Abstract
The AAA+ ATPase Vps4 disassembles ESCRT-III and is essential for HIV-1 budding and other pathways. Vps4 is a paradigmatic member of a class of hexameric AAA+ ATPases that disassemble protein complexes without degradation. To distinguish between local displacement versus global unfolding mechanisms for complex disassembly, we carried out hydrogen/deuterium exchange during Saccharomyces cerevisiae Vps4 disassembly of a chimeric Vps24-2 ESCRT-III filament. EX1 exchange behavior shows that Vps4 completely unfolds ESCRT-III substrates on a time scale consistent with the disassembly reaction. The established unfoldase ClpX showed the same pattern, thus demonstrating a common unfolding mechanism. Vps4 hexamers containing a single cysteine residue in the pore loops were cross-linked to ESCRT-III subunits containing unique cysteines within the folded core domain. These data support a mechanism in which Vps4 disassembles its substrates by completely unfolding them and threading them through the central pore.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Votteler, J. & Sundquist, W.I. Virus budding and the ESCRT pathway. Cell Host Microbe 14, 232–241 (2013).
Hanson, P.I. & Cashikar, A. Multivesicular body morphogenesis. Annu. Rev. Cell Dev. Biol. 28, 337–362 (2012).
Guizetti, J. et al. Cortical constriction during abscission involves helices of ESCRT-III-dependent filaments. Science 331, 1616–1620 (2011).
Agromayor, M. & Martin-Serrano, J. Knowing when to cut and run: mechanisms that control cytokinetic abscission. Trends Cell Biol. 23, 433–441 (2013).
Choudhuri, K. et al. Polarized release of T-cell-receptor-enriched microvesicles at the immunological synapse. Nature 507, 118–123 (2014).
Jimenez, A.J. et al. ESCRT machinery is required for plasma membrane repair. Science 343, 1247136 (2014).
Shields, S.B. & Piper, R.C. How ubiquitin functions with ESCRTs. Traffic 12, 1306–1317 (2011).
Wollert, T., Wunder, C., Lippincott-Schwartz, J. & Hurley, J.H. Membrane scission by the ESCRT-III complex. Nature 458, 172–177 (2009).
Lata, S. et al. Helical structures of ESCRT-III are disassembled by VPS4. Science 321, 1354–1357 (2008).
Hill, C.P. & Babst, M. Structure and function of the membrane deformation AAA ATPase Vps4. Biochim. Biophys. Acta 1823, 172–181 (2012).
Babst, M., Wendland, B., Estepa, E.J. & Emr, S.D. The Vps4p AAA ATPase regulates membrane association of a Vps protein complex required for normal endosome function. EMBO J. 17, 2982–2993 (1998).
Muzioł, T. et al. Structural basis for budding by the ESCRT-III factor CHMP3. Dev. Cell 10, 821–830 (2006).
Bajorek, M. et al. Structural basis for ESCRT-III protein autoinhibition. Nat. Struct. Mol. Biol. 16, 754–762 (2009).
Xiao, J. et al. Structural basis of Ist1 function and Ist1-Did2 interaction in the multivesicular body pathway and cytokinesis. Mol. Biol. Cell 20, 3514–3524 (2009).
Zamborlini, A. et al. Release of autoinhibition converts ESCRT-III components into potent inhibitors of HIV-1 budding. Proc. Natl. Acad. Sci. USA 103, 19140–19145 (2006).
Shim, S., Kimpler, L.A. & Hanson, P.I. Structure/function analysis of four core ESCRT-III proteins reveals common regulatory role for extreme C-terminal domain. Traffic 8, 1068–1079 (2007).
Lata, S. et al. Structural basis for autoinhibition of ESCRT-III CHMP3. J. Mol. Biol. 378, 818–827 (2008).
Ghazi-Tabatabai, S. et al. Structure and disassembly of filaments formed by the ESCRT-III subunit Vps24. Structure 16, 1345–1356 (2008).
