Abstract
The Ca2+ sensor synaptotagmin-1 is thought to trigger membrane fusion by binding to acidic membrane lipids and SNARE proteins. Previous work has shown that binding is mediated by electrostatic interactions that are sensitive to the ionic environment. However, the influence of divalent or polyvalent ions, at physiological concentrations, on synaptotagmin's binding to membranes or SNAREs has not been explored. Here we show that binding of rat synaptotagmin-1 to membranes containing phosphatidylinositol 4,5-bisphosphate (PIP2) is regulated by charge shielding caused by the presence of divalent cations. Surprisingly, polyvalent ions such as ATP and Mg2+ completely abrogate synaptotagmin-1 binding to SNAREs regardless of the presence of Ca2+. Altogether, our data indicate that at physiological ion concentrations Ca2+-dependent synaptotagmin-1 binding is confined to PIP2-containing membrane patches in the plasma membrane, suggesting that membrane interaction of synaptotagmin-1 rather than SNARE binding triggers exocytosis of vesicles.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Südhof, T.C. Neurotransmitter release: the last millisecond in the life of a synaptic vesicle. Neuron 80, 675–690 (2013).
Jahn, R. & Fasshauer, D. Molecular machines governing exocytosis of synaptic vesicles. Nature 490, 201–207 (2012).
Pobbati, A.V., Stein, A. & Fasshauer, D. N- to C-terminal SNARE complex assembly promotes rapid membrane fusion. Science 313, 673–676 (2006).
Perin, M.S., Fried, V.A., Mignery, G.A., Jahn, R. & Sudhof, T.C. Phospholipid binding by a synaptic vesicle protein homologous to the regulatory region of protein kinase C. Nature 345, 260–263 (1990).
Brose, N., Petrenko, A.G., Sudhof, T.C. & Jahn, R. Synaptotagmin: a calcium sensor on the synaptic vesicle surface. Science 256, 1021–1025 (1992).
Davletov, B.A. & Sudhof, T.C. A single C2 domain from synaptotagmin I is sufficient for high affinity Ca2+/phospholipid binding. J. Biol. Chem. 268, 26386–26390 (1993).
Herrick, D.Z., Sterbling, S., Rasch, K.A., Hinderliter, A. & Cafiso, D.S. Position of synaptotagmin I at the membrane interface: cooperative interactions of tandem C2 domains. Biochemistry 45, 9668–9674 (2006).
Fernandez, I. et al. Three-dimensional structure of the synaptotagmin 1 C2B-domain: synaptotagmin 1 as a phospholipid binding machine. Neuron 32, 1057–1069 (2001).
Araç, D. et al. Close membrane-membrane proximity induced by Ca2+-dependent multivalent binding of synaptotagmin-1 to phospholipids. Nat. Struct. Mol. Biol. 13, 209–217 (2006).
Wang, Z., Liu, H., Gu, Y. & Chapman, E.R. Reconstituted synaptotagmin I mediates vesicle docking, priming, and fusion. J. Cell Biol. 195, 1159–1170 (2011).
Honigmann, A. et al. Phosphatidylinositol 4,5-bisphosphate clusters act as molecular beacons for vesicle recruitment. Nat. Struct. Mol. Biol. 20, 679–686 (2013).
van den Bogaart, G. et al. Membrane protein sequestering by ionic protein-lipid interactions. Nature 479, 552–555 (2011).
Suh, B.C. & Hille, B. PIP2 is a necessary cofactor for ion channel function: how and why? Annu. Rev. Biophys. 37, 175–195 (2008).
Tucker, S.J. & Baukrowitz, T. How highly charged anionic lipids bind and regulate ion channels. J. Gen. Physiol. 131, 431–438 (2008).
McLaughlin, S. & Murray, D. Plasma membrane phosphoinositide organization by protein electrostatics. Nature 438, 605–611 (2005).
Helmholtz, H. Ueber einige Gesetze der Vertheilung elektrischer Ströme in körperlichen Leitern mit Anwendung auf die thierischelektrischen Versuche. Pogg. Ann. Physik Chemie 89, 211–233 (1853).
Manning, G.S. & Ray, J. Counterion condensation revisited. J. Biomol. Struct. Dyn. 16, 461–476 (1998).
