Abstract
Adenosine deaminases acting on RNA (ADARs) are editing enzymes that convert adenosine to inosine in duplex RNA, a modification reaction with wide-ranging consequences in RNA function. Understanding of the ADAR reaction mechanism, the origin of editing-site selectivity, and the effect of mutations is limited by the lack of high-resolution structural data for complexes of ADARs bound to substrate RNAs. Here we describe four crystal structures of the human ADAR2 deaminase domain bound to RNA duplexes bearing a mimic of the deamination reaction intermediate. These structures, together with structure-guided mutagenesis and RNA-modification experiments, explain the basis of the ADAR deaminase domain's dsRNA specificity, its base-flipping mechanism, and its nearest-neighbor preferences. In addition, we identified an ADAR2-specific RNA-binding loop near the enzyme active site, thus rationalizing differences in selectivity observed between different ADARs. Finally, our results provide a structural framework for understanding the effects of ADAR mutations associated with human disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Grosjean, H. Fine-Tuning of RNA Functions by Modification and Editing (Springer, 2005).
Bass, B.L. RNA editing by adenosine deaminases that act on RNA. Annu. Rev. Biochem. 71, 817–846 (2002).
Nishikura, K. Functions and regulation of RNA editing by ADAR deaminases. Annu. Rev. Biochem. 79, 321–349 (2010).
Wang, Q. et al. ADAR1 regulates ARHGAP26 gene expression through RNA editing by disrupting miR-30b-3p and miR-573 binding. RNA 19, 1525–1536 (2013).
Rueter, S.M., Dawson, T.R. & Emeson, R.B. Regulation of alternative splicing by RNA editing. Nature 399, 75–80 (1999).
Yeo, J., Goodman, R.A., Schirle, N.T., David, S.S. & Beal, P.A. RNA editing changes the lesion specificity for the DNA repair enzyme NEIL1. Proc. Natl. Acad. Sci. USA 107, 20715–20719 (2010).
Bass, B.L. et al. A standardized nomenclature for adenosine deaminases that act on RNA. RNA 3, 947–949 (1997).
Maas, S., Kawahara, Y., Tamburro, K.M. & Nishikura, K. A-to-I RNA editing and human disease. RNA Biol. 3, 1–9 (2006).
Slotkin, W. & Nishikura, K. Adenosine-to-inosine RNA editing and human disease. Genome Med. 5, 105 (2013).
Morabito, M.V. et al. Mice with altered serotonin 2C receptor RNA editing display characteristics of Prader-Willi syndrome. Neurobiol. Dis. 39, 169–180 (2010).
Rice, G.I. et al. Mutations in ADAR1 cause Aicardi-Goutières syndrome associated with a type I interferon signature. Nat. Genet. 44, 1243–1248 (2012).
Miyamura, Y. et al. Mutations of the RNA-specific adenosine deaminase gene (DSRAD) are involved in dyschromatosis symmetrica hereditaria. Am. J. Hum. Genet. 73, 693–699 (2003).
Zhang, X.J. et al. Seven novel mutations of the ADAR gene in Chinese families and sporadic patients with dyschromatosis symmetrica hereditaria (DSH). Hum. Mutat. 23, 629–630 (2004).
Chen, L. et al. Recoding RNA editing of AZIN1 predisposes to hepatocellular carcinoma. Nat. Med. 19, 209–216 (2013).
Gallo, A. RNA editing enters the limelight in cancer. Nat. Med. 19, 130–131 (2013).
Shimokawa, T. et al. RNA editing of the GLI1 transcription factor modulates the output of Hedgehog signaling. RNA Biol. 10, 321–333 (2013).
Goodman, R.A., Macbeth, M.R. & Beal, P.A. ADAR proteins: structure and catalytic mechanism. Curr. Top. Microbiol. Immunol. 353, 1–33 (2012).
Li, J.B. et al. Genome-wide identification of human RNA editing sites by parallel DNA capturing and sequencing. Science 324, 1210–1213 (2009).
