Abstract
We report crystal structures of the Thermus thermophilus ribosome at 2.3- to 2.5-Å resolution, which have enabled modeling of rRNA modifications. The structures reveal contacts of modified nucleotides with mRNA and tRNAs or protein pY, and contacts within the ribosome interior stabilizing the functional fold of rRNA. Our work provides a resource to explore the roles of rRNA modifications and yields a more comprehensive atomic model of a bacterial ribosome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Decatur, W.A. & Fournier, M.J. Trends Biochem. Sci. 27, 344–351 (2002).
Chow, C.S., Lamichhane, T.N. & Mahto, S.K. ACS Chem. Biol. 2, 610–619 (2007).
Sergiev, P.V. et al. in Ribosomes: Structure, Function, and Dynamics (eds. Rodnina, M.V. et al.) 97–110 (Springer, 2011).
Mengel-Jørgensen, J. et al. J. Biol. Chem. 281, 22108–22117 (2006).
Demirci, H. et al. RNA 16, 2319–2324 (2010).
Desaulniers, J.P. et al. Org. Biomol. Chem. 6, 3892–3895 (2008).
Sumita, M., Jiang, J., SantaLucia, J. Jr. & Chow, C.S. Biopolymers 97, 94–106 (2012).
Liang, X.H., Liu, Q. & Fournier, M.J. Mol. Cell 28, 965–977 (2007).
Baudin-Baillieu, A. et al. Nucleic Acids Res. 37, 7665–7677 (2009).
Baxter-Roshek, J.L., Petrov, A.N. & Dinman, J.D. PLoS ONE 2, e174 (2007).
Helser, T.L., Davies, J.E. & Dahlberg, J.E. Nat. New Biol. 233, 12–14 (1971).
Doi, Y. & Arakawa, Y. Clin. Infect. Dis. 45, 88–94 (2007).
Prokhorova, I.V. et al. Sci. Rep. 3, 3236 (2013).
Prince, J.B., Taylor, B.H., Thurlow, D.L., Ofengand, J. & Zimmermann, R.A. Proc. Natl. Acad. Sci. USA 79, 5450–5454 (1982).
Kimura, S. & Suzuki, T. Nucleic Acids Res. 38, 1341–1352 (2010).
Burakovsky, D.E. et al. Nucleic Acids Res. 40, 7885–7895 (2012).
Jemiolo, D.K., Taurence, J.S. & Giese, S. Nucleic Acids Res. 19, 4259–4265 (1991).
Arora, S. et al. Nucleic Acids Res. 41, 4963–4975 (2013).
Piekna-Przybylska, D., Decatur, W.A. & Fournier, M.J. Nucleic Acids Res. 36, D178–D183 (2008).
Polikanov, Y.S., Steitz, T.A. & Innis, C.A. Nat. Struct. Mol. Biol. 21, 787–793 (2014).
Polikanov, Y.S., Blaha, G.M. & Steitz, T.A. Science 336, 915–918 (2012).
Kabsch, W. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P.D. et al. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Acknowledgements
We thank J. Lin, C.A. Innis and N.V. Sumbatyan for bringing this project to our attention; R.L. Grodzicki for preparation of the unmodified tRNAs and for critical reading of the manuscript; and members of the T.A.S. and D.S. laboratories for discussions. We also thank the staff at the Advanced Photon Source (beamline 24ID), supported by award GM103403 from the US National Center for Research Resources at the US National Institutes of Health, the staff at the US National Synchrotron Light Source (beamline X25) for help during data collection and the staff at the Richards Center at Yale University for computational support. This work was supported by US National Institutes of Health grants GM022778 (T.A.S.) and GM022854 (D.S.).
Author information
Authors and Affiliations
Contributions
Y.S.P. and S.V.M. devised experiments; Y.S.P. crystallized ribosomal complexes and performed crystallographic data collection and processing, S.V.M. and Y.S.P. analyzed the data, and S.V.M., Y.S.P., D.S. and T.A.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Post-translational modification of ribosomal protein S12 and prosthetic group of protein S4.
