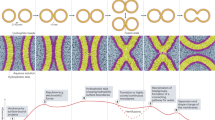
Abstract
SOS is a key activator of the small GTPase Ras. In cells, SOS-Ras signaling is thought to be initiated predominantly by membrane recruitment of SOS via the adaptor Grb2 and balanced by rapidly reversible Grb2-SOS binding kinetics. However, SOS has multiple protein and lipid interactions that provide linkage to the membrane. In reconstituted-membrane experiments, these Grb2-independent interactions were sufficient to retain human SOS on the membrane for many minutes, during which a single SOS molecule could processively activate thousands of Ras molecules. These observations raised questions concerning how receptors maintain control of SOS in cells and how membrane-recruited SOS is ultimately released. We addressed these questions in quantitative assays of reconstituted SOS-deficient chicken B-cell signaling systems combined with single-molecule measurements in supported membranes. These studies revealed an essentially one-way trafficking process in which membrane-recruited SOS remains trapped on the membrane and continuously activates Ras until being actively removed via endocytosis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Campbell, S.L., Khosravi-Far, R., Rossman, K.L., Clark, G.J. & Der, C.J. Increasing complexity of Ras signaling. Oncogene 17, 1395–1413 (1998).
Chang, L. & Karin, M. Mammalian MAP kinase signalling cascades. Nature 410, 37–40 (2001).
Karnoub, A.E. & Weinberg, R.A. Ras oncogenes: split personalities. Nat. Rev. Mol. Cell Biol. 9, 517–531 (2008).
Stephen, A.G., Esposito, D., Bagni, R.K. & McCormick, F. Dragging ras back in the ring. Cancer Cell 25, 272–281 (2014).
McCormick, F. Signal transduction: how receptors turn Ras on. Nature 363, 15–16 (1993).
Findlay, G.M. & Pawson, T. How is SOS activated? Let us count the ways. Nat. Struct. Mol. Biol. 15, 538–540 (2008).
Groves, J.T. & Kuriyan, J. Molecular mechanisms in signal transduction at the membrane. Nat. Struct. Mol. Biol. 17, 659–665 (2010).
Das, J. et al. Digital signaling and hysteresis characterize ras activation in lymphoid cells. Cell 136, 337–351 (2009).
Jun, J.E., Rubio, I. & Roose, J.P. Regulation of ras exchange factors and cellular localization of ras activation by lipid messengers in T cells. Front. Immunol. 4, 239 (2013).
Egan, S.E. et al. Association of Sos Ras exchange protein with Grb2 is implicated in tyrosine kinase signal transduction and transformation. Nature 363, 45–51 (1993).
Genot, E. & Cantrell, D.A. Ras regulation and function in lymphocytes. Curr. Opin. Immunol. 12, 289–294 (2000).
Kortum, R.L. et al. Targeted Sos1 deletion reveals its critical role in early T-cell development. Proc. Natl. Acad. Sci. USA 108, 12407–12412 (2011).
Baltanás, F.C. et al. Functional redundancy of Sos1 and Sos2 for lymphopoiesis and organismal homeostasis and survival. Mol. Cell. Biol. 33, 4562–4578 (2013).
Wang, D.Z. et al. Mutation in Sos1 dominantly enhances a weak allele of the EGFR, demonstrating a requirement for Sos1 in EGFR signaling and development. Genes Dev. 11, 309–320 (1997).
Findlay, G.M. et al. Interaction domains of Sos1/Grb2 are finely tuned for cooperative control of embryonic stem cell fate. Cell 152, 1008–1020 (2013).
Buday, L. & Downward, J. Epidermal growth factor regulates p21ras through the formation of a complex of receptor, Grb2 adapter protein, and Sos nucleotide exchange factor. Cell 73, 611–620 (1993).
Gale, N.W., Kaplan, S., Lowenstein, E.J., Schlessinger, J. & Bar-Sagi, D. Grb2 mediates the EGF-dependent activation of guanine nucleotide exchange on Ras. Nature 363, 88–92 (1993).
Li, N. et al. Guanine-nucleotide-releasing factor hSos1 binds to Grb2 and links receptor tyrosine kinases to Ras signalling. Nature 363, 85–88 (1993).
