Abstract
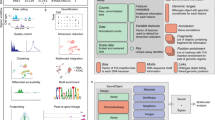
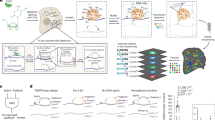
Under continuous, glucose-limited conditions, budding yeast exhibit robust metabolic cycles associated with major oscillations of gene expression. How such fluctuations are linked to changes in chromatin status is not well understood. Here we examine the correlated genome-wide transcription and chromatin states across the yeast metabolic cycle at unprecedented temporal resolution, revealing a 'just-in-time supply chain' by which components from specific cellular processes such as ribosome biogenesis become available in a highly coordinated manner. We identify distinct chromatin and splicing patterns associated with different gene categories and determine the relative timing of chromatin modifications relative to maximal transcription. There is unexpected variation in the chromatin modification and expression relationship, with histone acetylation peaks occurring with varying timing and 'sharpness' relative to RNA expression both within and between cycle phases. Chromatin-modifier occupancy reveals subtly distinct spatial and temporal patterns compared to those of the modifications themselves.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Accession codes
References
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Berger, S.L. The complex language of chromatin regulation during transcription. Nature 447, 407–412 (2007).
Pokholok, D.K. et al. Genome-wide map of nucleosome acetylation and methylation in yeast. Cell 122, 517–527 (2005).
Weiner, A. et al. Systematic dissection of roles for chromatin regulators in a yeast stress response. PLoS Biol. 10, e1001369 (2012).
Millar, C.B. & Grunstein, M. Genome-wide patterns of histone modifications in yeast. Nat. Rev. Mol. Cell Biol. 7, 657–666 (2006).
Brownell, J.E. et al. Tetrahymena histone acetyltransferase A: a homolog to yeast Gcn5p linking histone acetylation to gene activation. Cell 84, 843–851 (1996).
Takahashi, H., McCaffery, J.M., Irizarry, R.A. & Boeke, J.D. Nucleocytosolic acetyl-coenzyme a synthetase is required for histone acetylation and global transcription. Mol. Cell 23, 207–217 (2006).
Krebs, J.E. Moving marks: dynamic histone modifications in yeast. Mol. Biosyst. 3, 590–597 (2007).
Koike, N. et al. Transcriptional architecture and chromatin landscape of the core circadian clock in mammals. Science 338, 349–354 (2012).
Paige, S.L. et al. A temporal chromatin signature in human embryonic stem cells identifies regulators of cardiac development. Cell 151, 221–232 (2012).
Zhang, L., Ma, H. & Pugh, B.F. Stable and dynamic nucleosome states during a meiotic developmental process. Genome Res. 21, 875–884 (2011).
Tu, B.P., Kudlicki, A., Rowicka, M. & McKnight, S.L. Logic of the yeast metabolic cycle: temporal compartmentalization of cellular processes. Science 310, 1152–1158 (2005).
Tu, B.P. et al. Cyclic changes in metabolic state during the life of a yeast cell. Proc. Natl. Acad. Sci. USA 104, 16886–16891 (2007).
Cai, L., Sutter, B.M., Li, B. & Tu, B.P. Acetyl-CoA induces cell growth and proliferation by promoting the acetylation of histones at growth genes. Mol. Cell 42, 426–437 (2011).
Kaochar, S. & Tu, B.P. Gatekeepers of chromatin: small metabolites elicit big changes in gene expression. Trends Biochem. Sci. 37, 477–483 (2012).
Bergkessel, M., Whitworth, G.B. & Guthrie, C. Diverse environmental stresses elicit distinct responses at the level of pre-mRNA processing in yeast. RNA 17, 1461–1478 (2011).
Pleiss, J.A., Whitworth, G.B., Bergkessel, M. & Guthrie, C. Rapid, transcript-specific changes in splicing in response to environmental stress. Mol. Cell 27, 928–937 (2007).
Kurdistani, S.K., Tavazoie, S. & Grunstein, M. Mapping global histone acetylation patterns to gene expression. Cell 117, 721–733 (2004).