(a, b) β-methyl-thiolation of Asp88 (ms-Asp88) in ribosomal protein S12 is a bacteria-specific and essential modification with unknown role in protein synthesis (Kowalak, J.A. & Walsh, K.A. Beta-methylthio-aspartic acid: identification of a novel posttranslational modification in ribosomal protein S12 from Escherichia coli. Protein Sci. 5, 1625-1632, 1996). We show that this post-translational modification establishes an additional RNA-protein contact, mediated by the N7-methyl group of the modified nucleotide m7G527. This interaction, observed in a close proximity to the decoding center, suggests that the essential function of msD88 might be related to the assembly of the decoding center and that the effect of msD88 on the ribosome structure may depends on methylation of m7G527. (a) The 30S subunit is shown in light yellow with protein S12 highlighted in light blue. (b) Close-up view of panel (a). (c, d) The iron-sulfur cluster coordinated by protein S4 has been recently identified in a crystallographic study of T. thermophilus ribosomes (for details see Online Methods section in (Polikanov, Y.S., Steitz, T.A. & Innis, C.A. A proton wire to couple aminoacyl-tRNA accommodation and peptide-bond formation on the ribosome. Nat. Struct. Mol. Biol. 21, 787-793, 2014)). It is located in the vicinity of the mRNA entrance tunnel on the cytoplasmic side of the small ribosomal subunit. The covalent association of the 4Fe-4S cluster with protein S4 is mediated by four cysteine residues (Cys9, Cys12, Cys26, and Cys31) in the N-terminal globular domain. Presence of the iron-sulfur cluster within T. thermophilus ribosomes is consistent with metabolism of these bacteria that require high concentration of iron for optimal growth. It also explains why samples of pure ribosome have a yellow/red color. Importantly, it has been reported previously that protein S4 can associate with RNA polymerase in vivo (Torres, M., Condon, C., Balada, J.M., Squires, C. & Squires, C.L. Ribosomal protein S4 is a transcription factor with properties remarkably similar to NusA, a protein involved in both non-ribosomal and ribosomal RNA antitermination. Embo J. 20, 3811-3820, 2001). Although the function of this prosthetic group has not been examined, it is appealing to propose that protein S4 can mediate the ribosome interaction with RNA-polymerase during the coupled transcription-translation, where iron-sulfur cluster stabilizes the complex at high temperatures. Location of protein S4 on the 30S subunit is viewed from the cytoplasm with the 50S subunit removed, as indicated by the inset. (c) The 30S subunit is shown in light yellow with protein S4 highlighted in green. The mRNA is shown in blue, and arrows are pointing to its entrance and exit channels. (d) Close-up view of panel (c).
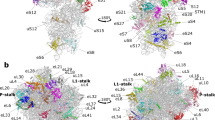
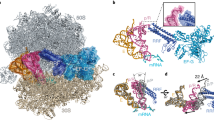
Supplementary Figure 2 Modified nucleotides form molecular contacts between the ribosomal subunits and within the ribosome interior.
(a) 2’-O-methyl groups of m5U1915 and Cm1920 of the 23S rRNA at the interface between the ribosomal subunits. Being specific to T. thermophilus, these modifications might adjust the strength of inter-subunit interactions at high temperatures. (b) m2A2503 methylation extends the stacking surface between the nucleotides A2059 and A2503 that maintains the fold of the two single-stranded rRNA segments forming the wall of the peptide exit tunnel. (c) The 2’-O-methyl group of the Um2552 of the 23S rRNA introduces hydrophobic contacts in the A-loop with the key residue G2553 that is involved in accommodation of the aminoacyl-tRNA. (d) Similarly to the A-loop, the 2’-O-methyl group of Gm2251 fills the cavity between the sugar of C2065 and the base of U2449 in the P-loop of the 23S rRNA.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2 and Supplementary Tables 1 and 2 (PDF 1469 kb)
Rights and permissions
About this article
Cite this article
Polikanov, Y., Melnikov, S., Söll, D. et al. Structural insights into the role of rRNA modifications in protein synthesis and ribosome assembly. Nat Struct Mol Biol 22, 342–344 (2015). https://doi.org/10.1038/nsmb.2992
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2992
This article is cited by
-
RNA modification-mediated mRNA translation regulation in liver cancer: mechanisms and clinical perspectives
Nature Reviews Gastroenterology & Hepatology (2024)
-
RIP-PEN-seq identifies a class of kink-turn RNAs as splicing regulators
Nature Biotechnology (2024)
-
Structural basis of Cfr-mediated antimicrobial resistance and mechanisms to evade it
Nature Chemical Biology (2024)
-
C2-methyladenosine in tRNA promotes protein translation by facilitating the decoding of tandem m2A-tRNA-dependent codons
Nature Communications (2024)
-
Structural insights into the mechanism of overcoming Erm-mediated resistance by macrolides acting together with hygromycin-A
Nature Communications (2023)