Abstract
ATX is a plasma lysophospholipase D that hydrolyzes lysophosphatidylcholine (LPC) and produces lysophosphatidic acid. To date, no ATX-inhibition-mediated treatment strategies for human diseases have been established. Here, we report anti-ATX DNA aptamers that inhibit ATX with high specificity and efficacy. We solved the crystal structure of ATX in complex with the anti-ATX aptamer RB011, at 2.0-Å resolution. RB011 binds in the vicinity of the active site through base-specific interactions, thus preventing the access of the choline moiety of LPC substrates. Using the structural information, we developed the modified anti-ATX DNA aptamer RB014, which exhibited in vivo efficacy in a bleomycin-induced pulmonary fibrosis mouse model. Our findings reveal the structural basis for the specific inhibition of ATX by the anti-ATX aptamer and highlight the therapeutic potential of anti-ATX aptamers for the treatment of human diseases, such as pulmonary fibrosis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Tuerk, C. & Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249, 505–510 (1990).
Ellington, A.D. & Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 346, 818–822 (1990).
Miyakawa, S. et al. Structural and molecular basis for hyperspecificity of RNA aptamer to human immunoglobulin G. RNA 14, 1154–1163 (2008).
Wang, J. et al. Inhibition of midkine alleviates experimental autoimmune encephalomyelitis through the expansion of regulatory T cell population. Proc. Natl. Acad. Sci. USA 105, 3915–3920 (2008).
Kishida, S. et al. Midkine promotes neuroblastoma through Notch2 signaling. Cancer Res. 73, 1318–1327 (2013).
Dyke, C.K. et al. First-in-human experience of an antidote-controlled anticoagulant using RNA aptamer technology: a phase 1a pharmacodynamic evaluation of a drug-antidote pair for the controlled regulation of factor IXa activity. Circulation 114, 2490–2497 (2006).
Gilbert, J.C. et al. First-in-human evaluation of anti von Willebrand factor therapeutic aptamer ARC1779 in healthy volunteers. Circulation 116, 2678–2686 (2007).
Keefe, A.D., Pai, S. & Ellington, A. Aptamers as therapeutics. Nat. Rev. Drug Discov. 9, 537–550 (2010).
Nakamura, Y., Ishiguro, A. & Miyakawa, S. RNA plasticity and selectivity applicable to therapeutics and novel biosensor development. Genes Cells 17, 344–364 (2012).
Ng, E.W.M. et al. Pegaptanib, a targeted anti-VEGF aptamer for ocular vascular disease. Nat. Rev. Drug Discov. 5, 123–132 (2006).
Oberthür, D. et al. Crystal structure of a mirror-image L-RNA aptamer (Spiegelmer) in complex with the natural L-protein target CCL2. Nat. Commun. 6, 6923 (2015).
Nomura, Y. et al. Conformational plasticity of RNA for target recognition as revealed by the 2.15 A crystal structure of a human IgG-aptamer complex. Nucleic Acids Res. 38, 7822–7829 (2010).
Jarvis, T.C. et al. Non-helical DNA triplex forms a unique aptamer scaffold for high affinity recognition of nerve growth factor. Structure 23, 1293–1304 (2015).
Umezu-Goto, M. et al. Autotaxin has lysophospholipase D activity leading to tumor cell growth and motility by lysophosphatidic acid production. J. Cell Biol. 158, 227–233 (2002).
Tokumura, A. et al. Identification of human plasma lysophospholipase D, a lysophosphatidic acid-producing enzyme, as autotaxin, a multifunctional phosphodiesterase. J. Biol. Chem. 277, 39436–39442 (2002).
Noguchi, K., Herr, D., Mutoh, T. & Chun, J. Lysophosphatidic acid (LPA) and its receptors. Curr. Opin. Pharmacol. 9, 15–23 (2009).
Moolenaar, W.H., van Meeteren, L.A. & Giepmans, B.N.G. The ins and outs of lysophosphatidic acid signaling. BioEssays 26, 870–881 (2004).
Pamuklar, Z. et al. Autotaxin/lysopholipase D and lysophosphatidic acid regulate murine hemostasis and thrombosis. J. Biol. Chem. 284, 7385–7394 (2009).
Kanda, H. et al. Autotaxin, an ectoenzyme that produces lysophosphatidic acid, promotes the entry of lymphocytes into secondary lymphoid organs. Nat. Immunol. 9, 415–423 (2008).
Contos, J.J., Fukushima, N., Weiner, J.A., Kaushal, D. & Chun, J. Requirement for the lpA1 lysophosphatidic acid receptor gene in normal suckling behavior. Proc. Natl. Acad. Sci. USA 97, 13384–13389 (2000).