Avvakumov, N., Nourani, A. & Côté, J. Histone chaperones: modulators of chromatin marks. Mol. Cell 41, 502–514 (2011).
Dang, W. et al. Histone H4 lysine 16 acetylation regulates cellular lifespan. Nature 459, 802–807 (2009).
Dion, M.F., Altschuler, S.J., Wu, L.F. & Rando, O.J. Genomic characterization reveals a simple histone H4 acetylation code. Proc. Natl. Acad. Sci. USA 102, 5501–5506 (2005).
Moazed, D. Enzymatic activities of Sir2 and chromatin silencing. Curr. Opin. Cell Biol. 13, 232–238 (2001).
Chern, M.K. et al. Yeast ribosomal protein L12 is a substrate of protein-arginine methyltransferase 2. J. Biol. Chem. 277, 15345–15353 (2002).
Kaplan, T. et al. Cell cycle- and chaperone-mediated regulation of H3K56ac incorporation in yeast. PLoS Genet. 4, e1000270 (2008).
Li, B., Carey, M. & Workman, J.L. The role of chromatin during transcription. Cell 128, 707–719 (2007).
Jorgensen, P. et al. A dynamic transcriptional network communicates growth potential to ribosome synthesis and critical cell size. Genes Dev. 18, 2491–2505 (2004).
Megee, P.C., Morgan, B.A. & Smith, M.M. Histone H4 and the maintenance of genome integrity. Genes Dev. 9, 1716–1727 (1995).
Zaslaver, A. et al. Just-in-time transcription program in metabolic pathways. Nat. Genet. 36, 486–491 (2004).
Laxman, S., Sutter, B.M. & Tu, B.P. Behavior of a metabolic cycling population at the single cell level as visualized by fluorescent gene expression reporters. PLoS ONE 5, e12595 (2010).
Morillon, A., Karabetsou, N., Nair, A. & Mellor, J. Dynamic lysine methylation on histone H3 defines the regulatory phase of gene transcription. Mol. Cell 18, 723–734 (2005).
Zhang, J.A., Mortazavi, A., Williams, B.A., Wold, B.J. & Rothenberg, E.V. Dynamic transformations of genome-wide epigenetic marking and transcriptional control establish T cell identity. Cell 149, 467–482 (2012).
Wamstad, J.A. et al. Dynamic and coordinated epigenetic regulation of developmental transitions in the cardiac lineage. Cell 151, 206–220 (2012).
Fischle, W., Wang, Y. & Allis, C.D. Histone and chromatin cross-talk. Curr. Opin. Cell Biol. 15, 172–183 (2003).
Strahl, B.D. & Allis, C.D. The language of covalent histone modifications. Nature 403, 41–45 (2000).
Dai, J. et al. Probing nucleosome function: a highly versatile library of synthetic histone H3 and H4 mutants. Cell 134, 1066–1078 (2008).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc 7, 562–578 (2012).
Li, C. & Wong, W.H. in The Analysis of Gene Expression Data: Methods and Software 120–141 (Springer, 2003).
Ji, H. et al. An integrated software system for analyzing ChIP-chip and ChIP-seq data. Nat. Biotechnol. 26, 1293–1300 (2008).
Xu, Z. et al. Bidirectional promoters generate pervasive transcription in yeast. Nature 457, 1033–1037 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Eilers, P.H. & Marx, B.D. Flexible smoothing with B-splines and penalties. Stat. Sci. 11, 89–102 (1996).
Schwarz, G. Estimating the dimension of a model. Ann. Stat. 6, 461–464 (1978).
Acknowledgements
We dedicate this paper to the memory of our colleague, Yu-yi Lin. We thank L. Huang for general data-analysis advice; J. Dai for advice on histone mutants; L. Shi, S. Uplekar and G. Rao for help with fermentor and YMC experiments; and S. Taverna, B. Cormack, J. Bader, P. Meluh, Y. Lin and members of the Boeke laboratory for helpful discussion. This work was supported by US National Institutes of Health grants R01HG006841 to H.J., R01GM094314 to B.P.T. and U54GM103520 to J.D.B.
