neudecker, P. et al. Structure of an intermediate state in protein folding and aggregation. Science 336, 362–366 (2012).
Sreedhar, A.S., Kalmár, E., Csermely, P. & Shen, Y.F. Hsp90 isoforms: functions, expression and clinical importance. FEBS Lett. 562, 11–15 (2004).
Park, S.J., Borin, B.N., Martinez-Yamout, M.A. & Dyson, H.J. The client protein p53 adopts a molten globule-like state in the presence of Hsp90. Nat. Struct. Mol. Biol. 18, 537–541 (2011).
Hagn, F. et al. Structural analysis of the interaction between Hsp90 and the tumor suppressor protein p53. Nat. Struct. Mol. Biol. 18, 1086–1093 (2011).
Lorenz, O.R. et al. Modulation of the Hsp90 chaperone cycle by a stringent client protein. Mol. Cell 53, 941–953 (2014).
Krukenberg, K.A., Förster, F., Rice, L.M., Sali, A. & Agard, D.A. Multiple conformations of E. coli Hsp90 in solution: insights into the conformational dynamics of Hsp90. Structure 16, 755–765 (2008).
Karagöz, G.E. et al. N-terminal domain of human Hsp90 triggers binding to the cochaperone p23. Proc. Natl. Acad. Sci. USA 108, 580–585 (2011).
Rosenzweig, R. & Kay, L.E. Bringing dynamic molecular machines into focus by methyl-TROSY NMR. Annu. Rev. Biochem. 83, 291–315 (2014).
Liu, W.M. et al. A pH-sensitive, colorful, lanthanide-chelating paramagnetic NMR probe. J. Am. Chem. Soc. 134, 17306–17313 (2012).
Keizers, P.H., Saragliadis, A., Hiruma, Y., Overhand, M. & Ubbink, M. Design, synthesis, and evaluation of a lanthanide chelating protein probe: CLaNP-5 yields predictable paramagnetic effects independent of environment. J. Am. Chem. Soc. 130, 14802–14812 (2008).
Pintacuda, G., John, M., Su, X.C. & Otting, G. NMR structure determination of protein-ligand complexes by lanthanide labeling. Acc. Chem. Res. 40, 206–212 (2007).
Keizers, P.H. & Ubbink, M. Paramagnetic tagging for protein structure and dynamics analysis. Prog. Nucl. Magn. Reson. Spectrosc. 58, 88–96 (2011).
Jackson, S.E. Hsp90: structure and function. Top. Curr. Chem. 328, 155–240 (2013).
Vaughan, C.K. et al. Structure of an Hsp90-Cdc37-Cdk4 complex. Mol. Cell 23, 697–707 (2006).
Chakraborty, R., Muchtar, E. & Gertz, M.A. Newer therapies for amyloid cardiomyopathy. Curr. Heart Fail. Rep. 13, 237–246 (2016).
Verba, K.A. et al. Atomic structure of Hsp90-Cdc37-Cdk4 reveals that Hsp90 traps and stabilizes an unfolded kinase. Science 352, 1542–1547 (2016).
Meyer, P. et al. Structural basis for recruitment of the ATPase activator Aha1 to the Hsp90 chaperone machinery. EMBO J. 23, 511–519 (2004).
Ali, M.M. et al. Crystal structure of an Hsp90-nucleotide-p23/Sba1 closed chaperone complex. Nature 440, 1013–1017 (2006).
Zhang, M., Kadota, Y., Prodromou, C., Shirasu, K. & Pearl, L.H. Structural basis for assembly of Hsp90-Sgt1-CHORD protein complexes: implications for chaperoning of NLR innate immunity receptors. Mol. Cell 39, 269–281 (2010).
Carman, A., Kishinevsky, S., Koren, J. III, Lou, W. & Chiosis, G. Chaperone-dependent neurodegeneration: a molecular perspective on therapeutic intervention. J. Alzheimers Dis. Parkinsonism 2013, 007 (2013).
Liu, K., Kelly, J.W. & Wemmer, D.E. Native state hydrogen exchange study of suppressor and pathogenic variants of transthyretin. J. Mol. Biol. 320, 821–832 (2002).
Tugarinov, V., Kanelis, V. & Kay, L.E. Isotope labeling strategies for the study of high-molecular-weight proteins by solution NMR spectroscopy. Nat. Protoc. 1, 749–754 (2006).
Bax, A. Multidimensional nuclear-magnetic-resonance methods for protein studies. Curr. Opin. Struct. Biol. 4, 738–744 (1994).
Sattler, M., Schleucher, J. & Griesinger, C. Heteronuclear multidimensional NMR experiments for the structure determination of proteins in solution employing pulsed field gradients. Prog. Nucl. Magn. Reson. Spectrosc. 34, 93–158 (1999).
Bax, A., Clore, G.M. & Gronenborn, A.M. H-1-H-1 correlation via isotropic mixing of C-13 magnetization, a new 3-dimensional approach for assigning H-1 and C-13 spectra of C-13-enriched proteins. J. Magn. Reson. 88, 425–431 (1990).
Wuthrich, K. NMR of Proteins and Nucleic Acids (Wiley Interscience, 1986).
Marion, D. et al. Overcoming the overlap problem in the assignment of 1H NMR spectra of larger proteins by use of three-dimensional heteronuclear 1H-15N Hartmann-Hahn-multiple quantum coherence and nuclear Overhauser-multiple quantum coherence spectroscopy: application to interleukin 1 beta. Biochemistry 28, 6150–6156 (1989).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Shen, Y. & Bax, A. Protein backbone and sidechain torsion angles predicted from NMR chemical shifts using artificial neural networks. J. Biomol. NMR 56, 227–241 (2013).
