Abstract
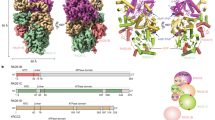
Ctp1 (also known as CtIP or Sae2) collaborates with Mre11–Rad50–Nbs1 to initiate repair of DNA double-strand breaks (DSBs), but its functions remain enigmatic. We report that tetrameric Schizosaccharomyces pombe Ctp1 contains multivalent DNA-binding and DNA-bridging activities. Through structural and biophysical analyses of the Ctp1 tetramer, we define the salient features of Ctp1 architecture: an N-terminal interlocking tetrameric helical dimer-of-dimers (THDD) domain and a central intrinsically disordered region (IDR) linked to C-terminal 'RHR' DNA-interaction motifs. The THDD, IDR and RHR are required for Ctp1 DNA-bridging activity in vitro, and both the THDD and RHR are required for efficient DSB repair in S. pombe. Our results establish non-nucleolytic roles of Ctp1 in binding and coordination of DSB-repair intermediates and suggest that ablation of human CtIP DNA binding by truncating mutations underlie the CtIP-linked Seckel and Jawad syndromes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Williams, R.S., Williams, J.S. & Tainer, J.A. Mre11-Rad50-Nbs1 is a keystone complex connecting DNA repair machinery, double-strand break signaling, and the chromatin template. Biochem. Cell Biol. 85, 509–520 (2007).
Pommier, Y. Drugging topoisomerases: lessons and challenges. ACS Chem. Biol. 8, 82–95 (2013).
You, Z. et al. CtIP links DNA double-strand break sensing to resection. Mol. Cell 36, 954–969 (2009).
Williams, R.S. et al. Nbs1 flexibly tethers Ctp1 and Mre11-Rad50 to coordinate DNA double-strand break processing and repair. Cell 139, 87–99 (2009).
Limbo, O. et al. Ctp1 is a cell-cycle-regulated protein that functions with Mre11 complex to control double-strand break repair by homologous recombination. Mol. Cell 28, 134–146 (2007).
Huertas, P., Cortes-Ledesma, F., Sartori, A.A., Aguilera, A. & Jackson, S.P. CDK targets Sae2 to control DNA-end resection and homologous recombination. Nature 455, 689–692 (2008).
Sartori, A.A. et al. Human CtIP promotes DNA end resection. Nature 450, 509–514 (2007).
Huertas, P. & Jackson, S.P. Human CtIP mediates cell cycle control of DNA end resection and double strand break repair. J. Biol. Chem. 284, 9558–9565 (2009).
Stracker, T.H. & Petrini, J.H. The MRE11 complex: starting from the ends. Nat. Rev. Mol. Cell Biol. 12, 90–103 (2011).
Chen, P.L. et al. Inactivation of CtIP leads to early embryonic lethality mediated by G1 restraint and to tumorigenesis by haploid insufficiency. Mol. Cell. Biol. 25, 3535–3542 (2005).
Qvist, P. et al. CtIP mutations cause Seckel and Jawad syndromes. PLoS Genet. 7, e1002310 (2011).
Zhou, Y., Caron, P., Legube, G. & Paull, T.T. Quantitation of DNA double-strand break resection intermediates in human cells. Nucleic Acids Res. 42, e19 (2014).
Westmoreland, J.W. & Resnick, M.A. Coincident resection at both ends of random, gamma-induced double-strand breaks requires MRX (MRN), Sae2 (Ctp1), and Mre11-nuclease. PLoS Genet. 9, e1003420 (2013).
Langerak, P., Mejia-Ramirez, E., Limbo, O. & Russell, P. Release of Ku and MRN from DNA ends by Mre11 nuclease activity and Ctp1 is required for homologous recombination repair of double-strand breaks. PLoS Genet. 7, e1002271 (2011).
Mimitou, E.P. & Symington, L.S. Sae2, Exo1 and Sgs1 collaborate in DNA double-strand break processing. Nature 455, 770–774 (2008).
Neale, M.J., Pan, J. & Keeney, S. Endonucleolytic processing of covalent protein-linked DNA double-strand breaks. Nature 436, 1053–1057 (2005).