Rozakis-Adcock, M., Fernley, R., Wade, J., Pawson, T. & Bowtell, D. The SH2 and SH3 domains of mammalian Grb2 couple the EGF receptor to the Ras activator mSos1. Nature 363, 83–85 (1993).
Waterman, H. et al. A mutant EGF-receptor defective in ubiquitylation and endocytosis unveils a role for Grb2 in negative signaling. EMBO J. 21, 303–313 (2002).
Chardin, P. et al. Human Sos1: a guanine nucleotide exchange factor for Ras that binds to GRB2. Science 260, 1338–1343 (1993).
Gureasko, J. et al. Membrane-dependent signal integration by the Ras activator Son of sevenless. Nat. Struct. Mol. Biol. 15, 452–461 (2008).
Gureasko, J. et al. Role of the histone domain in the autoinhibition and activation of the Ras activator Son of Sevenless. Proc. Natl. Acad. Sci. USA 107, 3430–3435 (2010).
Yadav, K.K. & Bar-Sagi, D. Allosteric gating of Son of sevenless activity by the histone domain. Proc. Natl. Acad. Sci. USA 107, 3436–3440 (2010).
Margarit, S.M. et al. Structural evidence for feedback activation by Ras.GTP of the Ras-specific nucleotide exchange factor SOS. Cell 112, 685–695 (2003).
Roberts, A.E. et al. Germline gain-of-function mutations in SOS1 cause Noonan syndrome. Nat. Genet. 39, 70–74 (2007).
Tumurkhuu, M., Saitoh, M., Takita, J., Mizuno, Y. & Mizuguchi, M. A novel SOS1 mutation in Costello/CFC syndrome affects signaling in both RAS and PI3K pathways. J. Recept. Signal Transduct. Res. 33, 124–128 (2013).
Kholodenko, B.N., Hoek, J.B. & Westerhoff, H.V. Why cytoplasmic signalling proteins should be recruited to cell membranes. Trends Cell Biol. 10, 173–178 (2000).
Karlovich, C.A. et al. In vivo functional analysis of the Ras exchange factor son of sevenless. Science 268, 576–579 (1995).
Corbalan-Garcia, S., Margarit, S.M., Galron, D., Yang, S.S. & Bar-Sagi, D. Regulation of Sos activity by intramolecular interactions. Mol. Cell. Biol. 18, 880–886 (1998).
McCollam, L. et al. Functional roles for the pleckstrin and Dbl homology regions in the Ras exchange factor Son-of-sevenless. J. Biol. Chem. 270, 15954–15957 (1995).
Wang, W. et al. The Grb2 binding domain of mSos1 is not required for downstream signal transduction. Nat. Genet. 10, 294–300 (1995).
Zhao, C., Du, G., Skowronek, K., Frohman, M.A. & Bar-Sagi, D. Phospholipase D2-generated phosphatidic acid couples EGFR stimulation to Ras activation by Sos. Nat. Cell Biol. 9, 706–712 (2007).
Roose, J.P., Mollenauer, M., Ho, M., Kurosaki, T. & Weiss, A. Unusual interplay of two types of Ras activators, RasGRP and SOS, establishes sensitive and robust Ras activation in lymphocytes. Mol. Cell. Biol. 27, 2732–2745 (2007).
Chen, R.H., Corbalan-Garcia, S. & Bar-Sagi, D. The role of the PH domain in the signal-dependent membrane targeting of Sos. EMBO J. 16, 1351–1359 (1997).
Groves, J.T., Ulman, N. & Boxer, S.G. Micropatterning fluid lipid bilayers on solid supports. Science 275, 651–653 (1997).
Groves, J.T. & Boxer, S.G. Micropattern formation in supported lipid membranes. Acc. Chem. Res. 35, 149–157 (2002).
Iversen, L. et al. Ras activation by SOS: allosteric regulation by altered fluctuation dynamics. Science 345, 50–54 (2014).
Lin, W.C. et al. H-Ras forms dimers on membrane surfaces via a protein-protein interface. Proc. Natl. Acad. Sci. USA 111, 2996–3001 (2014).
Bos, J.L., Rehmann, H. & Wittinghofer, A. GEFs and GAPs: critical elements in the control of small G proteins. Cell 129, 865–877 (2007).