Haudenschild, B.L. et al. A transition state analogue for an RNA-editing reaction. J. Am. Chem. Soc. 126, 11213–11219 (2004).
Phelps, K.J. et al. Recognition of duplex RNA by the deaminase domain of the RNA editing enzyme ADAR2. Nucleic Acids Res. 43, 1123–1132 (2015).
Macbeth, M.R. et al. Inositol hexakisphosphate is bound in the ADAR2 core and required for RNA editing. Science 309, 1534–1539 (2005).
Kuttan, A. & Bass, B.L. Mechanistic insights into editing-site specificity of ADARs. Proc. Natl. Acad. Sci. USA 109, E3295–E3304 (2012).
Wong, S.K., Sato, S. & Lazinski, D.W. Substrate recognition by ADAR1 and ADAR2. RNA 7, 846–858 (2001).
Klimasauskas, S., Kumar, S., Roberts, R.J. & Cheng, X. HhaI methyltransferase flips its target base out of the DNA helix. Cell 76, 357–369 (1994).
Daujotyte, D. et al. HhaI DNA methyltransferase uses the protruding Gln237 for active flipping of its target cytosine. Structure 12, 1047–1055 (2004).
Thiyagarajan, S., Rajan, S.S. & Gautham, N. Cobalt hexammine induced tautomeric shift in Z-DNA: the structure of d(CGCGCA)*d(TGCGCG) in two crystal forms. Nucleic Acids Res. 32, 5945–5953 (2004).
Slupphaug, G. et al. A nucleotide-flipping mechanism from the structure of human uracil-DNA glycosylase bound to DNA. Nature 384, 87–92 (1996).
Bruner, S.D., Norman, D.P. & Verdine, G.L. Structural basis for recognition and repair of the endogenous mutagen 8-oxoguanine in DNA. Nature 403, 859–866 (2000).
Lau, A.Y., Schärer, O.D., Samson, L., Verdine, G.L. & Ellenberger, T. Crystal structure of a human alkylbase-DNA repair enzyme complexed to DNA: mechanisms for nucleotide flipping and base excision. Cell 95, 249–258 (1998).
Brooks, S.C., Adhikary, S., Rubinson, E.H. & Eichman, B.F. Recent advances in the structural mechanisms of DNA glycosylases. Biochim. Biophys. Acta 1834, 247–271 (2013).
Eifler, T., Pokharel, S. & Beal, P.A. RNA-Seq analysis identifies a novel set of editing substrates for human ADAR2 present in Saccharomyces cerevisiae. Biochemistry 52, 7857–7869 (2013).
Eggington, J.M., Greene, T. & Bass, B.L. Predicting sites of ADAR editing in double-stranded RNA. Nat. Commun. 2, 319 (2011).
Peacock, H., Maydanovych, O. & Beal, P.A. N2-Modified 2-aminopurine ribonucleosides as minor-groove-modulating adenosine replacements in duplex RNA. Org. Lett. 12, 1044–1047 (2010).
Roberts, R.J. & Cheng, X. Base flipping. Annu. Rev. Biochem. 67, 181–198 (1998).
Hoang, C. & Ferré-D'Amaré, A.R. Cocrystal structure of a tRNA Psi55 pseudouridine synthase: nucleotide flipping by an RNA-modifying enzyme. Cell 107, 929–939 (2001).
Parikh, S.S. et al. Base excision repair initiation revealed by crystal structures and binding kinetics of human uracil-DNA glycosylase with DNA. EMBO J. 17, 5214–5226 (1998).
Yang, X., Gérczei, T., Glover, L.T. & Correll, C.C. Crystal structures of restrictocin-inhibitor complexes with implications for RNA recognition and base flipping. Nat. Struct. Biol. 8, 968–973 (2001).
Carter, A.P. et al. Crystal structure of an initiation factor bound to the 30S ribosomal subunit. Science 291, 498–501 (2001).
Réblová, K. et al. Structure, dynamics, and elasticity of free 16s rRNA helix 44 studied by molecular dynamics simulations. Biopolymers 82, 504–520 (2006).