Farah, J.A., Cromie, G.A. & Smith, G.R. Ctp1 and Exonuclease 1, alternative nucleases regulated by the MRN complex, are required for efficient meiotic recombination. Proc. Natl. Acad. Sci. USA 106, 9356–9361 (2009).
Hartsuiker, E., Neale, M.J. & Carr, A.M. Distinct requirements for the Rad32(Mre11) nuclease and Ctp1(CtIP) in the removal of covalently bound topoisomerase I and II from DNA. Mol. Cell 33, 117–123 (2009).
Lengsfeld, B.M., Rattray, A.J., Bhaskara, V., Ghirlando, R. & Paull, T.T. Sae2 is an endonuclease that processes hairpin DNA cooperatively with the Mre11/Rad50/Xrs2 complex. Mol. Cell 28, 638–651 (2007).
Lobachev, K.S., Gordenin, D.A. & Resnick, M.A. The Mre11 complex is required for repair of hairpin-capped double-strand breaks and prevention of chromosome rearrangements. Cell 108, 183–193 (2002).
Lloyd, J. et al. A supramodular FHA/BRCT-repeat architecture mediates Nbs1 adaptor function in response to DNA damage. Cell 139, 100–111 (2009).
Dubin, M.J. et al. Dimerization of CtIP, a BRCA1- and CtBP-interacting protein, is mediated by an N-terminal coiled-coil motif. J. Biol. Chem. 279, 26932–26938 (2004).
Dosztányi, Z., Csizmok, V., Tompa, P. & Simon, I. IUPred: web server for the prediction of intrinsically unstructured regions of proteins based on estimated energy content. Bioinformatics 21, 3433–3434 (2005).
Oates, M.E. et al. D2P2: database of disordered protein predictions. Nucleic Acids Res. 41, D508–D516 (2013).
Babu, M.M., Kriwacki, R.W. & Pappu, R.V. Structural biology: versatility from protein disorder. Science 337, 1460–1461 (2012).
Tompa, P. Intrinsically unstructured proteins. Trends Biochem. Sci. 27, 527–533 (2002).
Schiller, C.B. et al. Structure of Mre11–Nbs1 complex yields insights into ataxia-telangiectasia–like disease mutations and DNA damage signaling. Nat. Struct. Mol. Biol. 19, 693–700 (2012).
You, Z., Chahwan, C., Bailis, J., Hunter, T. & Russell, P. ATM activation and its recruitment to damaged DNA require binding to the C terminus of Nbs1. Mol. Cell. Biol. 25, 5363–5379 (2005).
Akamatsu, Y. et al. Molecular characterization of the role of the Schizosaccharomyces pombe nip1+/ctp1+ gene in DNA double-strand break repair in association with the Mre11-Rad50-Nbs1 complex. Mol. Cell. Biol. 28, 3639–3651 (2008).
Williams, R.S. et al. Mre11 dimers coordinate DNA end bridging and nuclease processing in double-strand-break repair. Cell 135, 97–109 (2008).
Hopfner, K.P. et al. The Rad50 zinc-hook is a structure joining Mre11 complexes in DNA recombination and repair. Nature 418, 562–566 (2002).
Deshpande, R.A. et al. ATP-driven Rad50 conformations regulate DNA tethering, end resection, and ATM checkpoint signaling. EMBO J. 33, 482–500 (2014).
Andres, S.N. et al. A human XRCC4-XLF complex bridges DNA. Nucleic Acids Res. 40, 1868–1878 (2012).
Hsiang, Y.H., Lihou, M.G. & Liu, L.F. Arrest of replication forks by drug-stabilized topoisomerase I-DNA cleavable complexes as a mechanism of cell killing by camptothecin. Cancer Res. 49, 5077–5082 (1989).
Pommier, Y. & Cherfils, J. Interfacial inhibition of macromolecular interactions: nature's paradigm for drug discovery. Trends Pharmacol. Sci. 26, 138–145 (2005).
Povirk, L.F. Processing of damaged DNA ends for double-strand break repair in mammalian cells. ISRN Mol. Biol. 2012, 345805 (2012).
Kim, H.S. et al. Functional interactions between Sae2 and the Mre11 complex. Genetics 178, 711–723 (2008).
Wang, H. et al. CtIP protein dimerization is critical for its recruitment to chromosomal DNA double-stranded breaks. J. Biol. Chem. 287, 21471–21480 (2012).