Boykevisch, S. et al. Regulation of ras signaling dynamics by Sos-mediated positive feedback. Curr. Biol. 16, 2173–2179 (2006).
Hall, B.E., Yang, S.S., Boriack-Sjodin, P.A., Kuriyan, J. & Bar-Sagi, D. Structure-based mutagenesis reveals distinct functions for Ras switch 1 and switch 2 in Sos-catalyzed guanine nucleotide exchange. J. Biol. Chem. 276, 27629–27637 (2001).
Sondermann, H. et al. Structural analysis of autoinhibition in the Ras activator Son of sevenless. Cell 119, 393–405 (2004).
Kubiseski, T.J., Chook, Y.M., Parris, W.E., Rozakis-Adcock, M. & Pawson, T. High affinity binding of the pleckstrin homology domain of mSos1 to phosphatidylinositol (4,5)-bisphosphate. J. Biol. Chem. 272, 1799–1804 (1997).
Oh-hora, M., Johmura, S., Hashimoto, A., Hikida, M. & Kurosaki, T. Requirement for Ras guanine nucleotide releasing protein 3 in coupling phospholipase C-gamma2 to Ras in B cell receptor signaling. J. Exp. Med. 198, 1841–1851 (2003).
Jun, J.E., Yang, M., Chen, H., Chakraborty, A.K. & Roose, J.P. Activation of extracellular signal-regulated kinase but not of p38 mitogen-activated protein kinase pathways in lymphocytes requires allosteric activation of SOS. Mol. Cell. Biol. 33, 2470–2484 (2013).
Sondermann, H., Nagar, B., Bar-Sagi, D. & Kuriyan, J. Computational docking and solution x-ray scattering predict a membrane-interacting role for the histone domain of the Ras activator son of sevenless. Proc. Natl. Acad. Sci. USA 102, 16632–16637 (2005).
Aronheim, A. et al. Membrane targeting of the nucleotide exchange factor Sos is sufficient for activating the Ras signaling pathway. Cell 78, 949–961 (1994).
Mossman, K.D., Campi, G., Groves, J.T. & Dustin, M.L. Altered TCR signaling from geometrically repatterned immunological synapses. Science 310, 1191–1193 (2005).
Davey, A.M., Liu, W., Sohn, H.W., Brzostowski, J. & Pierce, S.K. Understanding the initiation of B cell signaling through live cell imaging. Methods Enzymol. 506, 265–290 (2012).
Balagopalan, L., Sherman, E., Barr, V.A. & Samelson, L.E. Imaging techniques for assaying lymphocyte activation in action. Nat. Rev. Immunol. 11, 21–33 (2011).
Grakoui, A. et al. The immunological synapse: a molecular machine controlling T cell activation. Science 285, 221–227 (1999).
Weber, M. et al. Phospholipase C-gamma2 and Vav cooperate within signaling microclusters to propagate B cell spreading in response to membrane-bound antigen. J. Exp. Med. 205, 853–868 (2008).
Brdicka, T. et al. Non-T cell activation linker (NTAL): a transmembrane adaptor protein involved in immunoreceptor signaling. J. Exp. Med. 196, 1617–1626 (2002).
Janssen, E., Zhu, M., Zhang, W., Koonpaew, S. & Zhang, W. LAB: a new membrane-associated adaptor molecule in B cell activation. Nat. Immunol. 4, 117–123 (2003).
Pierce, S.K. & Liu, W. The tipping points in the initiation of B cell signalling: how small changes make big differences. Nat. Rev. Immunol. 10, 767–777 (2010).
Stenmark, H. Rab GTPases as coordinators of vesicle traffic. Nat. Rev. Mol. Cell Biol. 10, 513–525 (2009).
Iwig, J.S. et al. Structural analysis of autoinhibition in the Ras-specific exchange factor RasGRP1. eLife 2, e00813 (2013).
Daley, S.R. et al. Rasgrp1 mutation increases naive T-cell CD44 expression and drives mTOR-dependent accumulation of Helios+ T cells and autoantibodies. eLife 2, e01020 (2013).
Rojas, J.M., Oliva, J.L. & Santos, E. Mammalian son of sevenless guanine nucleotide exchange factors: old concepts and new perspectives. Genes Cancer 2, 298–305 (2011).
