Abstract
Liver cancer stem cells (CSCs) may contribute to the high rate of recurrence and heterogeneity of hepatocellular carcinoma (HCC); however, the molecular mechanisms underlying their self-renewal and differentiation remain largely unknown. Through analysis of transcriptome microarray data, we identified a long noncoding RNA (lncRNA) called lnc-β-Catm, which is highly expressed in human HCC tumors and liver CSCs. We found that lnc-β-Catm is required for self-renewal of liver CSCs and tumor propagation in mice. lnc-β-Catm associates with β-catenin and the methyltransferase EZH2, thereby promoting β-catenin methylation. Methylation suppresses the ubiquitination of β-catenin and promotes its stability, thus leading to activation of Wnt–β-catenin signaling. Accordingly, the expression of lnc-β-Catm, EZH2 and Wnt–β-catenin targets is positively correlated with cancer severity and prognosis of people with HCC.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Bruix, J., Gores, G.J. & Mazzaferro, V. Hepatocellular carcinoma: clinical frontiers and perspectives. Gut 63, 844–855 (2014).
Prieto, J., Melero, I. & Sangro, B. Immunological landscape and immunotherapy of hepatocellular carcinoma. Nat. Rev. Gastroenterol. Hepatol. 12, 681–700 (2015).
Kreso, A. & Dick, J.E. Evolution of the cancer stem cell model. Cell Stem Cell 14, 275–291 (2014).
Yamashita, T. et al. EpCAM-positive hepatocellular carcinoma cells are tumor-initiating cells with stem/progenitor cell features. Gastroenterology 136, 1012–1024 (2009).
Kurtova, A.V. et al. Blocking PGE2-induced tumour repopulation abrogates bladder cancer chemoresistance. Nature 517, 209–213 (2015).
Salmon, S.E. et al. Quantitation of differential sensitivity of human-tumor stem cells to anticancer drugs. N. Engl. J. Med. 298, 1321–1327 (1978).
Yang, Z.F. et al. Significance of CD90+ cancer stem cells in human liver cancer. Cancer Cell 13, 153–166 (2008).
Haraguchi, N. et al. CD13 is a therapeutic target in human liver cancer stem cells. J. Clin. Invest. 120, 3326–3339 (2010).
Visvader, J.E. & Lindeman, G.J. Cancer stem cells: current status and evolving complexities. Cell Stem Cell 10, 717–728 (2012).
Li, L. & Neaves, W.B. Normal stem cells and cancer stem cells: the niche matters. Cancer Res. 66, 4553–4557 (2006).
Cairo, S. et al. Hepatic stem-like phenotype and interplay of Wnt/β-catenin and Myc signaling in aggressive childhood liver cancer. Cancer Cell 14, 471–484 (2008).
Takebe, N. et al. Targeting Notch, Hedgehog, and Wnt pathways in cancer stem cells: clinical update. Nat. Rev. Clin. Oncol. 12, 445–464 (2015).
Zhu, P. et al. C8orf4 negatively regulates self-renewal of liver cancer stem cells via suppression of NOTCH2 signalling. Nat. Commun. 6, 7122 (2015).
Thompson, M.D. & Monga, S.P. WNT/β-catenin signaling in liver health and disease. Hepatology 45, 1298–1305 (2007).
Henderson, B.R. Nuclear-cytoplasmic shuttling of APC regulates β-catenin subcellular localization and turnover. Nat. Cell Biol. 2, 653–660 (2000).
Wu, G. et al. Structure of a β-TrCP1-Skp1-β-catenin complex: destruction motif binding and lysine specificity of the SCFβ-TrCP1 ubiquitin ligase. Mol. Cell 11, 1445–1456 (2003).
Korinek, V. et al. Constitutive transcriptional activation by a β-catenin-Tcf complex in APC-/- colon carcinoma. Science 275, 1784–1787 (1997).
Wang, K.C. et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature 472, 120–124 (2011).
Petruk, S. et al. Transcription of bxd noncoding RNAs promoted by trithorax represses Ubx in cis by transcriptional interference. Cell 127, 1209–1221 (2006).
Rapicavoli, N.A., Poth, E.M., Zhu, H. & Blackshaw, S. The long noncoding RNA Six3OS acts in trans to regulate retinal development by modulating Six3 activity. Neural Dev. 6, 32 (2011).
Liu, B. et al. A cytoplasmic NF-κB interacting long noncoding RNA blocks IκB phosphorylation and suppresses breast cancer metastasis. Cancer Cell 27, 370–381 (2015).
