Abstract
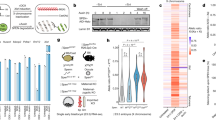
The nuclear long noncoding RNA (lncRNA) Xist ensures X-chromosome inactivation (XCI) in female placental mammals. Although Xist is one of the most intensively studied lncRNAs, the mechanisms associated with its capacity to trigger chromosome-wide gene silencing, the formation of facultative heterochromatin and an unusual 3D conformation of the inactive X chromosome (Xi) have remained elusive. Now researchers have identified novel functional partners of Xist in a series of breakthrough studies, using unbiased techniques to isolate Xist-bound proteins, as well as forward genetic screens. In addition, important insights into the 3D organization of Xi and its relation to gene expression have been obtained. In this Review, we discuss how this new information is providing a recipe for deciphering XCI mechanisms by which a multitasking RNA can structurally and functionally transform an active chromosome into uniquely organized facultative heterochromatin.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Galupa, R. & Heard, E. X-chromosome inactivation: new insights into cis and trans regulation. Curr. Opin. Genet. Dev. 31, 57–66 (2015).
Marahrens, Y., Panning, B., Dausman, J., Strauss, W. & Jaenisch, R. Xist-deficient mice are defective in dosage compensation but not spermatogenesis. Genes Dev. 11, 156–166 (1997).
Penny, G.D., Kay, G.F., Sheardown, S.A., Rastan, S. & Brockdorff, N. Requirement for Xist in X chromosome inactivation. Nature 379, 131–137 (1996).
van Bemmel, J.G., Mira-Bontenbal, H. & Gribnau, J. Cis- and trans-regulation in X inactivation. Chromosoma 125, 41–50 (2016).
Escamilla-Del-Arenal, M., da Rocha, S.T. & Heard, E. Evolutionary diversity and developmental regulation of X-chromosome inactivation. Hum. Genet. 130, 307–327 (2011).
Splinter, E. et al. The inactive X chromosome adopts a unique three-dimensional conformation that is dependent on Xist RNA. Genes Dev. 25, 1371–1383 (2011).
Brown, S.D. XIST and the mapping of the X chromosome inactivation centre. BioEssays 13, 607–612 (1991).
Nesterova, T.B. et al. Characterization of the genomic Xist locus in rodents reveals conservation of overall gene structure and tandem repeats but rapid evolution of unique sequence. Genome Res. 11, 833–849 (2001).
Yen, Z.C., Meyer, I.M., Karalic, S. & Brown, C.J. A cross-species comparison of X-chromosome inactivation in Eutheria. Genomics 90, 453–463 (2007).
Wutz, A., Rasmussen, T.P. & Jaenisch, R. Chromosomal silencing and localization are mediated by different domains of Xist RNA. Nat. Genet. 30, 167–174 (2002).
da Rocha, S.T. et al. Jarid2 is implicated in the initial Xist-induced targeting of PRC2 to the inactive X chromosome. Mol. Cell 53, 301–316 (2014).
Kohlmaier, A. et al. A chromosomal memory triggered by Xist regulates histone methylation in X inactivation. PLoS Biol. 2, E171 (2004).
Zhao, J., Sun, B.K., Erwin, J.A., Song, J.J. & Lee, J.T. Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 322, 750–756 (2008).
Brockdorff, N. et al. The product of the mouse Xist gene is a 15 kb inactive X-specific transcript containing no conserved ORF and located in the nucleus. Cell 71, 515–526 (1992).
Brown, C.J. et al. A gene from the region of the human X inactivation centre is expressed exclusively from the inactive X chromosome. Nature 349, 38–44 (1991).
Hasegawa, Y. et al. The matrix protein hnRNP U is required for chromosomal localization of Xist RNA. Dev. Cell 19, 469–476 (2010).
Jeon, Y. & Lee, J.T. YY1 tethers Xist RNA to the inactive X nucleation center. Cell 146, 119–133 (2011).
Maenner, S. et al. 2-D structure of the A region of Xist RNA and its implication for PRC2 association. PLoS Biol. 8, e1000276 (2010).
Sarma, K. et al. ATRX directs binding of PRC2 to Xist RNA and polycomb targets. Cell 159, 869–883 (2014).
McHugh, C.A., Russell, P. & Guttman, M. Methods for comprehensive experimental identification of RNA-protein interactions. Genome Biol. 15, 203 (2014).
Duszczyk, M.M., Wutz, A., Rybin, V. & Sattler, M. The Xist RNA A-repeat comprises a novel AUCG tetraloop fold and a platform for multimerization. RNA 17, 1973–1982 (2011).