Berjanskii, M.V. & Wishart, D.S. A simple method to predict protein flexibility using secondary chemical shifts. J. Am. Chem. Soc. 127, 14970–14971 (2005).
Güntert, P. Automated NMR structure calculation with CYANA. Methods Mol. Biol. 278, 353–378 (2004).
Bhattacharya, A., Tejero, R. & Montelione, G.T. Evaluating protein structures determined by structural genomics consortia. Proteins 66, 778–795 (2007).
Tugarinov, V., Hwang, P.M. & Kay, L.E. Nuclear magnetic resonance spectroscopy of high-molecular-weight proteins. Annu. Rev. Biochem. 73, 107–146 (2004).
Vranken, W.F. et al. The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59, 687–696 (2005).
Petoukhov, M.V. et al. New developments in the ATSAS program package for small-angle scattering data analysis. J. Appl. Crystallogr. 45, 342–350 (2012).
Acknowledgements
We thank M. Mizuguchi (University of Toyama, Toyama, Japan) for the transthyretin plasmid; C.A. Dickey (University of South Florida, Tampa, Florida, USA) for the Hsp90 plasmid; and C.A. Dickey, B.A. Nordhues and S.G. Rüdiger for useful discussions. We are grateful to M. Ubbink (Leiden University, Leiden, the Netherlands) for the CLanP-7 lanthanide tag, to P. Wysoczanski for help with NMR spectroscopy experiments recorded for Hsp90's M domain, and to A. Pérez-Lara for help with ITC experiments. This work was supported by the Alexander von Humboldt Foundation (fellowship to J.H.K.), the European Commission (Marie Curie Intra-European fellowship, project number 626526 to J.O.), the Fulbright Program (scholarship to B.J.C.) and the European Community's Seventh Framework Programme (FP7/2007-2013) under BioStruct-X (grant agreement 283570 to M.Z.).
Author information
Authors and Affiliations
Contributions
J.H.K. performed NMR spectroscopy and biochemical experiments on TTR variants, as well as structure calculations. J.O. performed NMR spectroscopy, SAXS and ITC experiments on Hsp90. B.J.C. and J.O. produced Hsp90 mutants for the assignment of isoleucine methyl groups. J.H.K., J.O. and M.Z. designed experiments. J.H.K., J.O. and M.Z. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 The structure of the misfolded monomeric conformation of TTR is well defined.
(a) Comparison of 1H-15N NMR spectra of M-TTR at ambient pressure (green) and 500 bar (blue). Working at 500 bar improves the spectral quality. (b) Experimentally observed NOEs (green dotted lines) are visualized on the 3D structure of M-TTR (blue).
Supplementary Figure 2 Reproducibility in calorimetric titrations of Hsp90 with M-TTR.
From left to right and top to bottom, raw data resulting from the titration of 25x1.5 μl aliquots of 1671 μM of M-TTR into 25 μM of Hsp90 (a), 783 μM of M-TTR into 30 μM of Hsp90 (b), 520 μM of M-TTR into 30 μM of Hsp90 (c), 313 μM of M-TTR into 25,5 μM of Hsp90 (d) and 166 μM of M-TTR into 22,75 μM of Hsp90 (e) are shown. The biphasic thermodynamic behavior was present in all tested conditions.
Supplementary Figure 3 Nucleotide binding does not promote allosteric changes in the Hsp90–M-TTR complex.
(a) As observed from the P(r) distribution obtained from SAXS scattering data, ADP binding does not change the global extended conformation of the Hsp90/M-TTR complex. While Hsp90 shows Rg= 6.34 ± 0.16 nm and DMAX= 20.94 ± 0.52 nm, the Hsp90+M-TTR complex has Rg= 6.58 ± 0.44 nm and DMAX= 21.77 ± 0.71 nm. (b) Comparison of NMR signal intensities of the methyl signals of M-TTR/Hsp90 in the absence and presence of 2 mM ADP. Samples contained 0.07 mM 13C-labeled and methyl-protonated M-TTR in 50 mM MES, pH 7, 100 mM NaCl, 5 mM DTT, 1 mM MgSO4, 0.1 mM DSS, and a 2-fold (blue) or 4-fold (green) molar excess of Hsp90.
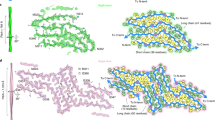
Supplementary Figure 4 Hsp90 uses several binding interfaces to bind M-TTR.
Residues in the N- and M-domain of Hsp90, which show the strongest PCSs in the presence of M-TTR, are highlighted in orange in the 3D structure. Hsp90’s N-domain is colored in light blue, M-domain in red and C-terminal domain in light green. The charged linker, which connects Hsp90’s N and M domain, is represented by a grey line.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Table 1 (PDF 868 kb)
Rights and permissions
About this article
Cite this article
Oroz, J., Kim, J., Chang, B. et al. Mechanistic basis for the recognition of a misfolded protein by the molecular chaperone Hsp90. Nat Struct Mol Biol 24, 407–413 (2017). https://doi.org/10.1038/nsmb.3380
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3380
This article is cited by
-
Molecular chaperones and their denaturing effect on client proteins
Journal of Biomolecular NMR (2021)
-
Probing conformational changes of monomeric transthyretin with second derivative fluorescence
Scientific Reports (2019)
-
Structure and pro-toxic mechanism of the human Hsp90/PPIase/Tau complex
Nature Communications (2018)
-
Structural ensemble-based docking simulation and biophysical studies discovered new inhibitors of Hsp90 N-terminal domain
Scientific Reports (2018)