Abstract
Ubiquitination is a post-translational modification that signals multiple processes, including protein degradation, trafficking and DNA repair. Polyubiquitin accumulates globally during the oxidative stress response, and this has been mainly attributed to increased ubiquitin conjugation and perturbations in protein degradation. Here we show that the unconventional Lys63 (K63)-linked polyubiquitin accumulates in the yeast Saccharomyces cerevisiae in a highly sensitive and regulated manner as a result of exposure to peroxides. We demonstrate that hydrogen peroxide inhibits the deubiquitinating enzyme Ubp2, leading to accumulation of K63 conjugates assembled by the Rad6 ubiquitin conjugase and the Bre1 ubiquitin ligase. Using linkage-specific isolation methods and stable isotope labeling by amino acids in cell culture (SILAC)-based quantitative proteomics, we identified >100 new K63-polyubiquitinated targets, which were substantially enriched in ribosomal proteins. Finally, we demonstrate that impairment of K63 ubiquitination during oxidative stress affects polysome stability and protein expression, rendering cells more sensitive to stress, and thereby reveal a new redox-regulatory role for this modification.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Herrero, E., Ros, J., Belli, G. & Cabiscol, E. Redox control and oxidative stress in yeast cells. Biochim. Biophys. Acta 1780, 1217–1235 (2008).
Apel, K. & Hirt, H. Reactive oxygen species: metabolism, oxidative stress, and signal transduction. Annu. Rev. Plant Biol. 55, 373–399 (2004).
Klaunig, J.E. & Kamendulis, L.M. The role of oxidative stress in carcinogenesis. Annu. Rev. Pharmacol. Toxicol. 44, 239–267 (2004).
Dröge, W. Free radicals in the physiological control of cell function. Physiol. Rev. 82, 47–95 (2002).
Finkel, T. & Holbrook, N.J. Oxidants, oxidative stress and the biology of ageing. Nature 408, 239–247 (2000).
Morano, K.A., Grant, C.M. & Moye-Rowley, W.S. The response to heat shock and oxidative stress in Saccharomyces cerevisiae. Genetics 190, 1157–1195 (2012).
Goldberg, A.L. Protein degradation and protection against misfolded or damaged proteins. Nature 426, 895–899 (2003).
Glickman, M.H. & Ciechanover, A. The ubiquitin-proteasome proteolytic pathway: destruction for the sake of construction. Physiol. Rev. 82, 373–428 (2002).
Reyes-Turcu, F.E., Ventii, K.H. & Wilkinson, K.D. Regulation and cellular roles of ubiquitin-specific deubiquitinating enzymes. Annu. Rev. Biochem. 78, 363–397 (2009).
Komander, D., Clague, M.J. & Urbe, S. Breaking the chains: structure and function of the deubiquitinases. Nat. Rev. Mol. Cell Biol. 10, 550–563 (2009).
Finley, D., Ulrich, H.D., Sommer, T. & Kaiser, P. The ubiquitin-proteasome system of Saccharomyces cerevisiae. Genetics 192, 319–360 (2012).
Hershko, A., Ciechanover, A., Heller, H., Haas, A.L. & Rose, I.A. Proposed role of ATP in protein breakdown: conjugation of protein with multiple chains of the polypeptide of ATP-dependent proteolysis. Proc. Natl. Acad. Sci. USA 77, 1783–1786 (1980).
Spence, J., Sadis, S., Haas, A.L. & Finley, D. A ubiquitin mutant with specific defects in DNA repair and multiubiquitination. Mol. Cell. Biol. 15, 1265–1273 (1995).
Komander, D. The emerging complexity of protein ubiquitination. Biochem. Soc. Trans. 37, 937–953 (2009).
Chen, Z.J. & Sun, L.J. Nonproteolytic functions of ubiquitin in cell signaling. Mol. Cell 33, 275–286 (2009).
Xu, P. et al. Quantitative proteomics reveals the function of unconventional ubiquitin chains in proteasomal degradation. Cell 137, 133–145 (2009).
Matsumoto, M.L. et al. K11-linked polyubiquitination in cell cycle control revealed by a K11 linkage-specific antibody. Mol. Cell 39, 477–484 (2010).