Delgado, A.V. et al. Measurement and interpretation of electrokinetic phenomena. J. Colloid Interface Sci. 309, 194–224 (2007).
Schiavo, G., Stenbeck, G., Rothman, J.E. & Sollner, T.H. Binding of the synaptic vesicle v-SNARE, synaptotagmin, to the plasma membrane t-SNARE, SNAP-25, can explain docked vesicles at neurotoxin-treated synapses. Proc. Natl. Acad. Sci. USA 94, 997–1001 (1997).
Rickman, C. & Davletov, B. Mechanism of calcium-independent synaptotagmin binding to target SNAREs. J. Biol. Chem. 278, 5501–5504 (2003).
Rickman, C. et al. Conserved prefusion protein assembly in regulated exocytosis. Mol. Biol. Cell 17, 283–294 (2006).
Chicka, M.C., Hui, E., Liu, H. & Chapman, E.R. Synaptotagmin arrests the SNARE complex before triggering fast, efficient membrane fusion in response to Ca2+. Nat. Struct. Mol. Biol. 15, 827–835 (2008).
Brewer, K.D. et al. Dynamic binding mode of a Synaptotagmin-1–SNARE complex in solution. Nat. Struct. Mol. Biol. 22, 555–564 (2015).
Li, L. et al. Phosphatidylinositol phosphates as co-activators of Ca2+ binding to C2 domains of synaptotagmin 1. J. Biol. Chem. 281, 15845–15852 (2006).
Park, Y. et al. Controlling synaptotagmin activity by electrostatic screening. Nat. Struct. Mol. Biol. 19, 991–997 (2012).
Chapman, E.R. How does synaptotagmin trigger neurotransmitter release? Annu. Rev. Biochem. 77, 615–641 (2008).
Rizo, J. & Rosenmund, C. Synaptic vesicle fusion. Nat. Struct. Mol. Biol. 15, 665–674 (2008).
Choi, U.B. et al. Single-molecule FRET–derived model of the synaptotagmin 1–SNARE fusion complex. Nat. Struct. Mol. Biol. 17, 318–324 (2010).
Tang, J. et al. A complexin/synaptotagmin 1 switch controls fast synaptic vesicle exocytosis. Cell 126, 1175–1187 (2006).
Bai, J., Tucker, W.C. & Chapman, E.R. PIP2 increases the speed of response of synaptotagmin and steers its membrane-penetration activity toward the plasma membrane. Nat. Struct. Mol. Biol. 11, 36–44 (2004).
Radhakrishnan, A., Stein, A., Jahn, R. & Fasshauer, D. The Ca2+ affinity of synaptotagmin 1 is markedly increased by a specific interaction of its C2B domain with phosphatidylinositol 4,5-bisphosphate. J. Biol. Chem. 284, 25749–25760 (2009).
Lee, H.K. et al. Dynamic Ca2+-dependent stimulation of vesicle fusion by membrane-anchored synaptotagmin 1. Science 328, 760–763 (2010).
Toner, M., Vaio, G., McLaughlin, A. & McLaughlin, S. Adsorption of cations to phosphatidylinositol 4,5-bisphosphate. Biochemistry 27, 7435–7443 (1988).
Suh, B.C. & Hille, B. Electrostatic interaction of internal Mg2+ with membrane PIP2 seen with KCNQ K+ channels. J. Gen. Physiol. 130, 241–256 (2007).
Knight, D.E. & Baker, P.F. Calcium-dependence of catecholamine release from bovine adrenal medullary cells after exposure to intense electric fields. J. Membr. Biol. 68, 107–140 (1982).
Wegenhorst, U., Gratzl, M., Fohr, K.J. & Ahnert-Hilger, G. Millimolar concentrations of free magnesium enhance exocytosis from permeabilized rat pheochromocytoma (PC 12) cells. Neurosci. Lett. 106, 300–304 (1989).
Koh, D.S. Carbon fiber amperometry in the study of ion channels and secretion. Methods Mol. Biol. 337, 139–153 (2006).
Kuo, W., Herrick, D.Z., Ellena, J.F. & Cafiso, D.S. The calcium-dependent and calcium-independent membrane binding of synaptotagmin 1: two modes of C2B binding. J. Mol. Biol. 387, 284–294 (2009).