Author information
Authors and Affiliations
Contributions
Z.K., L.C., B.P.T. and J.D.B. designed experiments; Z.K. and L.C. collected ChIP-seq and RNA-seq data; Z.K., X.Z. and H.J. performed the analysis; Z.K. made histone mutants and analyzed growth and YMC phenotypes; Z.K. wrote the manuscript with help from all other authors; all authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Dynamics of gene expression in YMC.
(a, b) RNA-seq experiment reveals the same three clusters (OX, RB and RC) discovered by previous microarray experiments. 3 cycles of microarray data are plotted in the first three joint columns, each containing 12 time points. 16 time-point RNA-seq data are plotted in the last joint column. (a) Enriched GO terms are labeled on the right. (b) 537 “non-cycling” genes defined previously show clearly elevated expression level in OX phase. Algorithm previously used to define cycling genes required highest expression of 2 or more time points and highest expression in all three cycle periods. As can be seen, these 537 genes form very sharp peaks with only one high time point and/or are represented in only one or two cycles. (c) Average 16 time-point RNA-seq data across genes in subclusters from Figure 1 (b-d). (d) Expression level of RTT109 and HST3 across 16 time points extracted from the RNA-seq experiment.
Supplementary Figure 2 Temporal patterns of RNA-seq signals at RP introns.
89 RP genes from Intron1 cluster were included in the analysis. (a) The ratio of intron to exon was calculated for every gene in the cluster and the means are displayed across 16 time points in the top panel. The bottom panel is total numbers of junction reads spanning the 3’ end of RP introns at 16 time points. Note that the numbers of 3' junction reads are consistently higher than the numbers of 5′ junction reads (Figure 2c), consistent with an arrest of splicing at the “2/3 intermediate” stage of splicing. Heat maps show the temporal patterns of junction reads spanning the 5’ end (b) and the 3’ end (c) of RP introns and exon-exon junctions (d) of RP genes. Data is centered to mean zero. “T4” or “T5” mark the time point where signals reach the peak. (e) The RNA levels of CK2 subunits, catalytic subunits CKA1 and CKA2, and regulatory subunits CKB1 and CKB2, are plotted across the 16 time points. Time point 4 and 5 are labeled by “T4” and “T5”. (f) RT-qPCRs of 4 Ribi (top panel), 4 RP (middle panel) and 4 RP intron (bottom panel) transcripts across 16 ultra-dense OX time points (1.5~2 min intervals). Signals are normalized by the level at time point 1. 16 values of each transcript are plotted in a smoothed line. The vertical lines in each panel mark the mean peak time of the 4 transcripts and the estimation values are labeled at the right side of the lines.
Supplementary Figure 3 Dynamic chromatin modifications across the YMC.
H3K9ac, H3K14ac, H4K5ac and H3K4me3 ChIP-seq signals at chrVII: 340669-389335 are displayed in cisgenome browser. 16 tracks represent 16 time points from top to bottom consecutively. 3-color bars on the left of each panel mark the three phases. The boxes with the same 3 colors in the last track mark the phases of expression of corresponding genes. Black boxes represent non-cycling genes.
Supplementary Figure 4 Temporal patterns of chromatin modifications at OX genes.
H3K9ac, H3K14ac, H4K5ac and H3K4me3 ChIP-seq signals at FUR1, RPL42B, RPS11B, RRB1, MET5 and LYS1 are displayed in CisGenome browser screenshots. 16 tracks represent 16 time points from top to bottom consecutively. 3-color bars on the left of each panel mark the three phases.
Supplementary Figure 5 Temporal association between chromatin states and gene expression.