Tanaka, M. et al. Autotaxin stabilizes blood vessels and is required for embryonic vasculature by producing lysophosphatidic acid. J. Biol. Chem. 281, 25822–25830 (2006).
van Meeteren, L.A. et al. Autotaxin, a secreted lysophospholipase D, is essential for blood vessel formation during development. Mol. Cell. Biol. 26, 5015–5022 (2006).
Inoue, M. et al. Initiation of neuropathic pain requires lysophosphatidic acid receptor signaling. Nat. Med. 10, 712–718 (2004).
Yang, S.Y. et al. Expression of autotaxin (NPP-2) is closely linked to invasiveness of breast cancer cells. Clin. Exp. Metastasis 19, 603–608 (2002).
Kishi, Y. et al. Autotaxin is overexpressed in glioblastoma multiforme and contributes to cell motility of glioblastoma by converting lysophosphatidylcholine to lysophosphatidic acid. J. Biol. Chem. 281, 17492–17500 (2006).
Tager, A.M. et al. The lysophosphatidic acid receptor LPA1 links pulmonary fibrosis to lung injury by mediating fibroblast recruitment and vascular leak. Nat. Med. 14, 45–54 (2008).
Oikonomou, N. et al. Pulmonary autotaxin expression contributes to the pathogenesis of pulmonary fibrosis. Am. J. Respir. Cell Mol. Biol. 47, 566–574 (2012).
Gierse, J. et al. A novel autotaxin inhibitor reduces lysophosphatidic acid levels in plasma and the site of inflammation. J. Pharmacol. Exp. Ther. 334, 310–317 (2010).
Albers, H.M.H.G. et al. Structure-based design of novel boronic acid-based inhibitors of autotaxin. J. Med. Chem. 54, 4619–4626 (2011).
Kawaguchi, M. et al. Screening and X-ray crystal structure-based optimization of autotaxin (ENPP2) inhibitors, using a newly developed fluorescence probe. ACS Chem. Biol. 8, 1713–1721 (2013).
Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003).
Burmeister, P.E. et al. Direct in vitro selection of a 2′-O-methyl aptamer to VEGF. Chem. Biol. 12, 25–33 (2005).
Nishimasu, H. et al. Crystal structure of autotaxin and insight into GPCR activation by lipid mediators. Nat. Struct. Mol. Biol. 18, 205–212 (2011).
Hausmann, J. et al. Structural basis of substrate discrimination and integrin binding by autotaxin. Nat. Struct. Mol. Biol. 18, 198–204 (2011).
Saga, H. et al. A novel highly potent autotaxin/ENPP2 inhibitor produces prolonged decreases in plasma lysophosphatidic acid formation in vivo and regulates urethral tension. PLoS One 9, e93230 (2014).
Tabata, S. et al. A rapid screening method for cell lines producing singly-tagged recombinant proteins using the “TARGET tag” system. J. Proteomics 73, 1777–1785 (2010).
Kato, K. et al. Expression, purification, crystallization and preliminary X-ray crystallographic analysis of Enpp1. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 68, 778–782 (2012).
Tokuhara, Y. et al. A new enzyme immunoassay for the quantitative determination of classical autotaxins (ATXα, ATXβ, and ATXγ) and novel autotaxins (ATXδ and ATXɛ). PLoS One 10, e0130074 (2015).
Fitter, S. & James, R. Deconvolution of a complex target using DNA aptamers. J. Biol. Chem. 280, 34193–34201 (2005).
Kishimoto, T., Matsuoka, T., Imamura, S. & Mizuno, K. A novel colorimetric assay for the determination of lysophosphatidic acid in plasma using an enzymatic cycling method. Clin. Chim. Acta 333, 59–67 (2003).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Vagin, A. & Teplyakov, A. Molecular replacement with MOLREP. Acta Crystallogr. D Biol. Crystallogr. 66, 22–25 (2010).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
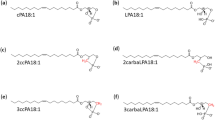
Okudaira, M. et al. Separation and quantification of 2-acyl-1-lysophospholipids and 1-acyl-2-lysophospholipids in biological samples by LC-MS/MS. J. Lipid Res. 55, 2178–2192 (2014).
Acknowledgements
We thank the beamline staff at BL32XU and BL41XU of SPring-8, Japan, for assistance with data collection. We thank T. Kishimoto for the LPA assays, S. Yamazaki for in vitro assays, E. Inomata for aptamer stability experiments and all other Ribomic members for discussions. We thank T. Nagano (University of Tokyo) for 3BoA. This work was supported by a grant from the Core Research for Evolutional Science and Technology Program, the Creation of Basic Chronic Inflammation, from the Japan Science and Technology Agency, to O.N. This work was also supported in part by a grant from NEDO.