Hanson, P.I., Roth, R., Lin, Y. & Heuser, J.E. Plasma membrane deformation by circular arrays of ESCRT-III protein filaments. J. Cell Biol. 180, 389–402 (2008).
Henne, W.M., Buchkovich, N.J., Zhao, Y. & Emr, S.D. The endosomal sorting complex ESCRT-II mediates the assembly and architecture of ESCRT-III helices. Cell 151, 356–371 (2012).
Cashikar, A.G. et al. Structure of cellular ESCRT-III spirals and their relationship to HIV budding. eLife 3, e02184 (2014).
Shen, Q.T. et al. Structural analysis and modeling reveals new mechanisms governing ESCRT-III spiral filament assembly. J. Cell Biol. 206, 763–777 (2014).
Obita, T. et al. Structural basis for selective recognition of ESCRT-III by the AAA ATPase Vps4. Nature 449, 735–739 (2007).
Stuchell-Brereton, M.D. et al. ESCRT-III recognition by VPS4 ATPases. Nature 449, 740–744 (2007).
Hurley, J.H. & Yang, D. MIT domainia. Dev. Cell 14, 6–8 (2008).
Kieffer, C. et al. Two distinct modes of ESCRT-III recognition are required for VPS4 functions in lysosomal protein targeting and HIV-1 budding. Dev. Cell 15, 62–73 (2008).
Monroe, N. et al. The oligomeric state of the active Vps4 AAA ATPase. J. Mol. Biol. 426, 510–525 (2014).
Scott, A. et al. Structural and mechanistic studies of VPS4 proteins. EMBO J. 24, 3658–3669 (2005).
Xiao, J., Xia, H., Yoshino-Koh, K., Zhou, J. & Xu, Z. Structural characterization of the ATPase reaction cycle of endosomal AAA protein Vps4. J. Mol. Biol. 374, 655–670 (2007).
Hartmann, C. et al. Vacuolar protein sorting: two different functional states of the AAA-ATPase Vps4p. J. Mol. Biol. 377, 352–363 (2008).
Gonciarz, M.D. et al. Biochemical and structural studies of yeast Vps4 oligomerization. J. Mol. Biol. 384, 878–895 (2008).
Yu, Z., Gonciarz, M.D., Sundquist, W.I., Hill, C.P. & Jensen, G.J. Cryo-EM structure of dodecameric Vps4p and its 2:1 complex with Vta1p. J. Mol. Biol. 377, 364–377 (2008).
Landsberg, M.J., Vajjhala, P.R., Rothnagel, R., Munn, A.L. & Hankamer, B. Three-dimensional structure of AAA ATPase Vps4: advancing structural insights into the mechanisms of endosomal sorting and enveloped virus budding. Structure 17, 427–437 (2009).
Zhang, X. et al. Structure of the AAA ATPase p97. Mol. Cell 6, 1473–1484 (2000).
Davies, B.A. et al. Coordination of substrate binding and ATP hydrolysis in Vps4-mediated ESCRT-III disassembly. Mol. Biol. Cell 21, 3396–3408 (2010).
Weber-Ban, E.U., Reid, B.G., Miranker, A.D. & Horwich, A.L. Global unfolding of a substrate protein by the Hsp100 chaperone ClpA. Nature 401, 90–93 (1999).
Baker, T.A. & Sauer, R.T. ClpXP, an ATP-powered unfolding and protein-degradation machine. Biochim. Biophys. Acta 1823, 15–28 (2012).
Abdelhakim, A.H., Sauer, R.T. & Baker, T.A. The AAA plus ClpX machine unfolds a keystone subunit to remodel the Mu transpososome. Proc. Natl. Acad. Sci. USA 107, 2437–2442 (2010).
Zhao, M. et al. Mechanistic insights into the recycling machine of the SNARE complex. Nature 518, 61–67 (2015).