Brockdorff, N. Noncoding RNA and Polycomb recruitment. RNA 19, 429–442 (2013).
Davidovich, C. et al. Toward a consensus on the binding specificity and promiscuity of PRC2 for RNA. Mol. Cell 57, 552–558 (2015).
Chu, C. et al. Systematic discovery of Xist RNA binding proteins. Cell 161, 404–416 (2015). This study revealed the Xist -protein interactome. The authors used a novel proteomic method (CHIRP-MS) to identify 81 Xist RBPs, including HNRNPU, HNRNPK and SPEN, which are involved in Xist -mediated silencing.
McHugh, C.A. et al. The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature 521, 232–236 (2015). This study was the first to use a UV-cross-linking-based proteomic method (RAP-MS) to reveal ten high-confidence Xist RBPs, including SPEN/SHARP. A mechanistic link between SPEN/SHARP and the SMRT/HDAC3 deacetylase complex in regulating XCI was demonstrated.
Minajigi, A. et al. A comprehensive Xist interactome reveals cohesin repulsion and an RNA-directed chromosome conformation. Science 349, aab2276 (2015). In this study, the authors used iDRiP to identify multiple Xist interactors, including architectural proteins. The paper describes a role for Xist in Xi-specific cohesin reorganization and chromosome conformation.
Wutz, A. & Jaenisch, R. A shift from reversible to irreversible X inactivation is triggered during ES cell differentiation. Mol. Cell 5, 695–705 (2000).
Agrelo, R. et al. SATB1 defines the developmental context for gene silencing by Xist in lymphoma and embryonic cells. Dev. Cell 16, 507–516 (2009).
Nechanitzky, R., Dávila, A., Savarese, F., Fietze, S. & Grosschedl, R. Satb1 and Satb2 are dispensable for X chromosome inactivation in mice. Dev. Cell 23, 866–871 (2012).
Wang, J. et al. Imprinted X inactivation maintained by a mouse Polycomb group gene. Nat. Genet. 28, 371–375 (2001).
Kalantry, S. et al. The Polycomb group protein Eed protects the inactive X-chromosome from differentiation-induced reactivation. Nat. Cell Biol. 8, 195–202 (2006).
Blewitt, M.E. et al. SmcHD1, containing a structural-maintenance-of-chromosomes hinge domain, has a critical role in X inactivation. Nat. Genet. 40, 663–669 (2008).
Gendrel, A.V. et al. Smchd1-dependent and -independent pathways determine developmental dynamics of CpG island methylation on the inactive X chromosome. Dev. Cell 23, 265–279 (2012).
Moindrot, B. et al. A pooled shRNA screen identifies Rbm15, Spen, and Wtap as factors required for Xist RNA-mediated silencing. Cell Rep. 12, 562–572 (2015). The RNAi-based forward screen used in this study identified novel factors involved in Xist -mediated gene silencing. The top hits were SPEN and RBM15, both RBPs containing a SPOC domain, and WTAP, a subunit of the m6A RNA methyltransferase complex.
Monfort, A. et al. Identification of Spen as a crucial factor for Xist function through forward genetic screening in haploid embryonic stem cells. Cell Rep. 12, 554–561 (2015). The authors of this study used a viral gene-trap screen on mouse haploid ESCs and successfully identified factors involved in in Xist -mediated gene silencing, including the SPEN RBP.
Rego, A., Sinclair, P.B., Tao, W., Kireev, I. & Belmont, A.S. The facultative heterochromatin of the inactive X chromosome has a distinctive condensed ultrastructure. J. Cell Sci. 121, 1119–1127 (2008).
Teller, K. et al. A top-down analysis of Xa- and Xi-territories reveals differences of higher order structure at ≥ 20 Mb genomic length scales. Nucleus 2, 465–477 (2011).
Smeets, D. et al. Three-dimensional super-resolution microscopy of the inactive X chromosome territory reveals a collapse of its active nuclear compartment harboring distinct Xist RNA foci. Epigenetics Chromatin 7, 8 (2014).
Chaumeil, J., Le Baccon, P., Wutz, A. & Heard, E. A novel role for Xist RNA in the formation of a repressive nuclear compartment into which genes are recruited when silenced. Genes Dev. 20, 2223–2237 (2006).
Clemson, C.M., Hall, L.L., Byron, M., McNeil, J. & Lawrence, J.B. The X chromosome is organized into a gene-rich outer rim and an internal core containing silenced nongenic sequences. Proc. Natl. Acad. Sci. USA 103, 7688–7693 (2006).