Author information
Authors and Affiliations
Contributions
K. Kato prepared and crystallized the protein and determined the crystal structure; H.I., S.F., Y. Nonaka, M.F., S.M., S.O., K. Kano and J.A. performed in vitro and in vivo experiments; H.I. and S.M. designed aptamers; J.M. prepared the protein; R.I. and H.N. assisted with the structural analysis; and K. Kato, R.I., H.N., Y. Nakamura and O.N. wrote the manuscript with help from all authors. Y. Nakamura and O.N. directed and supervised all of the research.
Corresponding authors
Ethics declarations
Competing interests
Except for K. Kato, J.A., J.M., R.I., H.N. and O.N., all authors are either employees of Ribomic Inc. and/or hold equity in Ribomic Inc. The remaining authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Characterization of RB011.
a, b, RB011 binding to ATX measured by SPR. Sensorgrams of different concentrations of RB011 (50, 25, 12.5, 6.25 and 3.12 nM from upper to lower sensorgrams) binding to human ATX (a) and mouse ATX (b). c, Sensorgrams of RB011 binding to mouse ATX (ENPP2) and ENPP1. d, Plasma pharmacokinetic profiles of the PEGylated RB011. The PEGylated RB011 was administered intravenously, subcutaneously, or intraperitoneally to C57BL/6J mice at 10 mg/kg. Data are mean ± S.E.M. (n = 3, number of mice). e, Pharmacokinetic parameters.
Supplementary Figure 2 Electron density map.
The 2mFO – DFC electron density map around the ATX–RB011 interface is shown as a gray mesh, contoured at 1σ. Water molecules are shown as red spheres.
Supplementary Figure 3 Multiple sequence alignment of ATX proteins from different animal species.
The zinc-coordinating and catalytic residues, gray triangles; the residues interacting with the stem loop, red triangles; the residues interacting with the corner junction, yellow triangles; and the residues interacting with the terminal stem, blue triangles.
Supplementary Figure 4 Multiple sequence alignment of the mouse ENPP family proteins.
The zinc-coordinating and catalytic residues and the residues interacting with RB011 are shown as in Supplementary Fig. 3.
Supplementary Figure 5 Recognition of ATX by RB011.
a, The catalytic domain of ATX in complex with RB011. b, The catalytic domain of ENPP1 with RB011 (model). The ATX–RB011 complex was superimposed on ENPP1 (PDB ID 4GTW). The yellow circle indicates predicted steric clashes between RB011 and a loop region (red) in the insertion subdomain. c, Sensorgrams of RB011 binding to the wild type and mutants of mouse ATX. d, Inhibition of the LysoPLD activities of the wild type and mutants of mouse ATX by RB011 (0.1 μM). Data are mean ± S.D (n = 4). Results are from two independent experiments with two technical replicates each.
Supplementary Figure 6 Structure-based optimization of anti-ATX aptamers.
a, Inhibition of LPA production in human serum after 3 h incubation at 37°C in the presence of 1 or 0.1 μM of the indicated inhibitors. Data are mean ± S.D. (n = 3, number of samples). b, Lung pharmacokinetic profiles of RB014. RB014 was administered intranasally to C57BL/6J mice at 20 μg/mouse. Data are mean ± S.D. (n = 3, number of mice). c, Inhibition of ATX activity by RB014 in BALF of BLM-induced pulmonary fibrosis model mice. The BALF of BLM-induced pulmonary fibrosis model mice was prepared, as in the experiments in Fig. 5, and then the LysoPLD activities were monitored. Data are mean ± S.E.M. (n = 5, number of mice). d, Inhibition of LPA production by RB014 in BALF of BLM-induced pulmonary fibrosis model mice. RB014 was administered intranasally to C57BL/6J mice at 20 μg/mouse. Data are mean ± S.E.M. (n = 5, number of mice). The variances between the groups were tested by the f-test. The p value was obtained by the Student’s t-test for the 18:2-LPA levels of Day 7.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Tables 1 and 2 (PDF 12936 kb)
Rights and permissions
About this article
Cite this article
Kato, K., Ikeda, H., Miyakawa, S. et al. Structural basis for specific inhibition of Autotaxin by a DNA aptamer. Nat Struct Mol Biol 23, 395–401 (2016). https://doi.org/10.1038/nsmb.3200
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3200