Wales, T.E. & Engen, J.R. Hydrogen exchange mass spectrometry for the analysis of protein dynamics. Mass Spectrom. Rev. 25, 158–170 (2006).
Weis, D.D., Wales, T.E., Engen, J.R., Hotchko, M. & Ten Eyck, L.F. Identification and characterization of EX1 kinetics in H/D exchange mass spectrometry by peak width analysis. J. Am. Soc. Mass Spectrom. 17, 1498–1509 (2006).
Martin, A., Baker, T.A. & Sauer, R.T. Pore loops of the AAA plus ClpX machine grip substrates to drive translocation and unfolding. Nat. Struct. Mol. Biol. 15, 1147–1151 (2008).
Martin, A., Baker, T.A. & Sauer, R.T. Diverse pore loops of the AAA plus ClpX machine mediate unassisted and adaptor-dependent recognition of ssrA-tagged substrates. Mol. Cell 29, 441–450 (2008).
Barkow, S.R., Levchenko, I., Baker, T.A. & Sauer, R.T. Polypeptide translocation by the AAA+ ClpXP protease machine. Chem. Biol. 16, 605–612 (2009).
Glynn, S.E., Martin, A., Nager, A.R., Baker, T.A. & Sauer, R.T. Structures of asymmetric ClpX hexamers reveal nucleotide-dependent motions in a AAA+ protein-unfolding machine. Cell 139, 744–756 (2009).
Maillard, R.A. et al. ClpX(P) generates mechanical force to unfold and translocate its protein substrates. Cell 145, 459–469 (2011).
Gottesman, S., Roche, E., Zhou, Y.N. & Sauer, R.T. The ClpXP and ClpAP proteases degrade proteins with carboxy-terminal peptide tails added by the SsrA-tagging system. Genes Dev. 12, 1338–1347 (1998).
Spurlino, J.C., Lu, G.Y. & Quiocho, F.A. Refined 1.8-Å structure reveals the mode of binding of beta-cyclodextrin to the maltodextrin binding protein. Biochemistry 32, 10553–10559 (1993).
Azmi, I.F. et al. ESCRT-III family members stimulate Vps4 ATPase activity directly or via Vta1. Dev. Cell 14, 50–61 (2008).
Babst, M., Sato, T.K., Banta, L.M. & Emr, S.D. Endosomal transport function in yeast requires a novel AAA-type ATPase, Vps4p. EMBO J. 16, 1820–1831 (1997).
Frickey, T. & Lupas, A.N. Phylogenetic analysis of AAA proteins. J. Struct. Biol. 146, 2–10 (2004).
Roll-Mecak, A. & Vale, R.D. Structural basis of microtubule severing by the hereditary spastic paraplegia protein spastin. Nature 451, 363–367 (2008).
Sharma, S., Hoskins, J.R. & Wickner, S. Binding and degradation of heterodimeric substrates by ClpAP and ClpXP. J. Biol. Chem. 280, 5449–5455 (2005).
Acknowledgements
We thank K. Nyquist (University of California, Berkeley) for samples of Escherichia coli ClpX and ClpP. This work was supported by grants R01AI112442 (J.H.H.) and R01GM094497 (A.M.) from the US National Institutes of Health.
Author information
Authors and Affiliations
Contributions
B.Y. conceived the project, created reagents, acquired data, analyzed data and wrote the manuscript; G.S. analyzed data; Q.S. acquired data; A.M. conceived the project and wrote the manuscript; J.H.H. conceived the project, analyzed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Vps24-2 filaments.
(a) Electron micrograph of negatively stained filaments of Vps24-2. Scale bar = 50 nm. (b) Sedimentation of Vps24-2 and MBP-Vps24-2-ssrA. Vps24-2 and MBP-Vps24-2-ssrA were concentrated to above 200 μM and incubated at 4 oC overnight before dilution to 5 μM and ultracentrifugation. The resulting pellet (P) and supernatant (S) fractions were analyzed by SDS-PAGE.