Yuan, J.H. et al. A long noncoding RNA activated by TGF-β promotes the invasion-metastasis cascade in hepatocellular carcinoma. Cancer Cell 25, 666–681 (2014).
Gupta, R.A. et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 464, 1071–1076 (2010).
Wang, Y. et al. The long noncoding RNA lncTCF7 promotes self-renewal of human liver cancer stem cells through activation of Wnt signaling. Cell Stem Cell 16, 413–425 (2015).
Zhu, P. et al. ZIC2-dependent OCT4 activation drives self-renewal of human liver cancer stem cells. J. Clin. Invest. 125, 3795–3808 (2015).
Ezhkova, E. et al. Ezh2 orchestrates gene expression for the stepwise differentiation of tissue-specific stem cells. Cell 136, 1122–1135 (2009).
Viré, E. et al. The Polycomb group protein EZH2 directly controls DNA methylation. Nature 439, 871–874 (2006).
Xu, K. et al. EZH2 oncogenic activity in castration-resistant prostate cancer cells is Polycomb-independent. Science 338, 1465–1469 (2012).
Kim, E. et al. Phosphorylation of EZH2 activates STAT3 signaling via STAT3 methylation and promotes tumorigenicity of glioblastoma stem-like cells. Cancer Cell 23, 839–852 (2013).
James, R.G. et al. WIKI4, a novel inhibitor of tankyrase and Wnt/ß-catenin signaling. PLoS One 7, e50457 (2012).
Roessler, S. et al. Integrative genomic identification of genes on 8p associated with hepatocellular carcinoma progression and patient survival. Gastroenterology 142, 957–966. e12 (2012).
Gupta, P.B., Chaffer, C.L. & Weinberg, R.A. Cancer stem cells: mirage or reality? Nat. Med. 15, 1010–1012 (2009).
Jia, Q., Zhang, X., Deng, T. & Gao, J. Positive correlation of Oct4 and ABCG2 to chemotherapeutic resistance in CD90+CD133+ liver cancer stem cells. Cell. Reprogram. 15, 143–150 (2013).
Zhao, W. et al. 1B50-1, a mAb raised against recurrent tumor cells, targets liver tumor-initiating cells by binding to the calcium channel α2δ1 subunit. Cancer Cell 23, 541–556 (2013).
Rinn, J.L. & Chang, H.Y. Genome regulation by long noncoding RNAs. Annu. Rev. Biochem. 81, 145–166 (2012).
Ulitsky, I. & Bartel, D.P. lincRNAs: genomics, evolution, and mechanisms. Cell 154, 26–46 (2013).
Yamashita, T., Budhu, A., Forgues, M. & Wang, X.W. Activation of hepatic stem cell marker EpCAM by Wnt-β-catenin signaling in hepatocellular carcinoma. Cancer Res. 67, 10831–10839 (2007).
Myant, K.B. et al. ROS production and NF-κB activation triggered by RAC1 facilitate WNT-driven intestinal stem cell proliferation and colorectal cancer initiation. Cell Stem Cell 12, 761–773 (2013).
Li, V.S. et al. Wnt signaling through inhibition of β-catenin degradation in an intact Axin1 complex. Cell 149, 1245–1256 (2012).
Liu, C. et al. Control of β-catenin phosphorylation/degradation by a dual-kinase mechanism. Cell 108, 837–847 (2002).
Gao, C., Xiao, G. & Hu, J. Regulation of Wnt/β-catenin signaling by posttranslational modifications. Cell Biosci. 4, 13 (2014).
Lu, L. et al. Kdm2a/b lysine demethylases regulate canonical Wnt signaling by modulating the stability of nuclear β-catenin. Dev. Cell 33, 660–674 (2015).
Asangani, I.A. et al. Characterization of the EZH2-MMSET histone methyltransferase regulatory axis in cancer. Mol. Cell 49, 80–93 (2013).
Zhang, X. et al. Coordinated silencing of MYC-mediated miR-29 by HDAC3 and EZH2 as a therapeutic target of histone modification in aggressive B-cell lymphomas. Cancer Cell 22, 506–523 (2012).
Lee, S.T. et al. Context-specific regulation of NF-κB target gene expression by EZH2 in breast cancers. Mol. Cell 43, 798–810 (2011).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Guichard, C. et al. Integrated analysis of somatic mutations and focal copy-number changes identifies key genes and pathways in hepatocellular carcinoma. Nat. Genet. 44, 694–698 (2012).
Davis, C.F. et al. The somatic genomic landscape of chromophobe renal cell carcinoma. Cancer Cell 26, 319–330 (2014).
Tseng, Y.Y. et al. PVT1 dependence in cancer with MYC copy-number increase. Nature 512, 82–86 (2014).