(a) Spatial distribution of chromatin markers. ChIP-seq reads are averaged from -1000 to +1000 bp of TSSs across the genome. (b) 14 clusters were determined for the whole genome, revealing complex combinatorial temporal patterns of H3K9ac, H3K14ac, H4K5ac, H3K56ac, H4K16ac, H3K4me3, H3K36me3 and H3. Each row represents centered signals of one gene and each column represents one time-point ChIP-seq corresponding to the indicated chromatin mark. We calculated reads at the 5′ end as gene-specific signals of H3K4me3, H3K9ac, H3K4ac, H4K5ac and H3K56ac, and reads across the full gene as signals of H3K36me3, H4K16ac and H3. (c) Estimation of time in YMC when RNA and histone modifications in “RNA_H3”, “RNA_H4”, “RNA_H5”, “RNA_H6”, “RNA_H7” reach the maximum. The middle of the horizontal colored lines on the O2 curves represent the mean peak value of genes in each cluster and the lengths represent standard deviations for genes in theses clusters.
Supplementary Figure 6 Dynamic analysis of chromatin modifiers and corresponding modifications.
(a) shows the numbers of binding sites and annotated genes by MACS for Gcn5, Esa1 and Set1. Areas of overlap among the three enzymes and numbers of annotated genes in each subset are labeled inside the Venn diagram. (b) 4 distinct patterns of the 1035 genes to which all three enzymes bind show the temporal combinatorial recruitment of the chromatin modifiers. Overrepresented GO terms for each cluster are listed on the right. (c) Gcn5-3FLAG 6HA-Set1 cells were collected at 8 time points from late RC to RB phase in YMC for ChIP-qPCR. (d) ChIP-qPCR of H3K9ac, H3K14ac, H4K5ac and H3K4me3 at examples of OX phase genes. RPS11b and RPL33b are two RP genes and MET5 and LYS1 are aa genes. H3K14ac and H4K5ac were detected earlier than H3K9ac on aa genes but not on RP genes. (e) ChIP-qPCR of Gcn5, Set1 and Esa1 at RPS11b and RPL33b. Esa1 and Set1 binding increase before Gcn5 binding.
Supplementary Figure 7 Analysis of histone-mutant O2 oscillation.
(a) Strains were plated (1:10 serial dilutions) on YPD plates and grown at 30°C. H3 mutants represent H3K9R, H3K(9,14,18)R, H3K(9,14,18,23)R, H3K(9,14,18,23,27)R, H3K(9,14,18,23,27)A, H3K(9,14,18,23,27)Q. H4 mutants represent H4K5R, H4K(5,8)R, H4K(5,8,12)R, H4K(5,8,12)A, H4K(5,8,12)Q. (b) O2 oscillation curves of H3K(9,14,18,23,27)A and H4K(5,8,12)A.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Note (PDF 8185 kb)
Supplementary Table 1
Analysis of the RNA-seq experiments in YMC (XLSX 4859 kb)
Supplementary Table 2
Clustering analysis of the RNA-seq reads at intron-containing genes (XLSX 253 kb)
Supplementary Table 3
Clustering analysis of histone-modification ChIP-seq experiments (XLSX 9794 kb)
Supplementary Table 4
Analysis of combinatorial histone modifications and RNA level (XLSX 160 kb)
Supplementary Table 5
Analysis of the chromatin-modifier ChIP-seq experiments (XLSX 795 kb)
Supplementary Table 6
Primers used in this study (XLSX 56 kb)
Rights and permissions
About this article
Cite this article
Kuang, Z., Cai, L., Zhang, X. et al. High-temporal-resolution view of transcription and chromatin states across distinct metabolic states in budding yeast. Nat Struct Mol Biol 21, 854–863 (2014). https://doi.org/10.1038/nsmb.2881
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2881
This article is cited by
-
Temporal segregation of biosynthetic processes is responsible for metabolic oscillations during the budding yeast cell cycle
Nature Metabolism (2023)
-
Single-cell imaging and RNA sequencing reveal patterns of gene expression heterogeneity during fission yeast growth and adaptation
Nature Microbiology (2019)
-
Stress response factors drive regrowth of quiescent cells
Current Genetics (2018)
-
GATOR1 regulates nitrogenic cataplerotic reactions of the mitochondrial TCA cycle
Nature Chemical Biology (2017)
-
Similarity-Based Segmentation of Multi-Dimensional Signals
Scientific Reports (2017)