Lauwers, E., Jacob, C. & Andre, B. K63-linked ubiquitin chains as a specific signal for protein sorting into the multivesicular body pathway. J. Cell Biol. 185, 493–502 (2009).
MacDonald, C., Buchkovich, N.J., Stringer, D.K., Emr, S.D. & Piper, R.C. Cargo ubiquitination is essential for multivesicular body intralumenal vesicle formation. EMBO Rep. 13, 331–338 (2012).
Hoege, C., Pfander, B., Moldovan, G.L., Pyrowolakis, G. & Jentsch, S. RAD6-dependent DNA repair is linked to modification of PCNA by ubiquitin and SUMO. Nature 419, 135–141 (2002).
Deng, L. et al. Activation of the IκB kinase complex by TRAF6 requires a dimeric ubiquitin-conjugating enzyme complex and a unique polyubiquitin chain. Cell 103, 351–361 (2000).
Zhou, H. et al. Bcl10 activates the NF-κB pathway through ubiquitination of NEMO. Nature 427, 167–171 (2004).
Dudek, E.J. et al. Selectivity of the ubiquitin pathway for oxidatively modified proteins: relevance to protein precipitation diseases. FASEB J. 19, 1707–1709 (2005).
Medicherla, B. & Goldberg, A.L. Heat shock and oxygen radicals stimulate ubiquitin-dependent degradation mainly of newly synthesized proteins. J. Cell Biol. 182, 663–673 (2008).
Shringarpure, R., Grune, T., Mehlhase, J. & Davies, K.J. Ubiquitin conjugation is not required for the degradation of oxidized proteins by proteasome. J. Biol. Chem. 278, 311–318 (2003).
Pickering, A.M. et al. The immunoproteasome, the 20S proteasome and the PA28αβ proteasome regulator are oxidative-stress-adaptive proteolytic complexes. Biochem. J. 432, 585–594 (2010).
Peterson, A.C., Russell, J.D., Bailey, D.J., Westphall, M.S. & Coon, J.J. Parallel reaction monitoring for high resolution and high mass accuracy quantitative, targeted proteomics. Mol. Cell. Proteomics 11, 1475–1488 (2012).
Kirkpatrick, D.S., Denison, C. & Gygi, S.P. Weighing in on ubiquitin: the expanding role of mass-spectrometry-based proteomics. Nat. Cell Biol. 7, 750–757 (2005).
Lee, B.H. et al. Enhancement of proteasome activity by a small-molecule inhibitor of USP14. Nature 467, 179–184 (2010).
Giannattasio, M., Lazzaro, F., Plevani, P. & Muzi-Falconi, M. The DNA damage checkpoint response requires histone H2B ubiquitination by Rad6-Bre1 and H3 methylation by Dot1. J. Biol. Chem. 280, 9879–9886 (2005).
Fleming, A.B., Kao, C.F., Hillyer, C., Pikaart, M. & Osley, M.A. H2B ubiquitylation plays a role in nucleosome dynamics during transcription elongation. Mol. Cell 31, 57–66 (2008).
Wood, A. et al. Bre1, an E3 ubiquitin ligase required for recruitment and substrate selection of Rad6 at a promoter. Mol. Cell 11, 267–274 (2003).
Hwang, C.S., Shemorry, A., Auerbach, D. & Varshavsky, A. The N-end rule pathway is mediated by a complex of the RING-type Ubr1 and HECT-type Ufd4 ubiquitin ligases. Nat. Cell Biol. 12, 1177–1185 (2010).
Dohmen, R.J., Madura, K., Bartel, B. & Varshavsky, A. The N-end rule is mediated by the UBC2(RAD6) ubiquitin-conjugating enzyme. Proc. Natl. Acad. Sci. USA 88, 7351–7355 (1991).
Lee, J.S. et al. Histone crosstalk between H2B monoubiquitination and H3 methylation mediated by COMPASS. Cell 131, 1084–1096 (2007).
Song, Y.H. & Ahn, S.H.A. Bre1-associated protein, large 1 (Lge1), promotes H2B ubiquitylation during the early stages of transcription elongation. J. Biol. Chem. 285, 2361–2367 (2010).