Justilien, V. et al. The PRKCI and SOX2 oncogenes are coamplified and cooperate to activate Hedgehog signaling in lung squamous cell carcinoma. Cancer Cell 25, 139–151 (2014).
Chen, J. et al. The endoplasmic reticulum adaptor protein ERAdP initiates NK cell activation via the Ubc13-mediated NF-κB pathway. J. Immunol. 194, 1292–1303 (2015).
Xia, P. et al. RNF2 is recruited by WASH to ubiquitinate AMBRA1 leading to downregulation of autophagy. Cell Res. 24, 943–958 (2014).
Xia, P. et al. WASH is required for the differentiation commitment of hematopoietic stem cells in a c-Myc-dependent manner. J. Exp. Med. 211, 2119–2134 (2014).
Xia, P. et al. Sox2 functions as a sequence-specific DNA sensor in neutrophils to initiate innate immunity against microbial infection. Nat. Immunol. 16, 366–375 (2015).
Hu, Y. & Smyth, G.K. ELDA: extreme limiting dilution analysis for comparing depleted and enriched populations in stem cell and other assays. J. Immunol. Methods 347, 70–78 (2009).
Acknowledgements
We thank J. Jia for technical support. This work was supported by the National Natural Science Foundation of China (91419308 to Z.F., 31530093 to Z.F., 81330047 to Z.F. and 81402459 to Y.W.); the State Projects of Essential Drug Research and Development (2012ZX09103301-041 to Z.F.); the 973 Program of the MOST of China (2015CB553705 to Z.F.); the Strategic Priority Research Programs of the Chinese Academy of Sciences (XDA01010407 to Z.F.); and the Beijing Natural Science Foundation (7162125 to Y.W.).
Author information
Authors and Affiliations
Contributions
P.Z. designed and performed experiments, analyzed data and wrote the paper; Y.W. performed experiments and analyzed data; G.H. and J.W. performed some experiments; L.H. provided HCC samples and analyzed data; B.Y., B.L. and Y.D. analyzed data. Z.F. initiated the study and organized, designed and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Characterization and high expression of Lnc-β-Catm in liver CSCs.
(a) Heatmap of differently expressed lncRNAs in Liver CSCs (CD13+CD133+) and non-CSCs (CD13-CD133-) according to transcriptome analyses. (b) 3’ and 5’ RACE for full length of lnc-β-Catm. The length of lnc-β-Catm was 2281 nucleotides verified by sequencing. Black arrowhead denotes the full length of lnc-β-Catm. (c) Histogram of lnc-β-Catm coding potential analyzed by CPC (left panel), CPAT (middle panel) and PhyloCSF (right panel). HOX transcript antisense RNA (Hotair), X inactivation-specific transcript (XIST) and lncTCF7 serve as control non-coding RNAs. β-actin (ACTB) and glyceraldehyde-3-phosphate dehydrogenase (GAPDH) serve as control coding genes. For CPC and phyloCSF, scores above 0 indicates coding potential, whereas scores below 0 represent no coding potential. CPAT scores indicate the possibility of coding. (d) Anti-Myc and anti-β-actin Western blots. Samples were shown in left panels. Full length of lnc-β-Catm was cloned into an eukaryotic expression vector pcDNA4-His-Myc B with transcription initiating codon ATG in three expression patterns. T, thymine. (e) Scatter plots (means±s.d.) of nomorlized lnc-β-Catm expression levels. eHCC, early HCC; aHCC, advanced HCC. (f, g) Histogram of lnc-β-Catm expression levels in liver CSCs (CD13+CD133+) and non-CSCs (CD13-CD133-) (f), or in oncospheres and non-spheres (g). Error bars, s.d. (n = 4 cell cultures). Two tailed Student’s t-test was used for statistical analysis, **, P < 0.01; ***, P<0.001. (h) Western blots (upper panel) and realtime PCR (lower panel) of Nucleocytoplasmic separation fractions. Error bars, s.d. (n = 4 cell cultures). U1 RNA serves as a positive control for nuclear location. EEA1, endosome antigen 1; H3, histone 3. For b, d, h, uncropped blots and gels can be found in Supplementary Data Set 1.
Supplementary Figure 2 Lnc-β-Catm enhances self-renewal of liver CSCs.