Xie, W., Liu, X. & Huang, R.H. Chemical trapping and crystal structure of a catalytic tRNA guanine transglycosylase covalent intermediate. Nat. Struct. Biol. 10, 781–788 (2003).
Erion, M.D. & Reddy, M.R. Calculation of relative hydration free energy differences for heteroaromatic compounds: use in the design of adenosine deaminase and cytidine deaminase inhibitors. J. Am. Chem. Soc. 120, 3295–3304 (1998).
Stefl, R. et al. The solution structure of the ADAR2 dsRBM-RNA complex reveals a sequence-specific readout of the minor groove. Cell 143, 225–237 (2010).
Fierro-Monti, I. & Mathews, M.B. Proteins binding to duplexed RNA: one motif, multiple functions. Trends Biochem. Sci. 25, 241–246 (2000).
Mannion, N.M. et al. The RNA-editing enzyme ADAR1 controls innate immune responses to RNA. Cell Rep. 9, 1482–1494 (2014).
Macbeth, M.R. & Bass, B.L. Large-scale overexpression and purification of ADARs from Saccharomyces cerevisiae for biophysical and biochemical studies. Methods Enzymol. 424, 319–331 (2007).
Pokharel, S. et al. Matching active site and substrate structures for an RNA editing reaction. J. Am. Chem. Soc. 131, 11882–11891 (2009).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Afonine, P.V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D Biol. Crystallogr. 68, 352–367 (2012).
Phelps, K.J., Ibarra-Soza, J.M., Tran, K., Fisher, A.J. & Beal, P.A. Click modification of RNA at adenosine: structure and reactivity of 7-ethynyl- and 7-triazolyl-8-aza-7-deazaadenosine in RNA. ACS Chem. Biol. 9, 1780–1787 (2014).
Acknowledgements
The authors acknowledge funding from the US National Institutes of Health (NIH) grant R01GM061115 (P.A.B.). A.I.S. was supported by NIH training grant T32 GM007377. C. Palumbo is acknowledged for technical assistance. Use of the Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, is supported by the US Department of Energy (DOE), Office of Science, Office of Basic Energy Sciences under contract no. DE-AC02-76SF00515. The SSRL Structural Molecular Biology Program is supported by the US Department of Energy, Office of Biological and Environmental Research, and by the NIH, US National Institute of General Medical Sciences (including P41GM103393). Part of this work is also based on research conducted at the Northeastern Collaborative Access Team beamlines, which are funded by the National Institute of General Medical Sciences from the NIH (P41 GM103403). The Pilatus 6M detector on the 24-ID-C beamline is funded by an NIH-ORIP HEI grant (S10 RR029205). This research used resources of the Advanced Photon Source, a DOE Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under contract no. DE-AC02-06CH11357. The contents of this publication are solely the responsibility of the authors and do not necessarily represent the official views of NIGMS or NIH.
Author information
Authors and Affiliations
Contributions
J.M.T., M.M.M., A.I.S., and Y.Z. purified protein. K.J.P. and J.M.T. designed and purified RNA for crystallography and characterized protein-RNA binding. M.M.M. and A.I.S. conducted crystallization trials. M.M.M. and A.J.F. collected diffraction data and solved and refined the crystal structures. J.M.T., Y.Z., and J.H. measured enzyme reaction rates. K.T. synthesized 8-azanebularane phosphoramidite. J.M.T. and A.I.S. conducted mutagenesis. J.M.T., M.M.M., P.A.B. and A.J.F. analyzed the structures. P.A.B. wrote the initial manuscript draft. J.M.T., M.M.M., P.A.B., and A.J.F. edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
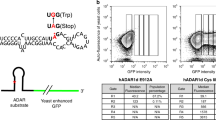
Supplementary Figure 1 8-Azanebularine-containing duplex RNAs form a tight and specific complex with hADAR2d.
(a) Representative electrophoretic mobility shift assay (EMSA) gel displaying tight and specific binding of hADARd E488Q mutant and N-containing hGLI1 24mer duplex. Lane 1: no protein added; Lanes 2-11: 0.25, 0.5, 1, 2, 4, 8, 16, 32, 64, and 128 nM hADAR2d E488Q. (b) Fitted plot of fraction RNA bound vs. hADAR2d E488Q concentration.