Wood, A., Schneider, J., Dover, J., Johnston, M. & Shilatifard, A. The Paf1 complex is essential for histone monoubiquitination by the Rad6-Bre1 complex, which signals for histone methylation by COMPASS and Dot1p. J. Biol. Chem. 278, 34739–34742 (2003).
Piro, A.S., Mayekar, M.K., Warner, M.H., Davis, C.P. & Arndt, K.M. Small region of Rtf1 protein can substitute for complete Paf1 complex in facilitating global histone H2B ubiquitylation in yeast. Proc. Natl. Acad. Sci. USA 109, 10837–10842 (2012).
Tercero, J.A. & Diffley, J.F. Regulation of DNA replication fork progression through damaged DNA by the Mec1/Rad53 checkpoint. Nature 412, 553–557 (2001).
Spence, J. et al. Cell cycle-regulated modification of the ribosome by a variant multiubiquitin chain. Cell 102, 67–76 (2000).
Tan, J.M. et al. Lysine 63-linked ubiquitination promotes the formation and autophagic clearance of protein inclusions associated with neurodegenerative diseases. Hum. Mol. Genet. 17, 431–439 (2008).
Saeki, Y. et al. Lysine 63-linked polyubiquitin chain may serve as a targeting signal for the 26S proteasome. EMBO J. 28, 359–371 (2009).
Grant, C.M. Regulation of translation by hydrogen peroxide. Antioxid. Redox Signal. 15, 191–203 (2011).
Lee, J.G., Baek, K., Soetandyo, N. & Ye, Y. Reversible inactivation of deubiquitinases by reactive oxygen species in vitro and in cells. Nat. Commun. 4, 1568 (2013).
Cotto-Rios, X.M., Bekes, M., Chapman, J., Ueberheide, B. & Huang, T.T. Deubiquitinases as a signaling target of oxidative stress. Cell Reports 2, 1475–1484 (2012).
Kee, Y., Munoz, W., Lyon, N. & Huibregtse, J.M. The deubiquitinating enzyme Ubp2 modulates Rsp5-dependent Lys63-linked polyubiquitin conjugates in Saccharomyces cerevisiae. J. Biol. Chem. 281, 36724–36731 (2006).
Komander, D. et al. Molecular discrimination of structurally equivalent Lys 63-linked and linear polyubiquitin chains. EMBO Rep. 10, 466–473 (2009).
Netto, L.E. et al. Reactive cysteine in proteins: protein folding, antioxidant defense, redox signaling and more. Comp. Biochem. Physiol. C Toxicol. Pharmacol. 146, 180–193 (2007).
Ritorto, M.S. et al. Screening of DUB activity and specificity by MALDI-TOF mass spectrometry. Nat. Commun. 5, 4763 (2014).
Tomanov, K., Luschnig, C. & Bachmair, A. Ubiquitin Lys 63 chains - second-most abundant, but poorly understood in plants. Front. Plant Sci. 5, 15 (2014).
Peng, J. et al. A proteomics approach to understanding protein ubiquitination. Nat. Biotechnol. 21, 921–926 (2003).
Herman, P.K. Stationary phase in yeast. Curr. Opin. Microbiol. 5, 602–607 (2002).
El-Sharoud, W.M. & Niven, G.W. The influence of ribosome modulation factor on the survival of stationary-phase Escherichia coli during acid stress. Microbiology 153, 247–253 (2007).
Yoshida, H., Ueta, M., Maki, Y., Sakai, A. & Wada, A. Activities of Escherichia coli ribosomes in IF3 and RMF change to prepare 100S ribosome formation on entering the stationary growth phase. Genes Cells 14, 271–280 (2009).
Kim, W. et al. Systematic and quantitative assessment of the ubiquitin-modified proteome. Mol. Cell 44, 325–340 (2011).
Shenton, D. et al. Global translational responses to oxidative stress impact upon multiple levels of protein synthesis. J. Biol. Chem. 281, 29011–29021 (2006).