Edelstein, A., Amodaj, N., Hoover, K., Vale, R. & Stuurman, N. Computer control of microscopes using μManager. Curr. Protoc. Mol. Biol. 92, 14.20 (2010).
Greene, A.C. et al. Spatial organization of EphA2 at the cell-cell interface modulates trans-endocytosis of ephrinA1. Biophys. J. 106, 2196–2205 (2014).
Acknowledgements
The authors thank the members of the laboratories of J.P.R., J.T.G., J. Kuriyan, and D.B.-S. for helpful comments and suggestions. The authors thank J. Kuriyan for insightful comments on the manuscript. In addition, the authors thank W.-C. Lin and L. Iversen for assistance with initial SLB experiments. This research was supported by a P01 Program grant from NIH-NIAID (AI091580 – to J.P.R. and J.T.G.). Further support came from R01-CA187318 NIH-NCI and R01-AI104789 (both to J.P.R.) and ARRA stimulus supplement GM078266 (to D.B.-S. and K.K.Y.) as well as a grant from the Danish Council for Independent Research, Natural Sciences (to S.M.C.). We thank T. Kurosaki (RIKEN) for providing wild-type and SOS1−2− DT40 B cells.
Author information
Authors and Affiliations
Contributions
S.M.C., H.-L.T., and J.E.J. performed experiments and analyzed data. S.A. and M.G.T. assisted with live-cell experiments. J.S.I. purified proteins. K.K.Y. performed COS1 cell experiments under supervision of D.B.-S. J.T.G., J.P.R., S.M.C., H.-L.T., and J.E.J. conceptualized and designed experiments. S.M.C., H.-L.T., J.E.J., J.P.R., and J.T.G. wrote the paper. J.T.G. and J.P.R. supervised the project. All authors discussed and commented on the results.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Quantification of two-dimensional Ras fluidity and surface density on SLBs and benchmarking of labeled SOS constructs.
(a) Fluorescence recovery after photobleaching (FRAP) experiment showing fluorescence from nucleotide loaded Ras at 0 s and 120s s after bleaching a circular region of a supported lipid bilayer. The recovery of the bleached signal demonstrates that Ras is laterally mobile at the membrane. The bottom panel shows the corresponding line-profiles of the fluorescence intensity across the bleached spot as indicated. FRAP analysis to verify lateral fluidity of the membrane was carried out for every supported lipid bilayer (SLB) sample employed in this work.
(b) Calibration curve to convert intensity extracted from microscopy images to Ras density. The curve was constructed from FCS measurements of Ras density (via the BODIPY labeled nucleotide) at a number of supported lipid bilayers combined with EPI imaging (plot ordinate) of the same sample positions.
(c) Labeling SOS with ATTO-647N does not perturb its enzymatic activity in the stopped-flow assay. The activities of the shown SOS constructs were benchmarked by performing stopped flow nucleotide exchange assays on Ras functionalized supported lipid bilayers with comparable Ras densities and using 100 nM of the respective SOS construct. For SOSCat, DPC and HDPC no significant alteration of the activity was observed upon labeling. The data indicates a slightly decreased activity for the labeled HDPC(R552G) construct. Of note, the minor decrease in activity for labeled HDPC(R552G) would not affect the conclusions on the data presented in Figure 3b,c. The activity for each construct was evaluated by fitting the fluorescence decay (see caption for d below) in the chase phase of the stopped flow assay with a monoexponential decay function. The decay rate observed for the respective unlabeled construct was set to 1 and the rate for the corresponding labeled variant was accordingly normalized. The bars represent the fitted decay coefficients (one trace for each condition) except for SOSCat where the bars indicate the average of two such measurements. Error bars indicate the estimated standard deviations of the fit coefficients.
(d) Processive turnover of Ras by SOSCat in the stopped flow assay. Shown intensity traces were collected immediately after the initiation of the chase phase of the stopped flow assay where unlabeled nucleotide is flowed through the reaction chamber (GDP or GTP, as indicated, see also Figure 2a, right). The shown traces were collected after first populating the bilayers with SOSCat and washing away unbound SOS molecules. It is evident that the recruited SOSCat is highly processive at the membrane surface, as evidenced by sustained Ras turnover throughout the nucleotide chase (resulting in decline of fluorescence as labeled nucleotides are exchanged for unlabeled nucleotides).