Vrljic, M. et al. Post-translational modifications and lipid binding profile of insect cell-expressed full-length mammalian synaptotagmin 1. Biochemistry 50, 9998–10012 (2011).
Jeschke, G., Koch, A., Jonas, U. & Godt, A. Direct conversion of EPR dipolar time evolution data to distance distributions. J. Magn. Reson. 155, 72–82 (2002).
Pannier, M., Veit, S., Godt, A., Jeschke, G. & Spiess, H.W. Dead-time free measurement of dipole-dipole interactions between electron spins. J. Magn. Reson. 142, 331–340 (2000).
Beis, I. & Newsholme, E.A. The contents of adenine nucleotides, phosphagens and some glycolytic intermediates in resting muscles from vertebrates and invertebrates. Biochem. J. 152, 23–32 (1975).
London, R.E. Methods for measurement of intracellular magnesium: NMR and fluorescence. Annu. Rev. Physiol. 53, 241–258 (1991).
Hess, P., Metzger, P. & Weingart, R. Free magnesium in sheep, ferret and frog striated muscle at rest measured with ion-selective micro-electrodes. J. Physiol. (Lond.) 333, 173–188 (1982).
Rickman, C. et al. Synaptotagmin interaction with the syntaxin/SNAP-25 dimer is mediated by an evolutionarily conserved motif and is sensitive to inositol hexakisphosphate. J. Biol. Chem. 279, 12574–12579 (2004).
Gaffaney, J.D., Dunning, F.M., Wang, Z., Hui, E. & Chapman, E.R. Synaptotagmin C2B domain regulates Ca2+-triggered fusion in vitro: critical residues revealed by scanning alanine mutagenesis. J. Biol. Chem. 283, 31763–31775 (2008).
Vennekate, W. et al. Cis- and trans-membrane interactions of synaptotagmin-1. Proc. Natl. Acad. Sci. USA 109, 11037–11042 (2012).
Shao, X. et al. Synaptotagmin-syntaxin interaction: the C2 domain as a Ca2+-dependent electrostatic switch. Neuron 18, 133–142 (1997).
Sarhan, S. & Seiler, N. On the subcellular localization of the polyamines. Biol. Chem. Hoppe Seyler 370, 1279–1284 (1989).
Dai, H., Shen, N., Arac, D. & Rizo, J. A quaternary SNARE-synaptotagmin-Ca2+-phospholipid complex in neurotransmitter release. J. Mol. Biol. 367, 848–863 (2007).
Lynch, K.L. et al. Synaptotagmin C2A loop 2 mediates Ca2+-dependent SNARE interactions essential for Ca2+-triggered vesicle exocytosis. Mol. Biol. Cell 18, 4957–4968 (2007).
Lai, A.L., Huang, H., Herrick, D.Z., Epp, N. & Cafiso, D.S. Synaptotagmin 1 and SNAREs form a complex that is structurally heterogeneous. J. Mol. Biol. 405, 696–706 (2011).
Lai, Y., Lou, X., Diao, J. & Shin, Y.K. Molecular origins of synaptotagmin 1 activities on vesicle docking and fusion pore opening. Sci. Rep. 5, 9267 (2015).
Lai, A.L., Tamm, L.K., Ellena, J.F. & Cafiso, D.S. Synaptotagmin 1 modulates lipid acyl chain order in lipid bilayers by demixing phosphatidylserine. J. Biol. Chem. 286, 25291–25300 (2011).
Martens, S., Kozlov, M.M. & McMahon, H.T. How synaptotagmin promotes membrane fusion. Science 316, 1205–1208 (2007).
Hui, E., Johnson, C.P., Yao, J., Dunning, F.M. & Chapman, E.R. Synaptotagmin-mediated bending of the target membrane is a critical step in Ca2+-regulated fusion. Cell 138, 709–721 (2009).
McMahon, H.T., Kozlov, M.M. & Martens, S. Membrane curvature in synaptic vesicle fusion and beyond. Cell 140, 601–605 (2010).
van den Bogaart, G. et al. Synaptotagmin-1 may be a distance regulator acting upstream of SNARE nucleation. Nat. Struct. Mol. Biol. 18, 805–812 (2011).
Imig, C. et al. The morphological and molecular nature of synaptic vesicle priming at presynaptic active zones. Neuron 84, 416–431 (2014).