Cannavo, E. & Cejka, P. Sae2 promotes dsDNA endonuclease activity within Mre11–Rad50–Xrs2 to resect DNA breaks. Nature 514, 122–125 (2014).
Eid, W. et al. DNA end resection by CtIP and exonuclease 1 prevents genomic instability. EMBO Rep. 11, 962–968 (2010).
Zhang, Y. & Jasin, M. An essential role for CtIP in chromosomal translocation formation through an alternative end-joining pathway. Nat. Struct. Mol. Biol. 18, 80–84 (2011).
Moreno-Herrero, F. et al. Mesoscale conformational changes in the DNA-repair complex Rad50/Mre11/Nbs1 upon binding DNA. Nature 437, 440–443 (2005).
Lobachev, K., Vitriol, E., Stemple, J., Resnick, M.A. & Bloom, K. Chromosome fragmentation after induction of a double-strand break is an active process prevented by the RMX repair complex. Curr. Biol. 14, 2107–2112 (2004).
Chen, L., Trujillo, K., Ramos, W., Sung, P. & Tomkinson, A.E. Promotion of Dnl4-catalyzed DNA end-joining by the Rad50/Mre11/Xrs2 and Hdf1/Hdf2 complexes. Mol. Cell 8, 1105–1115 (2001).
Stols, L. et al. A new vector for high-throughput, ligation-independent cloning encoding a tobacco etch virus protease cleavage site. Protein Expr. Purif. 25, 8–15 (2002).
Sreerama, N. & Woody, R.W. A self-consistent method for the analysis of protein secondary structure from circular dichroism. Anal. Biochem. 209, 32–44 (1993).
Sreerama, N., Venyaminov, S.Y. & Woody, R.W. Estimation of the number of alpha-helical and beta-strand segments in proteins using circular dichroism spectroscopy. Protein Sci. 8, 370–380 (1999).
Whitmore, L. & Wallace, B.A. DICHROWEB, an online server for protein secondary structure analyses from circular dichroism spectroscopic data. Nucleic Acids Res. 32, W668–W673 (2004).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Thépaut, M. et al. Crystal structure of the coiled-coil dimerization motif of geminin: structural and functional insights on DNA replication regulation. J. Mol. Biol. 342, 275–287 (2004).
Winn, M.D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Terwilliger, T.C. Maximum-likelihood density modification. Acta Crystallogr. D Biol. Crystallogr. 56, 965–972 (2000).
Moreno, S., Klar, A. & Nurse, P. Molecular genetic analysis of fission yeast Schizosaccharomyces pombe. Methods Enzymol. 194, 795–823 (1991).
Tournier, S., Gachet, Y. & Hyams, J.S. Identification and preliminary characterization of p31, a new PSTAIRE-related protein in fission yeast. Yeast 13, 727–734 (1997).
Williams, J.S. et al. γH2A binds Brc1 to maintain genome integrity during S-phase. EMBO J. 29, 1136–1148 (2010).
Birren, B. & Lai, E. Pulsed Field Gel Electrophoresis: a Practical Guide (Academic Press, San Diego, 1993).
Acknowledgements
Our studies are supported by the US National Institute of Health Intramural Program, US National Institute of Environmental Health Sciences (NIEHS) grants 1Z01ES102765 (R.S.W.) and 1Z01ES021016 (M.A.R.). We thank L. Pedersen of the NIEHS Collaborative crystallography group and the Advanced Photon Source (APS) Southeast Regional Collaborative Access Team (SER-CAT) for beamline access. Use of the APS was supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences, under contract no. W-31-109-Eng-38. We thank M. Junop (University of Western Ontario) for nicked plasmid substrate, J. Williams of the NIEHS protein microcharacterization core for MS analysis, R. Dutcher (NIEHS) for help with MALS analysis and G. Mueller (NIEHS) and B. Wallace (NIEHS) for comments on the manuscript.