Nora, E.P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Darrow, E.M. et al. Deletion of DXZ4 on the human inactive X chromosome alters higher-order genome architecture. Proc. Natl. Acad. Sci. USA 113, E4504–E4512 (2016). In this study, the authors dissected the role of the DXZ4 boundary element in maintaining the megadomain structure of the human Xi. Deletion of DXZ4 revealed the importance of the element for the 3D Xi conformation, but not for Xi silencing.
Deng, X. et al. Bipartite structure of the inactive mouse X chromosome. Genome Biol. 16, 152 (2015).
Giorgetti, L. et al. Structural organization of the inactive X chromosome in the mouse. Nature 535, 575–579 (2016). This study—the most comprehensive study of the Xi structure to date—reveals organization into two megadomains and a general lack of TAD segmentation, except at regions of clustered escapees. Xist RNA coating is required for the formation of Xi's bipartite structure, whereas DXZ4 boundary deletion disrupts this structure, with little effect on Xi silencing.
Rao, S.S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Wang, S. et al. Spatial organization of chromatin domains and compartments in single chromosomes. Science 353, 598–602 (2016).
Siomi, H., Matunis, M.J., Michael, W.M. & Dreyfuss, G. The pre-mRNA binding K protein contains a novel evolutionarily conserved motif. Nucleic Acids Res. 21, 1193–1198 (1993).
Ong, S.E. & Mann, M. Stable isotope labeling by amino acids in cell culture for quantitative proteomics. Methods Mol. Biol. 359, 37–52 (2007).
Ariyoshi, M. & Schwabe, J.W. A conserved structural motif reveals the essential transcriptional repression function of Spen proteins and their role in developmental signaling. Genes Dev. 17, 1909–1920 (2003).
Cooper, S. et al. Jarid2 binds mono-ubiquitylated H2A lysine 119 to mediate crosstalk between Polycomb complexes PRC1 and PRC2. Nat. Commun. 7, 13661 (2016).
Kolodziej, P.A., Jan, L.Y. & Jan, Y.N. Mutations that affect the length, fasciculation, or ventral orientation of specific sensory axons in the Drosophila embryo. Neuron 15, 273–286 (1995).
Arieti, F. et al. The crystal structure of the Split End protein SHARP adds a new layer of complexity to proteins containing RNA recognition motifs. Nucleic Acids Res. 42, 6742–6752 (2014).
Shi, Y. et al. Sharp, an inducible cofactor that integrates nuclear receptor repression and activation. Genes Dev. 15, 1140–1151 (2001).
You, S.H. et al. Nuclear receptor co-repressors are required for the histone-deacetylase activity of HDAC3 in vivo. Nat. Struct. Mol. Biol. 20, 182–187 (2013).
Lu, Z. et al. RNA duplex map in living cells reveals higher-order transcriptome structure. Cell 165, 1267–1279 (2016).
Kuroda, K. et al. Regulation of marginal zone B cell development by MINT, a suppressor of Notch/RBP-J signaling pathway. Immunity 18, 301–312 (2003).
Gallardo, M. et al. hnRNP K is a haploinsufficient tumor suppressor that regulates proliferation and differentiation programs in hematologic malignancies. Cancer Cell 28, 486–499 (2015).
Patil, D.P. et al. m6A RNA methylation promotes XIST-mediated transcriptional repression. Nature 537, 369–373 (2016). This was the first study to report a functional link between m6A RNA methylation and Xist RNA function.
Lee, J.H. & Skalnik, D.G. Rbm15-Mkl1 interacts with the Setd1b histone H3-Lys4 methyltransferase via a SPOC domain that is required for cytokine-independent proliferation. PLoS One 7, e42965 (2012).
Zolotukhin, A.S. et al. Nuclear export factor RBM15 facilitates the access of DBP5 to mRNA. Nucleic Acids Res. 37, 7151–7162 (2009).
Ma, Z. et al. Fusion of two novel genes, RBM15 and MKL1, in the t(1;22)(p13;q13) of acute megakaryoblastic leukemia. Nat. Genet. 28, 220–221 (2001).
Horiuchi, K. et al. Identification of Wilms' tumor 1-associating protein complex and its role in alternative splicing and the cell cycle. J. Biol. Chem. 288, 33292–33302 (2013).
Little, N.A., Hastie, N.D. & Davies, R.C. Identification of WTAP, a novel Wilms' tumour 1-associating protein. Hum. Mol. Genet. 9, 2231–2239 (2000).
Horiuchi, K. et al. Wilms' tumor 1-associating protein regulates G2/M transition through stabilization of cyclin A2 mRNA. Proc. Natl. Acad. Sci. USA 103, 17278–17283 (2006).