Source data for plots and graphs are available online.
Supplementary Figure 2 Analysis of surface CD69 expression in transfected Jurkat T cells and spontaneous localization of SOSCat to the membrane in resting COS-1 cells.
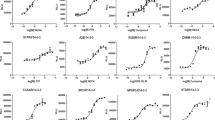
(a) Analysis of Ras-signal dependent CD69 expression in transfected Jurkat T cells. The indicated SOSCat construct or empty vector were cotransfected with a GFP-expression construct and 20 hours later CD69 expression was assessed in unstimulated Jurkat T Cells. Surface CD69 (sCD69) MFI’s (mean fluorescence intensity) are plotted as a function of GFP expression level. Source data available online.
(b) Raw data for the panel in a. Jurkat T cells were transiently transfected for 20 hours with 10 µg of wild-type or allosteric mutant (W729E) SOSCat- encoding plasmid together with 10 µg of GFP plasmid. The activity of Ras-ERK pathway was measured by FACS staining of surface CD69 (sCD69) together with GFP intensity measurement. GFP-positive cells were sub-gated into 9 fractions. Geometric mean fluorescence of CD69 level was determined for each fraction. Data shown in a and b are representative of five independent experiments.
(c) Analysis of SOSCat (red) and H-Ras(A59G D38E) (green) localization in unstimulated, transfected COS-1 cells. The panels to the right show data for a mutant impaired in allosteric Ras binding (SOSCat-L687E R688A). H-Ras(A59G D38E) is a mutant that only binds the allosteric site on SOS. The data show that spontaneous membrane-localization of SOSCat requires an intact allosteric pocket. Data in figure 2c are a reproduction of previous published work (Gureasko, J. et al., Nat. Struct. Mol. Biol. 15, 452-61, 2008).
(d) Model of SOSCat membrane recruitment via its allosteric pocket.
(e) HDPC recruitment, as measured in the stopped-flow assay, with and without Ras at the membrane. Relative recruitment was calculated by normalizing the SOS counts at the membrane to the concentration of SOS injected during the pulse. Data, as well as averages (red lines) for two SLB samples for each condition are shown. Note that other constructs in this study are recruited at least six times (in the case of HDPC(R552G)) more avidly compared to HDPC (Fig. 3b) and in this regard the control establish a baseline for the reported observations. Source data available online.
Supplementary Figure 3 Additional SOS recruitment data.
(a) Recruitment probability for DPC, HDPC and HDPC(R552G) on bilayers containing RasGDP or RasGTP. RasGDP data are replotted from Figure 3b for comparison. Note the generally increased recruitment probabilities across the panel of SOS constructs for binding to RasGTP. Each bar represents the average of data collected at N SLB samples, for RasGDP;RasGTP: DPC N=3;N=3, HDPC N=2;N=2, HDPC(R552G) N=3;N=3. Each sample was imaged at least at 15 different positions. Error bars represent SEM. For HDPC the individual data points from the two samples are shown instead of an average.
(b) Desorption traces of SOS constructs on RasGDP displaying bilayers. The shown traces represent averages over N SLB samples: SOSCat N=5, DPC N=4, HDPC N=2, HDPC(R552G) N=3. The data were fitted (solid lines) in accordance with a monoexponential decay process to extract the mean residency period for each construct (data presented in main Fig. 3c). In the case of SOSCat we observed a fraction of rapidly desorbing species during the first few seconds of the chase (Fig. 2c). This fast desorbing fraction was not contributing substantially to processive Ras turnover (Supplementary Fig. 1d) and for the comparison with other constructs shown here we cropped the first 10 s of the trace.
(c) Desorption traces of SOS constructs on RasGTP displaying bilayers. The shown traces represent averages over N SLB samples: SOSCat N=3, DPC N=2, HDPC N=2, HDPC(R552G) N=3. Same data format as in b. Note the generally slower desorption kinetics when SOS is bound via RasGTP (compare with RasGDP data presented in b).