Gerashchenko, M.V., Lobanov, A.V. & Gladyshev, V.N. Genome-wide ribosome profiling reveals complex translational regulation in response to oxidative stress. Proc. Natl. Acad. Sci. USA 109, 17394–17399 (2012).
Vogel, C., Silva, G.M. & Marcotte, E.M. Protein expression regulation under oxidative stress. Mol. Cell Proteomics 10, M111.009217 (2011).
Ben-Shem, A., Jenner, L., Yusupova, G. & Yusupov, M. Crystal structure of the eukaryotic ribosome. Science 330, 1203–1209 (2010).
Liu, C., Apodaca, J., Davis, L.E. & Rao, H. Proteasome inhibition in wild-type yeast Saccharomyces cerevisiae cells. Biotechniques 42, 158,160,162 (2007).
Ramírez-Valle, F., Braunstein, S., Zavadil, J., Formenti, S.C. & Schneider, R.J. eIF4GI links nutrient sensing by mTOR to cell proliferation and inhibition of autophagy. J. Cell Biol. 181, 293–307 (2008).
Acknowledgements
We thank J.R. Cussiol (Cornell University) and M. Smolka (Cornell University) for the yeast deletion templates, support in yeast genetics and comments on the manuscript. We are grateful to G. Hsu and R. Schneider for technical support in polysome analysis, and V. Subramanian and A. Hochwagen in fluorescence-activated cell sorting analysis. We thank N. Brandt (New York University), D. Gresham (New York University), M. Hochstrasser (Yale University), M.A. Osley (University of New Mexico) and R. Ratan (Burke Medical Research Institute) for sharing yeast strains and HT22 cells. We thank J.R. Chapman for assistance with targeted MS, and the Pride Team (http://www.ebi.ac.uk/services/teams/pride) for assistance with MS data deposition. We are indebted to D. Gresham, J. Davis, E. Miraldi and T. Rock for feedback on the manuscript. This work was supported in part by US National Science Foundation EAGER grant MCB-1355462 (G.M.S. and C.V.), the Zegar Family Foundation Fund for Genomics Research at New York University (G.M.S. and C.V.) and US National Institutes of Health grant GM43601 (D.F.).
Author information
Authors and Affiliations
Contributions
G.M.S. conceived of the project. G.M.S., D.F. and C.V. designed the experiments, and G.M.S. conducted the experiments. G.M.S. and C.V. wrote the manuscript. G.M.S., D.F. and C.V. discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Characterization of K63 ubiquitination under oxidative stress.
(a,b) To demonstrate appropriate interpretation of the data, we provide western blots with multiple exposure times for both anti-K63 (a) and anti-K48 ubiquitin antibody (b) as for Figure. 1a. (c) Anti-K11 ubiquitin blot of lysate from wild-type (WT) cells upon treatment with, and subsequent recovery from, 0.6 mM H2O2. (d–f) Quantification of ubiquitin linkages by parallel reaction monitoring (PRM). Plots showing elution peaks with respective retention time for quantification of K48 and K63 ubiquitin signature peptides. SILAC analysis was performed using untreated WT cells (heavy labeled - blue line) combined with 0.6 mM H2O2-treated (d) or 2 h recovery cells (light labeled - pink line) (e). Peptide quantification was normalized using unmodified TLS and TITLE ubiquitin peptides. Mass list and peptide sequences used in PRM analysis are presented in Supplementary Table 5. (f) Elution peak of the three selected fragment ions for K48 and K63 ubiquitin peptides. (g) Histogram showing K48 ubiquitin quantification using +2 and +3 peptide charge states. Although the dynamic trends of both peptides are similar, we chose the +3 peptide for quantification as it has a smaller variance and only minor interference with other contaminant peaks. Histogram shows the mean of two different biological replicates (with two technical replicates each); error bars indicate the range of values across the replicates. (h,i) Anti-K63 ubiquitin western blot of lysate from WT cells treated with H2O2 for different amounts of time (h) or different concentrations of H2O2 (i). (j) Growth curve of WT (blue line) and K63R (pink line) cells in the presence (closed symbol) or in the absence (open symbol) of 0.6 mM H2O2. Cells were transferred to fresh medium after 45 min for recovery. (k) Histogram shows cellular viability measured by incubation of WT cells with 5 μM FUN1 fluorescent probe (Life Technologies) after treatment with H2O2. Error bars show standard deviation, n = 3 independent cellular growth. (l) Anti-K63 ubiquitin western blot of lysate from mouse neuronal HT22 cells exposed to H2O2 for different amounts of time. WT, wild-type SUB280 yeast strain. K63R, ubiquitin K63R mutant SUB413 yeast strain. GAPDH and actin, detected with anti-GAPDH and anti-actin antibodies, were used as loading control.