Supplementary Figure 2 Overall structure of the hADAR2d–RNA complex.
(a) (Left) A simulated-annealed composite omit electron density map calculated in PHENIX (Afonine, P.V. et al. Acta Crystallogr D Biol Crystallogr 71, 646-666, 2015) is shown in blue mesh. The map is contoured at 1σ at 2.75Å resolution. The inositol hexakisphosphate is shown in white-colored-carbon sticks, and the active site zinc atom as a gray sphere near the flipped out base. (Right) View of hADAR2d E488Q-Bdf2 complex showing protein surface electrostatic potential. The electrostatic potential was calculated use the Adaptive Poisson-Boltzmann Solver (APBS) plugin in PyMol (Baker, N.A et al. Proc Natl Acad Sci USA 98, 10037-10041, 2001). Electrostatic potential was calculated using the default values, and displayed on the protein surface with blue and red representing the positive and negative potentials respectively. (b) Summary of protein-RNA contacts observed in the Gli1-hADAR2d E488Q complex. (c) Summary of protein-RNA contacts observed in the hADAR2d WT-Bdf2-C complex. (d) Summary of protein-RNA contacts observed in the hADAR2d WT-Bdf2-U complex.
Supplementary Figure 3 Close-up views of the active site and 5′-nearest-neighbor nucleotides with electron density map.
(a) (Left) Close-up view of the flipped out base with electron density. The flipped out 8-azanebularine nucleotide (hADAR2d E488Q + Bdf2-C) is shown as sticks with white-colored carbon atoms. Residues that ligate the active site zinc are shown with brown-colored carbon atoms. Simulated annealed composite omit map shown in blue mesh contoured at 1σ. Numbers next to yellow dashed lines represent ligation distances in Å. (Right) Simulated annealed omit map of the flipped out base. The 8-azanebularine nucleotide was omitted from the structure and a simulated annealed omit map calculated in PHENIX (Adams, P.D et al. Acta Crystallogr D Biol Crystallogr 66, 213-221, 2010). Positive electron density contoured at 3σ is displayed in green. This clearly shows the presence of oxygen O6 (extending from carbon 6) ligating to the active site zinc atom. This definitively reveals the hydrated intermediate is being trapped in the crystal. (b) Close up view of 5’ UA pair from (hADARd WT + Bdf2-U) shown with composite omit-map electron density at 1σ. The geometry of the 5’ nearest neighbor base pair from hADAR2d WT+ Bdf2-U (sticks) and electron density overlaid with structure of hADAR2d WT+ Bdf2-C (lines) suggests a non-canonical base pair between the A and U.
Supplementary Figure 4 Intercalating residue and adjacent RNA.
(a) Overlay of intercalating residue and RNA from three structures. Overlay of structures of wild type and E488Q hADAR2d with Bdf2-C RNA and wild type hADAR2d with Bdf2-U RNA show high degree of similarity in both RNA and enzyme conformations (b) Comparison of ADAR2 and HhaI Mtase orphan base recognition. Shown is the interaction between intercalating residue side chain and orphan nucleotide in ADAR2 (Left) and HhaI cytidine methyltransferase (Right). For ADAR2 the side chain approaches the nucleotide from the minor groove of the A form helix while for HhaI Mtase the approach is made from the major groove of the B form DNA (Klimasauskas, S. et al. Cell, 76, 357-369 1994).
Supplementary Figure 5 Comparison of protein–nucleic acid complexes with flipped nucleotides.
(a) Structures of DNA modifying enzymes with base flipping dependent mechanisms approaching from the minor groove. Protein in blue, DNA in yellow, which is bent, flipped nucleotide in red. (b) Structures of protein–RNA complexes with flipped nucleotides. Protein in blue, RN in pink, flipped nucleotide in red.
Supplementary Figure 6 Access of residue 488 into the minor grooves of A-form and B-form helices.