Supplementary Figure 2 Peptide coverage of Vps24-2 and MBP–Vps24-2–ssrA.
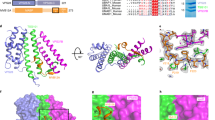
Sequence coverage map for (a) Vps24-2 and (b) MBP-Vps24-2-ssrA. Solid green lines above the protein sequence denote the pepsin digest fragments identified in the study. The two residues mutated to cysteine for crosslinking experiments are labeled with asterisks.
Supplementary Figure 3 Filamentous Vps24-2 has a stable core.
(a) Total deuteron incorporation into undigested Vps24-2 monomers (triangles) and filaments (squares) over time. (b) Deuteron incorporation over time for the Vps24-2 part of MBP-Vps24-2-ssrA, mapped onto the Vps24-2 structural model. (c) and (d) Deuteron incorporation at 5 sec, 20 sec, 2min and 5 min for Vps24-2 monomers (c) and Vps24-2 filaments (d), mapped onto the Vps24-2 structural model. The color-coding for different percentages of deuteron incorporation is the same as in Fig. 2b.
Supplementary Figure 4 Vps4-dependent EX1 exchange in helices α1, α2 and α3.
(a), (c) and (e) Mass spectra of the indicated peptides from helices α1, α2, and α3 of Vps24-2 monomers (left), filaments (middle) or filaments plus Vps4 (right). Controls and time points are indicated. Arrows above the spectra indicate regions of the spectrum representing EX2 and EX1 behavior. (b), (d) and (f) Peak-width analysis of the selected peptides at 5, 10, 20, 30, 40 and 60 sec. Open circles, Vps24-2 monomers; filled squares, Vps24-2 filaments; filled triangles, Vps24-2 filaments plus Vps4. The grey bar denotes the 2 Da peak-width change allowance for peptides undergoing EX2 kinetics 41. Each peptide is color coded as in Fig. 2a.
Supplementary Figure 5 ClpX- and Vps4-dependent EX1 exchange in MBP.
(a-d) Mass spectra of four additional peptides from the MBP portion of MBP-Vps24-2-ssrA prior to incubation in D2O (top row), alone (second row), and in the presence of ClpX (third row) or Vps4 (fourth row), or after unfolding in 6 M GdnHCl followed by complete deuteration (bottom row). Each peptide is color coded as in Fig. 4a.
Supplementary Figure 6 ATPase activity of Vps4 constructs.
(a) SDS-PAGE gel of Vps4CF-mediated disassembly of Vps24-2 filament. (b) Bar graph showing the ATPase activity of Vps4 constructs in the absence or presence of substrate Vps24-2. (c) Structural model of Vps24 with position of I92C and M136C mutation highlighted in yellow.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 (PDF 6427 kb)
Supplementary Data Set 1
Uncropped gels and western blot (PDF 35772 kb)
Rights and permissions
About this article
Cite this article
Yang, B., Stjepanovic, G., Shen, Q. et al. Vps4 disassembles an ESCRT-III filament by global unfolding and processive translocation. Nat Struct Mol Biol 22, 492–498 (2015). https://doi.org/10.1038/nsmb.3015
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3015
This article is cited by
-
The peroxisomal AAA-ATPase Pex1/Pex6 unfolds substrates by processive threading
Nature Communications (2018)
-
HDX-MS reveals dysregulated checkpoints that compromise discrimination against self RNA during RIG-I mediated autoimmunity
Nature Communications (2018)
-
Katanin spiral and ring structures shed light on power stroke for microtubule severing
Nature Structural & Molecular Biology (2017)
-
Dynamic subunit turnover in ESCRT-III assemblies is regulated by Vps4 to mediate membrane remodelling during cytokinesis
Nature Cell Biology (2017)
-
HDX reveals the conformational dynamics of DNA sequence specific VDR co-activator interactions
Nature Communications (2017)