Jeschke, G. et al. DeerAnalysis2006: a comprehensive software package for analyzing pulsed ELDOR data. Appl. Magn. Reson. 30, 473–498 (2006).
Weber, T. et al. SNAREpins: minimal machinery for membrane fusion. Cell 92, 759–772 (1998).
Stein, A., Radhakrishnan, A., Riedel, D., Fasshauer, D. & Jahn, R. Synaptotagmin activates membrane fusion through a Ca2+-dependent trans interaction with phospholipids. Nat. Struct. Mol. Biol. 14, 904–911 (2007).
Park, Y. et al. α-SNAP interferes with the zippering of the SNARE protein membrane fusion machinery. J. Biol. Chem. 289, 16326–16335 (2014).
Zhou, Z. & Misler, S. Action potential-induced quantal secretion of catecholamines from rat adrenal chromaffin cells. J. Biol. Chem. 270, 3498–3505 (1995).
Pannier, M., Veit, S., Godt, A., Jeschke, G. & Spiess, H.W. Dead-time free measurement of dipole-dipole interactions between electron spins. J. Magn. Reson. 142, 331–340 (2000).
Stein, A., Weber, G., Wahl, M.C. & Jahn, R. Helical extension of the neuronal SNARE complex into the membrane. Nature 460, 525–528 (2009).
Warshaviak, D.T., Serbulea, L., Houk, K.N. & Hubbell, W.L. Conformational analysis of a nitroxide side chain in an α-helix with density functional theory. J. Phys. Chem. B 115, 397–405 (2011).
Polyhach, Y., Bordignon, E. & Jeschke, G. Rotamer libraries of spin labelled cysteines for protein studies. Phys. Chem. Chem. Phys. 13, 2356–2366 (2011).
Cypionka, A. et al. Discrimination between docking and fusion of liposomes reconstituted with neuronal SNARE-proteins using FCS. Proc. Natl. Acad. Sci. USA 106, 18575–18580 (2009).
Fogolari, F., Brigo, A. & Molinari, H. The Poisson-Boltzmann equation for biomolecular electrostatics: a tool for structural biology. J. Mol. Recognit. 15, 377–392 (2002).
Li, C., Li, L., Petukh, M. & Alexov, E. Progress in developing Poisson-Boltzmann equation solvers. Mol. Based Math. Biol. 1, 42–62 (2013).
Hill, A.V. The combinations of haemoglobin with oxygen and with carbon monoxide. I. Biochem. J. 7, 471–480 (1913).
Augustine, G.J. & Neher, E. Calcium requirements for secretion in bovine chromaffin cells. J. Physiol. (Lond.) 450, 247–271 (1992).
Acknowledgements
We thank B. Hille for helpful discussion. This research was supported by US National Institutes of Health grant DK080840 (to D.-S.K.); the National Research Foundation of Korea through the Basic Science Research Program, grants 2012R1A2A4A01004419 and 2011-0012331 (to M.C.); and US National Institutes of Health grant 2 P01 GM072694 (to R.J. and D.S.C.).
Author information
Authors and Affiliations
Contributions
Y.P. designed and performed most of the biochemical and biophysical study, including fusion assays, FRET, floatation assays and anisotropy. Y.P. collected and analyzed data. J.B.S., under the supervision of D.-S.K., carried out dual recording of amperometry and patch-clamping. A.F. and D.S.C. performed DEER. K.H., M.C. and S.-R.J. developed the mathematical model and performed numerical simulation. A.P.-L. carried out stopped-flow spectroscopy. I.K., under the supervision of P.J.W., performed fluorescence cross-correlation spectroscopy. H.Y. assisted with protein labeling. Y.P., K.H., M.C., D.S.C., D.-S.K. and R.J. wrote the manuscript. All authors read and provided their comments on the draft. D.S.C., D.-S.K. and R.J. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 High PIP2 concentrations but no SNAREs are required for Ca2+-independent binding of the C2AB domain.