(a) Relative lnc-β-Catm expression levels of lnc-β-Catm and control cells. Error bars, s.d. (n = 3 cell cultures). Two tailed Student’s t-test was used for statistical analysis, **, P < 0.01; ***, P<0.001. (b, c) Serial sphere formation (b) and tumor propagation (c) with lnc-β-Catm overexpressing and control cells. Error bars, s.d. (n = 3 cell cultures for b, n = 5 mice for c). Two tailed Student’s t-test was used for statistical analysis, *, P < 0.05; **, P<0.01; ***, P<0.001. (d) Tumor-free mice ratios after 3 months’ tumor formation with lnc-β-Catm overexpressing (oeLnc) and control (oeVec) cells. n = 6 mice for each group.
Supplementary Figure 3 Lnc-β-Catm associates with β-catenin and EZH2.
(a) Relative mRNA expression levels in lnc-β-Catm silenced and control cells (lower panels), 16 nearby genes of lnc-β-Catm locus (less than 2 Mb) (upper panels) were analyzed. Error bars, s.d. (n = 4 cell cultures). Two tailed Student’s t-test was used for statistical analysis, **, P < 0.01. (b, c) MS-MS profiles of β-catenin (b) and EZH2 (c), corresponding peptide sequences are listed on the top of the corresponding graphs. (d) Different regions of lnc-β-Catm (left panel) were labeled with biotin and incubated with PLC sphere lysates, followed by RNA pulldown assays (right panel). (e) Stem-loop structures of full length (1-2281 nt), segment #6 (1181-1437 nt) and segment #9 (1938-2281 nt) of lnc-β-Catm. Predictions were based on minimum free energy (MFE) and partition function. Color scales denote confidence of predictions for each base with shades of red indicating strong confidence (http://rna.tbi.univie.ac.at/). (f) Anti-β-catenin and anti-EZH2 Western blots. Samples were derived from co-immunoprecipitation (co-IP) with PLC spheres. (g, h) Anti-β-catenin, anti-EZH2, anti-β-actin (loading control) and anti-Oct4 (serum treated control) Western blots. Samples were immunoprecipitates from spheres (S) and non-spheres (N) (g), or serum treated spheres (h). (i) Intensity profiles along the diagonal from upper left to lower right. Green profiles indicate lnc-β-Catm gray value (intensity), red profiles indicate β-catenin intensity, and blue EZH2. (j) Flag-β-catenin truncates (upper panels) overexpressing spheres were established, followed by co-IP assays and Western blots (lower panels). (k) Anti-Myc and anti-His Western blots (right panels) with EZH2 truncates overepressing spheres, as in j. (l, m) Histogram of lnc-β-Catm enrichment after RNA immunoprecipitation assays. β-catenin truncates (l) and EZH2 truncates (m) overexpressing PLC spheres were used. Error bars, s.d. (n = 4 independent experiments). Throughout figure, uncropped blots and gels can be found in Supplementary Data Set 1.
Supplementary Figure 4 Characterization of β-catenin methylation.
(a) Anti-Methylated lysine, anti-β-catenin, anti-H3 and anti-EEA1 Western blots. Samples were nuclear (N) and cytoplasmic (C) fractions of the indicated spheres. EEA1, endosome antigen 1; H3, histone 3. (b) Western blots for β-catenin methylation signals in HCC tumor tissues (T) and peri-tumor tissues (P). (c) Methylation observation of β-catenin in peri-tumor and tumor tissues. β-catenin (green), methylated lysine (red), and EZH2 (blue). Scale bars, 20 μm. Uncropped blots in a and b can be found in Supplementary Data Set 1.
Supplementary Figure 5 β-catenin methylation promotes its stability.
(a) Anti-phosphorylated β-catenin (p-β-catenin), anti-β-catenin, anti-EZH2 and anti-β-actin (control) Western blots using EZH2 overexpressing (oeEZH2) and control (oeVec) spheres. (b) Anti-K48 linkage ubiquitylation (K48-Ub), anti-β-catenin and anti-β-actin (control) Western blots. EZH2 inhibitors (GSK126 and GSK343) treated and control (DMSO) spheres were used for β-catenin immunoprecipitation. (c) Anti-β-catenin and anti-β-actin (control) Western blots of methylated and non-methylated β-catenin supplemented with sphere lysates (left panels). Relative β-catenin levels in the right panel. Error bars, s.d. (n = 3 cell cultures). Throughout figure, uncropped Western blot results can be found in Supplementary Data Set 1.
Supplementary Figure 6 Lnc-β-Catm promotes Wnt signaling by increasing β-catenin stability.