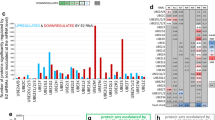
Supplementary Figure 2 New role for Rad6-Bre1 during the oxidative stress response.
(a) Scheme depicting the E2 ubiquitin conjugase Rad6 with its interacting E3 ligases (Bre1, Rad18 and Ubr1), their respective targets, and intracellular roles. (b–d) Anti-K63 and/or anti-K48 ubiquitin western blot of lysates from cells deleted for genes essential to support Rad6-Bre1-dependent monoubiquitination of histone H2B (b), yeast strain Y133 containing the histone H2B K123R mutation (c), WT cells treated with MMS (methyl methanesulfonate) or H2O2 for indicated times (d). (e) FACS analysis of asynchronously grown yeast cells (WTcol, rad6Δ and bre1Δ) untreated (blue) or treated with 0.6 mM H2O2 (red). (f–i) Anti-K63 and/or anti-K48 ubiquitin western blot of lysate from WT cells treated with 2.5 μg/ml actinomycin D (f), 100 μg/ml cycloheximide (g), 75 μM MG-132 (h) or 5 mM 3-methyladenine (3-MA) (i). (j) Anti-K63 ubiquitin western blot of cell lysate from WTcol and atg7Δ cells. Lysates were prepared from cells upon treatment with, and subsequent recovery from, 0.6 mM H2O2. GAPDH, detected with anti-GAPDH antibody, was used as loading control. WTcol, wild-type cells S288c used with the yeast deletion collection.
Supplementary Figure 3 Ubp2 regulates K63 ubiquitin reversal.
(a) Western blots anti-K63, -K48, and -K11 ubiquitin of lysate from WT cells treated with PR-619 for different amounts of time. Patterns of K63 ubiquitination induced by H2O2 and by PR-619 might differ since PR-619 is a wide range DUB inhibitor while H2O2 triggers a specific and transitory response. (b) Anti-K63 ubiquitin of lysate from WT cells treated with H2O2. Cell lysate was then incubated with (+) or without (–) 15 mM DTT for 2h at 30 °C after incubation with 20 mM IAM, 100 μM PR-619, 10 mM EDTA, or 1x EMD protease inhibitor set I. (c) Anti-K63 ubiquitin of cell lysate from yeast strains deleted for indicated DUBs from the UBP family. (d) anti-K63 ubiquitin of lysates from DUB-deleted strains treated with H2O2 for different amounts of time. (e) Plot shows activity of TAP-tagged purified Ubp2 incubated with 0.75 μM Ub-AMC fluorogenic substrate after 10 mM DTT (blue) or 0.5 mM H2O2 (red) at 30 °C for the indicated times. After measurement, 10 mM DTT was added to H2O2-treated samples (green). WTDUBs, wild-type yeast strain SUB62. GAPDH, detected with anti-GAPDH antibody, was used as loading control. a.u., fluorescence arbitrary units.
Supplementary Figure 4 K63 polyubiquitination targets and functional role.