(d) The single molecule SOS activity assay reported in Iversen et al. (Iversen, L. et al., Science. 345, 50-4, 2014) was adapted with the modification that here SOS was continuously incubated over the bilayers during imaging (as opposed to populating bilayers with SOS by providing a transient pulse through the flow cell). SOSHDPC (unlabeled) was incubated at a Ras-functionalized patterned supported lipid bilayer (3x3 µm corrals) at 12.5 pM. Ras was initially loaded with fluorescent nucleotide analog. The nucleotide label was imaged before and after incubation with SOS. After incubation a few corrals exhibited a depleted nucleotide signal relative to the neighboring corrals, indicating that a SOS molecule got activated at these particular sites of the membrane.
(e) The same experiment as in d, but with fluorescently labeled SOSDPC at 100 pM and dual imaging of the SOS and nucleotide labels. The 3 panels display time-traces of SOS and nucleotide on Ras for selected corrals. Step increases in the SOS channel signifies stable association of a SOS molecule with the membrane in the particular corral. As expected, arrival of SOS is followed by depletion of the initially labeled nucleotides on Ras as these are exchanged for unlabeled counterparts. In trace 1 and 3 the single dye label on SOS undergoes step photobleaching. The data illustrates that robust Ras activation requires stable association of SOS with the membrane and that catalysis takes place in a highly processive manner.
Supplementary Figure 4 Multiparameter p-FLOW assay of the Ras-ERK pathway.
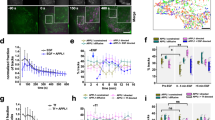
(a) Western blot for GFP confirming correct expression size of EGFP fusion SOS1 variants.
(b) Transfected cells expressing intermediary level of SOS1-EGFP exhibit signaling kinetics comparable to wild-type DT40 cells. BCR-induced fold increase in mean fluorescence intensity (MFI) of p-ERK relative to resting level (t= 0 min) is plotted for wild-type DT40 and full-length hSOS1-transfected SOS1-2- DT40 cells. Alternating subsets out of a total of 9 bins are compared to whole population of WT DT40 cells. Each data point represents four independent experiments (mean +/- SEM).
(c) SOS-EGFP can rescue ERK activation defect in SOS1-2- DT40. Representative scattered (left most column) and gated FACS data (right columns) at different time points after M4 stimulation are shown. Note that gate 9 has higher noise due to fewer acquired events (< 100 events).
(d) Two dimensional representation of p-ERK versus hSOS1-EGFP intensity across various time points after BCR stimulation comparing SOSFL (top row) and SOSCat (bottom row).
Supplementary Figure 5 SOSCat flanking domains block spontaneous activation in the basal state but promote Ras-ERK signal transduction after receptor stimulation.
(a,b) Incremental suppression by additional N-terminal domains (see also Fig. 5a-c).
(c,d) Incremental suppression by additional N-terminal domains is consistent to similar variants lacking PR domain. The ratio of pERK MFI for longer to shorter SOS variants is plotted against increasing SOS concentration for unstimulated cells (basal state). Addition of SOSCat- flanking domains inhibits spontaneous activation of ERK: DH-PH domain (c) and HF domain (d) (see also Fig. 5a-c)
(e-g) Time dependence of pERK MFI ratio after BCR stimulation is plotted for indicated constructs: Full length with a truncated HF domain (f) and full length with a truncated C-terminal PR domain (g). In each structural diagram, truncated domains are greyed out. p-FLOW assay data in this figure were obtained from BCR stimulated cells with intermediate SOS expression level as indicated in e.
All shown data are based on three independent cell cultures and pFLOW experiments. Error bars represent SEM. Yellow plane on the cubes in a and e indicates the corresponding subspace of the 3D parameter space in the pFLOW assay (c.f., Fig. 4c,d). Δ highlights the difference between the compared constructs.
Supplementary Figure 6 Membrane-targeting lipid modification of full-length (FL) SOS1 allows spontaneous activation of ERK and prolonged response after BCR stimulation.
(a) Farnesylation tag engineered to FL WT hSOS1 (SOS1-F) and point-mutated inactive control (SOS1-F*).
(b) Western blotting shows mobility shift by lipid modification.
(c) Ligand-independent activation of pERK. To note, membrane-anchored full-length SOS1 signals to ERK independent of BCR stimulation as observed on SOSCat-expressing cells (see also, Fig 4e,h)
(d) Time trace of p-ERK response upon BCR stimulation at intermediary SOS1-expressing cells.