A labeled variant of the C2AB fragment (Syt97-421, Alexa Fluor 488-labeled at S342C) was used to monitor binding to liposomes (Lip.) by fluorescence anisotropy. 50 μM EGTA was included in all incubations. (a) Increasing Ca2+ concentration from 1 mM to 15 mM had no effects on anisotropy signal in the absence of liposomes. (b) No binding was observable to liposomes containing 1% PIP2 and the Q-SNARE complex consisting of full-length syntaxin-1A (1-288) and SNAP-25A (Syx-SN25). (c) Binding was observable to protein-free liposomes containing 5% PIP2, which was reversed by the addition of 1 mM MgCl2/ATP. a.u., arbitrary unit.
Supplementary Figure 2 Inhibitory effect of Mg2+ on Ca2+-independent binding of the C2AB to liposomes.
Fluorescence anisotropy as given in Supplementary Fig. 1 for Ca2+-independent binding of the C2AB. (a–c) Liposomes (Lip.) and 30 μM free Ca2+ were subsequently added to observe Ca2+-independent and Ca2+-dependent binding of C2AB domain, respectively. Mg2+ and ATP were absent in this reaction. WT, wild-type C2AB; KAKA, a mutant containing alanine substitutions of two cationic lysine residues in the polybasic region (K326A and K327A). (c) C2AB binding was plotted as a net change of anisotropy signal (no MgCl2). (d,e) Increasing MgCl2 concentrations reduced Ca2+-independent binding of C2AB domain. (e) C2AB binding was presented as percentage of maximum value of C2AB binding in the absence of Mg2+. Data for c,e are mean ± SD (n=4~5).
Supplementary Figure 3 Effects of Mg2+, spermine and neomycin on exocytosis of chromaffin granules (CGs).
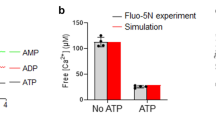
(a) CG fusion using a lipid-mixing assay. Purified native CGs were incubated with proteoliposomes reconstituted with a stabilized Q-SNARE complex (see Fig. 1 for details). The free Ca2+ concentration was adjusted to 100 μM in the presence of 1 mM MgCl2 and 3 mM ATP using the Maxchelator simulation program (http://maxchelator.stanford.edu). ATP was also included in control (without Ca2+). (b) Chromaffin cells were patched with internal solutions containing 50 μM free Ca2+ and different concentrations of Mg2+, spermine, or neomycin. All solutions except 15 mM MgCl2 solution contained 1 mM MgCl2. The exocytotic events were detected by a carbon fiber microelectrode and the number of events within 30 s-time window was counted over 8 min. Average rate of exocytosis was analyzed as events per 30 s by counting exocytotic events for 7.5 min (0.5 - 8 min after whole-cell patch clamping). To evaluate the effect of the cations after equilibrium between the patch pipette and cytoplasm, we additionally analyzed the time period between 5 and 8 min whole-cell patch clamping (for 3 min). Number of cells measured was indicated in brackets. ** P < 0.01 and *** P < 0.001 compared to control (1 mM MgCl2) without spermine or neomycin.
Supplementary Figure 4 Direct interaction of ATP with the polybasic region of the C2B domain.
(a, b) FRET measurements in which the C2AB fragment (Syt97-421) and ATP were labeled with Alexa Fluor 488 at S342C (green) and BODIPY Texas Red (red) as a donor and an acceptor, respectively (see Online Methods for details). Donor fluorescence was quenched by increasing BODIPY Texas Red-labeled ATP concentrations (a), whereas no quenching was observed with the KAKA-mutant (b). The fluorescence at 620 nm was from BODIPY Texas Red-ATP. Polybasic region is shown in blue and Ca2+ in magenta. (c,d) Binding was also observed when ATP labeled with a different dye (Alexa Fluor 647) was used. Shown are emission spectra of donor fluorescence originating from Alexa Fluor 488-labeled C2AB. Mg2+ was not included. a.u., arbitrary unit.
Supplementary Figure 5 Interaction of synaptotagmin-1 and the SNARE complex, monitored by DEER.