Overlay of intercalating residue 488 with idealized A form RNA (Left) shows that the geometry of the minor groove in an A form helix allows the side chain to occupy the position of the adenosine base (yellow). Overlay of residue 488 with idealized B form DNA (Right) shows that the greater depth of the minor groove in a B form helix prevents the side chain from intercalating into the space occupied by an adenosine base (yellow). Idealized A-RNA and B-DNA of Bdf2 sequence were superimposed onto the RNA duplex observed in the hADAR2d complexed with RNA (as shown in Figure 4B).
Supplementary Figure 7 Nearest-neighbor effects of hADAR2d.
(a) Predicted secondary structure of GLI1 mRNA editing site showing 3’ nearest neighbor and complementary mutations. Edited A is highlighted in red. (b) Comparison of deamination rates in 3’ nearest neighbor mutants in hGLI1 mRNA. Observed in vitro deamination rate constants for deamination of hGLI1 mRNA by hADAR2d and 3’ nearest neighbor mutants.
(c) Space filling models of 5’ nearest neighbor base pair and hADAR2d G489. Minor groove edge of U11-A13’ base pair is in close proximity to the protein backbone at G489 in the Bdf2-hADAR2d complex (top left). U11 is located on the 5’ side of the editing site (i.e. 5’ U). This site appears to accommodate a 5’ A (top right) but when either a C-G (bottom left) or a G-C (bottom right) base pair is modeled into this site, a clash is apparent between the G 2-amino group and the alpha carbon of G489.
Supplementary Figure 8 hADAR1 residues associated with AGS, mapped to analogous positions in hADAR2d.
(a) Space filling model of hADAR2d E488Q + Bdf2-C RNA highlighting the locations corresponding to residues in hADAR1 where mutations cause Aicardi–Goutières syndrome. Modeling positions of surface residues of hADAR1 with mutations previously associated with Aicardi–Goutières syndrome (Rice, G.I. et al. Nat Genetics, 44, 1243-1248, 2012) provides insight into the basis for the mutations’ effect on editing. hADAR2 residues corresponding to hADAR1 R892 and G1007 (K376 and G487 respectively) are found at the protein/RNA binding interface. K999 maps to ADAR2’s 5’ major groove binding loop (Q479). While Y1112 and D1113 are not directly involved in the protein/RNA interface, they appear to be in position to stabilize an RNA binding loop. E588 in hADAR2 (corresponding to D1113 in hADAR1) is involved in an intramolecular salt bridge with highly conserved R349 located adjacent to RNA contact residue R348. (b) hADAR2 G487 is located at the protein-RNA interface. Mutation to arginine (corresponding to the G1007R mutation in hADAR1 associated with the human diseases AGS and DHS) would cause a severe clash with the RNA, especially with orphan C11’ and A12’. The G487R mutation is shown with transparent CPK spheres with magenta-colored carbons.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Tables 1 and 2 (PDF 3473 kb)
RNA morph
This file contains a morphing movie of the change in conformation of the RNA before and after hADAR2d binding (assuming an ideal A-form duplex RNA starting point). (MOV 1557 kb)
hADAR2 morph
This file contains a morphing movie of the change in hADAR2d conformation before and after RNA binding. (MOV 988 kb)
Rights and permissions
About this article
Cite this article
Matthews, M., Thomas, J., Zheng, Y. et al. Structures of human ADAR2 bound to dsRNA reveal base-flipping mechanism and basis for site selectivity. Nat Struct Mol Biol 23, 426–433 (2016). https://doi.org/10.1038/nsmb.3203
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3203
This article is cited by
-
RNA modification: mechanisms and therapeutic targets
Molecular Biomedicine (2023)
-
Host-mediated RNA editing in viruses
Biology Direct (2023)
-
Precision RNA base editing with engineered and endogenous effectors
Nature Biotechnology (2023)
-
Novel insights into double-stranded RNA-mediated immunopathology
Nature Reviews Immunology (2023)
-
Compact RNA editors with small Cas13 proteins
Nature Biotechnology (2022)