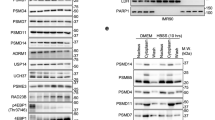
(a) Anti-K63 and anti-K48 ubiquitin western blot of K63-TUBE pull down (LifeSensors) from WT and the K63R mutant cell lysate after treatment with 0.6 mM H2O2. (b) Frequency distribution of normalized SILAC log2-ratios (WT/K63R) of K63-TUBE pull-downs from untreated (blue) and H2O2-treated (red) samples. A 5% false discovery rate was employed to select significantly changing targets (Supplementary Table 1). Further details on protein identification are provided in the Methods section and in the Supplementary Notes. (c) Frequency distribution of absolute protein abundances as log10 of molecules/cell obtained from SGD (Saccharomyces Genome Database). Graph shows abundance of all proteins quantified in the yeast proteome (white – average 3.4 ± 0.7), all proteins identified by the whole-cell LC-MS/MS analysis (gray - average 4.1 ± 0.7), and all proteins from the K63 pull down experiments (black – average 4.3 ± 0.9). Although mass spectrometry favors abundant proteins, our method was able to capture proteins within a wide dynamic range of concentrations. (d) Surface 3D structure of the large 60S and the small 40S ribosome subunit (PDB ID 4V7R)59 in light and dark gray, respectively. K63 ubiquitinated proteins identified by mass spectrometry analysis are labeled in blue. WT, wild-type SILAC GMS280 yeast strain. K63R, ubiquitin K63R mutant SILAC GMS413 yeast strain.
Supplementary Figure 5 Loading controls and K48-K63 antibody cross-reactivity.
(a) Sucrose sedimentation profiles and Ponceau-S staining for loading control of membranes used in the experiment shown in Figure 5c and 5d. (b) Ponceau-S staining for loading control of membranes used in the experiment shown in Figure 5e. * indicates that half the sample volume was loaded for better visualization. (c) Anti-K48 ubiquitin western blot of polysome profile fraction from WT and K63R mutant cells after H2O2 treatment. WT, wild-type SUB280 yeast strain. K63R, ubiquitin K63R mutant SUB413 yeast strain. WTcol, wild-type cells S288c used with the yeast deletion collection.
Supplementary Figure 6 Protein oxidation and protein abundance changes in response to oxidative stress.
(a) anti-dinitrophenyl (DNP) (oxidation) western blot of lysates from WT and K63R mutant cells upon treatment with, and subsequent recovery from, 0.6 mM H2O2. (b) Frequency distribution of the log2 SILAC K63R/WT ratio of the total cell lysate from two biological replicates analyzed by mass spectrometry. GAPDH, detected with anti-GAPDH antibody, was used as loading control. WT, wild-type SUB280 yeast strain. K63R, ubiquitin K63R mutant SUB413 yeast strain.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Note (PDF 3884 kb)
K63 ubiquitinated targets
Description of the K63 targets identified by high-resolution mass spectrometry (XLS 79 kb)
K63 ubiquitinated core targets
Description of the high-confidence K63 targets identified by two different search engines (XLS 45 kb)
Whole cell lysate protein abundance
Description of protein abundance changes in the cellular lysate measured by high-resolution mass spectrometry (XLS 365 kb)
Yeast strains
Table contains the genotypes and reference sources for the yeast Saccharomyces cerevisiae strains used in the study (XLS 39 kb)
Ubiquitin signature peptides for PRM analysis
List of the signature peptides used in the parallel reaction monitoring analysis to quantify linkage specific polyubiquitination (XLS 38 kb)
Supplementary Data Set 1
Original gels and blots - Uncropped images of gels and blots used in the main figures of this study (PDF 510 kb)
Rights and permissions
About this article
Cite this article
Silva, G., Finley, D. & Vogel, C. K63 polyubiquitination is a new modulator of the oxidative stress response. Nat Struct Mol Biol 22, 116–123 (2015). https://doi.org/10.1038/nsmb.2955
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2955
This article is cited by
-
Post-ischemic ubiquitination at the postsynaptic density reversibly influences the activity of ischemia-relevant kinases
Communications Biology (2024)
-
Molecular basis for recognition and deubiquitination of 40S ribosomes by Otu2
Nature Communications (2023)
-
Current methodologies in protein ubiquitination characterization: from ubiquitinated protein to ubiquitin chain architecture
Cell & Bioscience (2022)
-
Transcriptome analysis of Auricularia fibrillifera fruit-body responses to drought stress and rehydration
BMC Genomics (2022)
-
Rho family GTPase 1 (RND1), a novel regulator of p53, enhances ferroptosis in glioblastoma
Cell & Bioscience (2022)