(a) Six additional sets of Q-band DEER data obtained from the 1:1 complex formed from C2AB of synaptotagmin-1 (syt1) and the core SNAREs each at a concentration of 50 µM in the presence of Ca2+. The time domain signals (grey traces, left panels) were obtained from a single label in the core SNARE complex at either site 228 or 238 in syntaxin-1A H3 and a second label in the C2B domain of synaptotagmin-1. The distance distribution that yielded the best fit (blue trace) is shown in the corresponding panel to the right. The grey error bars in the distance distributions represent uncertainty due to the background subtraction form factor that produce fits within 15% RMSD of the best fit. The R1 label was incorporated into surface exposed sites of the SNAREs to minimize interference by the label, and efficient labeling of the SNARE proteins was confirmed by mass spectrometry. In every case, the distributions are broad and indicate that there is no specific mode of binding between synaptotagmin-1 and the core SNARE complex under normal ionic strength conditions. No interactions were detected between C2AB and site 211R1 in syntaxin on the N-terminal side of the core SNARE complex. The red histograms represent predictions based on the model generated from single-molecule FRET data (Choi, U.B. et al., Nat Struct Mol Biol 17, 318-24, 2010). These predictions are specific to the R1 side chain and take into account the sterically available conformers of R1 at the labeled sites (Hagelueken, G. et al., Appl Magn Reson 42, 377-391, 2012). (b) DEER measurements were run on three mutants of the core SNARE complex each containing 1 pair of spin labels. One spin label was incorporated into site 48 on SNAP-25A, with a second label at site 194, 228 or 242 of the syntaxin-1A SNARE motif. The time domain DEER signals (grey traces) are shown in the left panels along with the best fits (red traces) using the model-free approach in DeerAnalysis (Jeschke, G. et al., Applied Magnetic Resonance 30, 473-498, 2006). The corresponding panels on the right show the experimental distance distributions that represent the best fits (blue traces) to the data. Also shown (grey error bars) is the uncertainty in the distribution based upon variation in the background subtraction form that produces a fit within 15% RMSD of the best fit. The red histograms in the distribution represent predictions based upon the crystal structure of the core SNARE complex (PDB ID: 1SFC). These were obtained using the PyMol plugin MTSSL Wizard (Hagelueken, G. et al., Appl Magn Reson 42, 377-391, 2012). The distances measured between SNAP-25A 48R1 and either syntaxin-1A 194R1 or 228R1 agree well with the predictions from the crystal structure and indicate that the SNARE complex is well-formed. The last pair in the figure (syntaxin-1A 242R1/SNAP25A 48R1) produced the correct distance, but with a broader distribution than expected, suggesting that C-terminal end of the SNARE complex may have some disorder. (c) DEER distributions for the synaptotagmin-1–SNARE interaction plotted along with predictions of distance distributions based upon the 5 MD-derived structures in PDB ID: 2N1T (Brewer, K.D. et al., Nat Struct Mol Biol 22, 555-64, 2015). The distribution with the best fit to the DEER data is shown in blue and the grey shading indicates the range of distributions that produce fits within 15% RMSD of the best fit. The histograms (magenta) represent the expected distance distributions generated from the heterogeneous synaptotagmin-1 C2B–SNARE complex included in PDB ID: 2N1T (Brewer, K.D. et al., Nat Struct Mol Biol 22, 555-64, 2015). The distributions were calculated from a DFT based rotamer library (Warshaviak, D.T.. et al., J Phys Chem B 115, 397-405, 2011) using the program MMM (Polyhach, Y.. et al., Phys Chem Chem Phys 13, 2356-66, 2011). In most cases, this MD simulation captures the shorter range of these distributions but fails to capture the longer distance components.
Supplementary Figure 6 No C2AB-SNARE interaction at physiological ionic strength.