(a) Relative expression levels of Wnt-β-catenin target genes in lnc-β-Catm silenced and control spheres. (b) Relative β-catenin protein levels in lnc-β-Catm depleted spheres and control spheres. Error bars, s.d. (n=3 cell cultures). Two tailed Student’s t-test was used for statistical analysis, **, P < 0.01; ***, P < 0.001. (c) Anti-methylated lysine and anti-β-catenin (immunoprecipitation control) Western blots. Lnc-β-Catm overexpressing (oeLnc) and control (oeVec) HCC primary spheres were used. (d, e) Realtime PCR (d) and Western blots (e) of Wnt-β-catenin target genes in lnc-β-Catm KO cells (lnc-β-Catm KO) and rescued cells (lnc-β-Catm KO+rescueLnc). Error bars, s.d. (n=3 cell cultures). (f-i) Anti-methylated lysine, anti-β-catenin, anti-ubiquitination and anti-actin (control) Western blots for β-catenin methylation (f), phosphorylation (g), ubiquitination (h) and stability (i). Samples were lnc-β-Catm KO cells, rescued cells and control cells. For i, relative β-catenin protein levels were calculated and shown in the right panel. Error bars, s.d. (n = 3 cell cultures). Two tailed Student’s t-test was used for statistical analysis, *, P < 0.05; **, P < 0.01; ***, P < 0.001. (j, k) Sphere formation (j) and xenograft tumor growth (k). Samples were lnc-β-Catm silenced cells rescued with Wnt-β-catenin target genes (c-Myc, Ccnd1, and Pttg1). Scale bar, 500 μm. Error bars, s.d. (n = 3 independent experiments). Two tailed Student’s t-test was used for statistical analysis, *, P < 0.05. (i) Sphere formation of lnc-β-Catm overexpressed spheres supplemented with Wnt-β-catenin inhibitor WIKI4. Typical images were shown in left panels and sphere formation ratios were calculated (right panels). Error bars, s.d. (n = 3 independent experiments). Two tailed Student’s t-test was used for statistical analysis, *, P < 0.05; **, P < 0.01. ns, not significant. Throughout figure, uncropped blots and gels can be found in Supplementary Data Set 1.
Supplementary Figure 7 Lnc-β-Catm plays a predominant role in HCC and liver CSCs.
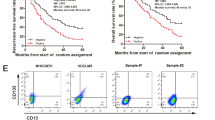
(a, b) Expression levels of EZH2 and Wnt-β-catenin target genes in HCC tumors (a) and metastasis patients (b) derived from Wang’s cohort. Data are shown as box-and-whisker plots. Whiskers: 5th and 95th percentiles; Horizontal lines: median levels; Boxes: interquartile range (IQR); upper and lower edges: 75th and 25th percentiles. (c) Kaplan-Meier survival analysis of Wnt-β-catenin target genes. HCC samples were divided into 2 groups according to the indicated gene expression levels. (d) Sphere formation of lnc-β-Catm and lncTCF7 silenced and control HCC primary cells. 31 HCC primary cells were used. *, **, ***, lncRNA shRNA versus control shRNA. #, ##, lnc-β-Catm shRNA versus lncTCF7 shRNA. Error bars, s.d. (n = 3 cell cultures). Two tailed Student’s t-test was used for statistical analysis, *, P < 0.05; **, P < 0.01; ***, P < 0.001; #, P < 0.05; ##, P < 0.01. (e, f) Confocal observation with CD133 antibody (e), Oct4 antibody and c-Myc antibody (f). Control, lnc-β-Catm silenced and lncTCF7 silenced spheres were used. Scale bar, 20 μm.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–3 (PDF 2331 kb)
Supplementary Data Set 1
Uncropped blots and gels (PDF 2659 kb)
Rights and permissions
About this article
Cite this article
Zhu, P., Wang, Y., Huang, G. et al. lnc-β-Catm elicits EZH2-dependent β-catenin stabilization and sustains liver CSC self-renewal. Nat Struct Mol Biol 23, 631–639 (2016). https://doi.org/10.1038/nsmb.3235
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3235
This article is cited by
-
Crosstalk between colorectal CSCs and immune cells in tumorigenesis, and strategies for targeting colorectal CSCs
Experimental Hematology & Oncology (2024)
-
EZH2 in hepatocellular carcinoma: progression, immunity, and potential targeting therapies
Experimental Hematology & Oncology (2023)
-
The epigenetic factor CHD4 contributes to metastasis by regulating the EZH2/β-catenin axis and acts as a therapeutic target in ovarian cancer
Journal of Translational Medicine (2023)
-
LncRNA H19-EZH2 interaction promotes liver fibrosis via reprogramming H3K27me3 profiles
Acta Pharmacologica Sinica (2023)
-
ENO2-derived phosphoenolpyruvate functions as an endogenous inhibitor of HDAC1 and confers resistance to antiangiogenic therapy
Nature Metabolism (2023)