Author information
Authors and Affiliations
Contributions
S.N.A., R.S.W., M.A.R., J.W. and J.S.W. designed the experiments. S.N.A. and P.D.R. performed crystallization experiments. S.N.A. and R.S.W. solved and refined the Ctp1 X-ray structure. S.N.A. and R.S.W. carried out SAXS experiments. S.N.A., Y.N. and C.D.A. performed biochemical experiments. S.N.A., J.W., C.D.A. and J.S.W. performed S. pombe experiments. R.S.W. and S.N.A. wrote the manuscript with input from all authors. R.S.W. managed the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Ctp1 small-angle X-ray scattering (SAXS).
SAXS profiles of full length Ctp1 (blue), MBP-Ctp11–60 (green) and MBP-Ctp115–60 (black). Inset: Guinier plots, colored as for scattering profiles, indicating the absence of aggregated protein in SAXS samples.
Supplementary Figure 2 A hydrophobic core stabilizes the Ctp1 tetrameric helical dimer-of-dimers (THDD) domain.
(a) Electrostatic surface potential representation illustrating the Ctp1 tetramer hydrophobic core and hydrophilic exterior. (b) Helical wheel diagram of Ctp1 parallel dimeric coiled-coil region. The heptad repeat with additional salt bridges (dotted lines) between coils is shown. Amino acid positions are numbered for “a” and “d” positions of the heptad repeat.
Supplementary Figure 3 Experimental electron density of the Ctp1 THDD domain.
A model phased, σ-A weighted 2Fo-Fc electron density map (blue, contoured at 1.0σ) was calculated from the initial molecular replacement polyalanine model solution. The corresponding σ-A weighted Fo-Fc map (green, contoured at 2.0σ) shows positive difference density for unmodeled helical regions, and amino acid side-chains.
Supplementary Figure 4 Oligomerization states of Ctp1 truncations and mutants.
(a) SEC-MALS traces of differential refractive index and molar mass for Ctp161–294 (blue) and Ctp1 THDD mutation (R32A K41A) (red). (b) SEC-MALS traces of differential refractive index and molar mass for N-terminal MBP-tagged Ctp11–60 (green) (as in Fig. 1d) and MBP-tagged Ctp11–60 (H11A W12A Y16A) (brown).
Supplementary Figure 5 Ctp1 binds DNA.
(a) Quantification of Ctp1–DNA binding. Mean values shown. Error bars, s.d. (n=3). (b) Ctp1FL binding variable lengths of double-stranded DNA. Arrow indicates 40–50bp. (c) Purified Ctp1 deletion and internal deletion protein constructs. (d) Ctp1FL and Ctp1 protein halves binding DNA. (e) Ctp1FL and Ctp1 internal deletion DNA binding assays. WT, wildtype. Ctp1 concentrations expressed as tetrameric (tet) or monomeric (mono) as labeled. Experiments were repeated three times for (a), (b), (d), (e), with representative gels shown.
Supplementary Figure 6 Purified Ctp1FL N- and C-terminal mutations.
(a) Ctp1 N-terminal THDD domain point mutations used in DNA binding studies. (b) Ctp1 C-terminal point mutations used in DNA binding studies. WT, wildtype.
Supplementary Figure 7 Titration of DNA binding by Ctp1 mutants of the conserved RHR motif.
Ctp1 concentrations expressed as tetrameric (tet). Experiment was repeated 3 times with representative gels shown.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–4 (PDF 3777 kb)
Supplementary Data Set 1
Uncropped gels and blots (PDF 2633 kb)
Rights and permissions
About this article
Cite this article
Andres, S., Appel, C., Westmoreland, J. et al. Tetrameric Ctp1 coordinates DNA binding and DNA bridging in DNA double-strand-break repair. Nat Struct Mol Biol 22, 158–166 (2015). https://doi.org/10.1038/nsmb.2945
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2945
This article is cited by
-
Inositol pyrophosphates promote the interaction of SPX domains with the coiled-coil motif of PHR transcription factors to regulate plant phosphate homeostasis
Nature Communications (2021)
-
Phosphopeptide interactions of the Nbs1 N-terminal FHA-BRCT1/2 domains
Scientific Reports (2021)
-
Regulatory control of DNA end resection by Sae2 phosphorylation
Nature Communications (2018)
-
The end-joining factor Ku acts in the end-resection of double strand break-free arrested replication forks
Nature Communications (2017)
-
Cullin3-KLHL15 ubiquitin ligase mediates CtIP protein turnover to fine-tune DNA-end resection
Nature Communications (2016)