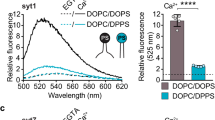
(a-d) FRET experiments to monitor C2AB binding to syntaxin-1A, in which C2AB fragment and full-length syntaxin-1A (1-288) were labeled with Alexa Fluor 488 at S342C (green) and Alexa Fluor 594 at T197C (magenta) as a donor and an acceptor, respectively. Syntaxin-1A was incorporated in liposomes containing 80% PC and 20% PE (no acidic phospholipids). Full-length syntaxin-1A is shown in green and SNARE motif in red. (a) Ca2+-independent and -dependent (1 mM free Ca2+) C2AB binding to syntaxin-1A was exaggerated at low ionic strength (50 mM KCl), compared to normal ionic strength (150 mM KCl, b). (c,d) C2AB binding to syntaxin-1A was completely disrupted by 1 mM MgCl2 and 3 mM ATP. Donor fluorescence was normalized as percentage of maximum value with C2AB alone. Shown are the typical traces of donor emission fluorescence from 3 to 5 independent experiments. PDB ID: 3C98 for syntaxin-1A. (e-h) No C2AB-SNARE interaction even in the presence of complexin II. FRET experiments to monitor C2AB binding to the SNARE complex, in which C2AB fragment and SNAP-25A were labeled with Alexa Fluor 488 at 342C (green) and Texas Red at 130C (magenta) as a donor and an acceptor, respectively. The SNARE complex consisting of synaptobrevin-2 (1-96, blue), SNAP-25A (green), and syntaxin-1A (183–288, red) was incorporated in liposomes containing 80% PC and 20% PE. (e,f) Ca2+-independent and -dependent (1 mM free Ca2+) C2AB binding to SNAREs was observed at low ionic strength (50 mM KCl) in the absence or presence of 1 μM complex II. (g,h) C2AB binding to SNAREs was not observed at physiological ionic strength (150 mM KCl, 1 mM MgCl2, and 3 mM ATP) regardless of whether complexin II was present or not. Donor fluorescence was normalized as percentage of maximum value with C2AB alone. Structures were visualized with the program PyMOL (PDB ID: 1KIL for complexin and 1SFC for SNARE complex).
Supplementary Figure 7 No C2AB-SNARE interaction at physiological ionic strength.
(a) Liposome co-flotation assay to test the C2AB-SNARE interaction. Physiological ionic environment (150 mM KCl, 1 mM MgCl2, and 5 mM ATP) completely reduced the C2AB interaction with the SNAREs in buffer fraction regardless of whether Ca2+ was present or not. (b,c) Fluorescence cross-correlation spectroscopy (FCCS) to test liposome tethering by proteins. (b) Incubation of liposomes (synaptotagmin-1, 80% PC, 19% PE, and 1% Oregon Green-DHPE) with 1% Texas Red-containing liposomes which incorporate the SNARE complex (full-length syntaxin-1A, SNAP-25A, and soluble synaptobrevin-2 (aa 49-96)) led to liposome tethering in low ionic strength (50 mM KCl). Physiological ionic strength (150 mM KCl/1 mM MgCl2/5 mM ATP) reduced the SNARE-synaptotagmin-1 interaction, independently of Ca2+. (c) SNARE-free liposomes containing 10% PS, 1% PIP2, and 1% Texas Red-DHPE (see Online Methods for details of lipid composition) was incubated with synaptotagmin-1-containing liposomes. 100 μM Ca2+ specifically increased liposome tethering in a physiological ionic strength (150 mM KCl, 1 mM MgCl2, and 5 mM ATP). Data are mean ± SD (n = 3). (d) Simulation of PIP2 by divalent cations. The fractions Pp (screened PIP2, red line) and Pf (free PIP2, blue line) are plotted based on parameters given in Supplementary Table 1. (e) Kinetics of the C2AB binding to liposome membranes. Averaged time courses of the rhodamine emission at different concentrations of C2AB. Liposomes were rapidly mixed with an equal volume of solutions containing different C2AB concentrations under pseudo-first order conditions. a.u., arbitrary unit.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Table 1 (PDF 1219 kb)
Rights and permissions
About this article
Cite this article
Park, Y., Seo, J., Fraind, A. et al. Synaptotagmin-1 binds to PIP2-containing membrane but not to SNAREs at physiological ionic strength. Nat Struct Mol Biol 22, 815–823 (2015). https://doi.org/10.1038/nsmb.3097
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3097
This article is cited by
-
Mechanisms of SNARE proteins in membrane fusion
Nature Reviews Molecular Cell Biology (2024)
-
Vesicle trafficking and vesicle fusion: mechanisms, biological functions, and their implications for potential disease therapy
Molecular Biomedicine (2022)
-
Electrostatic regulation of the cis- and trans-membrane interactions of synaptotagmin-1
Scientific Reports (2022)
-
Molecular landscape of BoNT/B bound to a membrane-inserted synaptotagmin/ganglioside complex
Cellular and Molecular Life Sciences (2022)
-
Conserved arginine residues in synaptotagmin 1 regulate fusion pore expansion through membrane contact
Nature Communications (2021)