BCR-induced ERK activity is prolonged compared to unmodified SOS1 or inactive SOS1-F*.
Shown data are representative of two independent cell cultures and pFLOW experiments. Source data are available online.
Supplementary Figure 7 Additional hybrid live-cell SLB assay data and replotted overlays.
(a) The anti-BCR antibody was coupled to the bilayer via Cy5-labeled streptavidin thus providing a readout for the location of activated BCR microclusters whereas SOS was visualized via EGFP tagging.
(b) Representative microscopy images collected from live cell experiments for various SOS constructs. Bright field (top row), RICM (second row), EGFP (third row), and Cy5 (bottom row) channels of cells expressing different SOS-EGFP constructs (one cell for each column). The images were taken 20-40 minutes after cell landing and spread on the antibody-functionalized supported lipid bilayers. Information on number of samples for each construct is given in Figure 6b. Bar: 3 µm.
(c) Overlay micrographs from Figure 6c replotted as separate image channels. Bar: 3 µm.
(d) Overlay from the inset in Figure 7d replotted as separate image channels.
Supplementary Figure 8 Monomeric SH2 fusion of HDPC restores ERK activation at later time points, and interaction between SOS1 and Grb2 is prolonged during antigen-receptor-induced Ras-ERK activation and human cancer– and Noonan syndrome–associated alterations of SOS1.
(a,b) Time dependence of pERK MFI ratio after BCR stimulation is plotted for indicated constructs: Single SH2 domain from human GrRB2 was fused to C-terminus of HDPC (HDPC-SH2). p-FLOW assay data in this figure were obtained from BCR stimulated cells with intermediate SOS expression level. Shown data are based on four independent cell cultures and pFLOW experiments. Error bars represent SEM. Source data are available online.
(c,d) Mouse CD4+ T cell blasts and hSOS1 stably transfected DT40 B cells were stimulated with antigen receptor crosslinking antibodies for the indicated time. SOS1 was immunoprecipitated from resulting lysates. Immunoprecipitated SOS1 and associated Grb2 were visualized by Western blotting. Phosphorylated ERK was used to indicate the status of Ras-ERK pathway. For panel (d), the percentage to max pERK2 at the indicated time is calculated after normalization to total ERK2 amount. Shown results are representative of two independent replicate experiments of each cell type.
(e) Human cancer- and Noonan-syndrome associated alterations of SOS1. Each bar represents a SOS alteration detected in patients. Green bars indicate missense point mutations. Alterations disrupting the C-terminus are indicated with red bars (fs: frame shift, spl: aberrant splicing, *: STOP codon,). Yellow boxes indicate PxxP motifs in C-terminal interacting with Grb2. Modified from cBioPortal for Cancer Genomics Database (Gao, J. et al, Sci. Signal., 6, p. l1, 2013).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Notes 1–7 (PDF 2541 kb)
SOS-EGFP expressing B cell interacting with SLB presenting cross-linking antibody for the BCR
Time-lapse movie showing trajectories of BCR (red) and SOSFL (green) movement at the cell-bilayer interface. (AVI 284 kb)
Source data
Rights and permissions
About this article
Cite this article
Christensen, S., Tu, HL., Jun, J. et al. One-way membrane trafficking of SOS in receptor-triggered Ras activation. Nat Struct Mol Biol 23, 838–846 (2016). https://doi.org/10.1038/nsmb.3275
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3275
This article is cited by
-
Membrane anchoring facilitates colocalization of enzymes in plant cytochrome P450 redox systems
Communications Biology (2021)
-
Stability Analysis of a Bulk–Surface Reaction Model for Membrane Protein Clustering
Bulletin of Mathematical Biology (2020)
-
Novel regulation of Ras proteins by direct tyrosine phosphorylation and dephosphorylation
Cancer and Metastasis Reviews (2020)
-
The CSN3 subunit of the COP9 signalosome interacts with the HD region of Sos1 regulating stability of this GEF protein
Oncogenesis (2019)
-
Mechanism of SOS PR-domain autoinhibition revealed by single-molecule assays on native protein from lysate
Nature Communications (2017)