Ping, X.L. et al. Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res. 24, 177–189 (2014).
Schwartz, S. et al. Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5′ sites. Cell Rep. 8, 284–296 (2014).
Meyer, K.D. & Jaffrey, S.R. The dynamic epitranscriptome: N6-methyladenosine and gene expression control. Nat. Rev. Mol. Cell Biol. 15, 313–326 (2014).
Yamada, N. et al. Xist exon 7 contributes to the stable localization of Xist RNA on the inactive X-chromosome. PLoS Genet. 11, e1005430 (2015).
Pullirsch, D. et al. The Trithorax group protein Ash2l and Saf-A are recruited to the inactive X chromosome at the onset of stable X inactivation. Development 137, 935–943 (2010).
Kolpa, H.J., Fackelmayer, F.O. & Lawrence, J.B. SAF-A requirement in anchoring XIST RNA to chromatin varies in transformed and primary cells. Dev. Cell 39, 9–10 (2016).
Sakaguchi, T. et al. Control of chromosomal localization of Xist by hnRNP U family molecules. Dev. Cell 39, 11–12 (2016).
Olins, A.L., Rhodes, G., Welch, D.B., Zwerger, M. & Olins, D.E. Lamin B receptor: multi-tasking at the nuclear envelope. Nucleus 1, 53–70 (2010).
Chen, C.K. et al. Xist recruits the X chromosome to the nuclear lamina to enable chromosome-wide silencing. Science 354, 468–472 (2016).
Yang, F. et al. The lncRNA Firre anchors the inactive X chromosome to the nucleolus by binding CTCF and maintains H3K27me3 methylation. Genome Biol. 16, 52 (2015).
Zhang, L.F., Huynh, K.D. & Lee, J.T. Perinucleolar targeting of the inactive X during S phase: evidence for a role in the maintenance of silencing. Cell 129, 693–706 (2007).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Chadwick, B.P. DXZ4 chromatin adopts an opposing conformation to that of the surrounding chromosome and acquires a novel inactive X-specific role involving CTCF and antisense transcripts. Genome Res. 18, 1259–1269 (2008).
Horakova, A.H. et al. The mouse DXZ4 homolog retains Ctcf binding and proximity to Pls3 despite substantial organizational differences compared to the primate macrosatellite. Genome Biol. 13, R70 (2012).
Dixon, J.R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Merkenschlager, M. & Nora, E.P. CTCF and cohesin in genome folding and transcriptional gene regulation. Annu. Rev. Genomics Hum. Genet. 17, 17–43 (2016).
Fang, R., Moss, W.N., Rutenberg-Schoenberg, M. & Simon, M.D. Probing Xist RNA structure in cells using Targeted Structure-Seq. PLoS Genet. 11, e1005668 (2015).
Smola, M.J. et al. SHAPE reveals transcript-wide interactions, complex structural domains, and protein interactions across the Xist lncRNA in living cells. Proc. Natl. Acad. Sci. USA 113, 10322–10327 (2016).
Acknowledgements
We thank I. Pinheiro, E. Nora and S.F. de Almeida for their critical reading of the manuscript. This work was supported by Fundação para a Ciência e Tecnologia (grants PTDC/BEX-BCM/2612/2014 and IF/00242/2014 to S.T.d.R.). Research in the Heard laboratory was supported by Labex DEEP (ANR-11-LBX-0044), part of the IDEX Idex PSL (ANR-10-IDEX-0001-02 PSL), ERC Advanced Investigator award 250367 and “La Ligue Contre le Cancer” (Equipe Labelisée).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
da Rocha, S., Heard, E. Novel players in X inactivation: insights into Xist-mediated gene silencing and chromosome conformation. Nat Struct Mol Biol 24, 197–204 (2017). https://doi.org/10.1038/nsmb.3370
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3370
This article is cited by
-
Long noncoding RNA ADAMTS9-AS1 represses ferroptosis of endometrial stromal cells by regulating the miR-6516-5p/GPX4 axis in endometriosis
Scientific Reports (2022)
-
RNA proximity sequencing data and analysis pipeline from a human neuroblastoma nuclear transcriptome
Scientific Data (2020)
-
Disruption of ATRX-RNA interactions uncovers roles in ATRX localization and PRC2 function
Nature Communications (2020)
-
Regulation of gene expression by cis-acting long non-coding RNAs
Nature Reviews Genetics (2020)
-
Global chromatin conformation differences in the Drosophila dosage compensated chromosome X
Nature Communications (2019)