Abstract
Mitogen-activated protein kinases (MAPKs) are essential to intracellular signal transduction. MAPKs anchor their pathway-specific substrates through so-called 'docking interactions' at locations distal from the active site. Docking interactions ensure efficient substrate recognition, but their contribution to the kinase reaction itself remains unclear. Herein, we use solution NMR to analyze the interaction between dually phosphorylated, active human p38α and the C-terminal fragments of its substrate MK2. p38α phosphorylation and ATP loading collaboratively induce the active conformation; subsequently, p38α accommodates MK2 phosphoacceptor residues in its active site. The docking interaction enhances binding of ATP and the phosphoacceptor to p38α, accelerating the phosphotransfer reaction. Thus, the docking interaction enhances p38α's enzymatic activity toward pathway-specific substrates allosterically as well as by the anchor effect. These findings clarify how MAPK cascades are organized in cells, even under ATP-depleted conditions often associated with environmental stress.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Raman, M., Chen, W. & Cobb, M.H. Differential regulation and properties of MAPKs. Oncogene 26, 3100–3112 (2007).
Avruch, J. MAP kinase pathways: the first twenty years. Biochim. Biophys. Acta 1773, 1150–1160 (2007).
Cargnello, M. & Roux, P.P. Activation and function of the MAPKs and their substrates, the MAPK-activated protein kinases. Microbiol. Mol. Biol. Rev. 75, 50–83 (2011).
Plotnikov, A., Zehorai, E., Procaccia, S. & Seger, R. The MAPK cascades: signaling components, nuclear roles and mechanisms of nuclear translocation. Biochim. Biophys. Acta 1813, 1619–1633 (2011).
Marshall, C.J. Specificity of receptor tyrosine kinase signaling: transient versus sustained extracellular signal-regulated kinase activation. Cell 80, 179–185 (1995).
Cuadrado, A. & Nebreda, A.R. Mechanisms and functions of p38 MAPK signalling. Biochem. J. 429, 403–417 (2010).
Schindler, J.F., Monahan, J.B. & Smith, W.G. p38 pathway kinases as anti-inflammatory drug targets. J. Dent. Res. 86, 800–811 (2007).
Dodeller, F. & Schulze-Koops, H. The p38 mitogen-activated protein kinase signaling cascade in CD4 T cells. Arthritis Res. Ther. 8, 205 (2006).
Johnson, G.L. & Lapadat, R. Mitogen-activated protein kinase pathways mediated by ERK, JNK, and p38 protein kinases. Science 298, 1911–1912 (2002).
Anderson, N.G., Maller, J.L., Tonks, N.K. & Sturgill, T.W. Requirement for integration of signals from two distinct phosphorylation pathways for activation of MAP kinase. Nature 343, 651–653 (1990).
Zhang, Y.Y., Mei, Z.Q., Wu, J.W. & Wang, Z.X. Enzymatic activity and substrate specificity of mitogen-activated protein kinase p38α in different phosphorylation states. J. Biol. Chem. 283, 26591–26601 (2008).
Keyse, S.M. Protein phosphatases and the regulation of mitogen-activated protein kinase signalling. Curr. Opin. Cell Biol. 12, 186–192 (2000).
Goldsmith, E.J., Cobb, M.H. & Chang, C.I. Structure of MAPKs. Methods Mol. Biol. 250, 127–144 (2004).
Wang, Z. et al. The structure of mitogen-activated protein kinase p38 at 2.1-A resolution. Proc. Natl. Acad. Sci. USA 94, 2327–2332 (1997).
Wilson, K.P. et al. Crystal structure of p38 mitogen-activated protein kinase. J. Biol. Chem. 271, 27696–27700 (1996).
Zhang, F., Strand, A., Robbins, D., Cobb, M. & Goldsmith, E. Atomic structure of the MAP kinase ERK2 at 2.3 A resolution. Nature 367, 704–711 (1994).
Bellon, S., Fitzgibbon, M.J., Fox, T., Hsiao, H.M. & Wilson, K.P. The structure of phosphorylated p38γ is monomeric and reveals a conserved activation-loop conformation. Structure 7, 1057–1065 (1999).
Canagarajah, B.J., Khokhlatchev, A., Cobb, M.H. & Goldsmith, E.J. Activation mechanism of the MAP kinase ERK2 by dual phosphorylation. Cell 90, 859–869 (1997).
Zhang, Y.Y., Wu, J.W. & Wang, Z.X. Mitogen-activated protein kinase (MAPK) phosphatase 3-mediated cross-talk between MAPKs ERK2 and p38α. J. Biol. Chem. 286, 16150–16162 (2011).
Weston, C.R., Lambright, D.G. & Davis, R.J. Signal transduction. MAP kinase signaling specificity. Science 296, 2345–2347 (2002).
Ubersax, J.A. & Ferrell, J.E. Mechanisms of specificity in protein phosphorylation. Nat. Rev. Mol. Cell Biol. 8, 530–541 (2007).
Tanoue, T. & Nishida, E. Docking interactions in the mitogen-activated protein kinase cascades. Pharmacol. Ther. 93, 193–202 (2002).
Tanoue, T., Adachi, M., Moriguchi, T. & Nishida, E. A conserved docking motif in MAP kinases common to substrates, activators and regulators. Nat. Cell Biol. 2, 110–116 (2000).
Gupta, S., Campbell, D., Derijard, B. & Davis, R.J. Transcription factor ATF2 regulation by the JNK signal transduction pathway. Science 267, 389–393 (1995).
Kallunki, T. et al. JNK2 contains a specificity-determining region responsible for efficient c-Jun binding and phosphorylation. Genes Dev. 8, 2996–3007 (1994).
Yang, S.-H., Galanis, A. & Sharrocks, A.D. Targeting of p38 mitogen-activated protein kinases to MEF2 transcription factors. Mol. Cell. Biol. 19, 4028–4038 (1999).
Lukas, S.M. et al. Catalysis and function of the p38αMK2a signaling complex. Biochemistry 43, 9950–9960 (2004).
Chang, C.I., Xu, B.E., Akella, R., Cobb, M.H. & Goldsmith, E.J. Crystal structures of MAP kinase p38 complexed to the docking sites on its nuclear substrate MEF2A and activator MKK3b. Mol. Cell 9, 1241–1249 (2002).
ter Haar, E., Prabhakar, P., Prabakhar, P., Liu, X. & Lepre, C. Crystal structure of the p38α-MAPKAP kinase 2 heterodimer. J. Biol. Chem. 282, 9733–9739 (2007).
Tanoue, T., Maeda, R., Adachi, M. & Nishida, E. Identification of a docking groove on ERK and p38 MAP kinases that regulates the specificity of docking interactions. EMBO J. 20, 466–479 (2001).
Reményi, A., Good, M.C., Bhattacharyya, R.P. & Lim, W.A. The role of docking interactions in mediating signaling input, output, and discrimination in the yeast MAPK network. Mol. Cell 20, 951–962 (2005).
Songyang, Z. et al. A structural basis for substrate specificities of protein Ser/Thr kinases: primary sequence preference of casein kinases I and II, NIMA, phosphorylase kinase, calmodulin-dependent kinase II, CDK5, and Erk1. Mol. Cell. Biol. 16, 6486–6493 (1996).
White, A., Pargellis, C.A., Studts, J.M., Werneburg, B.G. & Farmer, B.T. Molecular basis of MAPK-activated protein kinase 2:p38 assembly. Proc. Natl. Acad. Sci. USA 104, 6353–6358 (2007).
Szafranska, A.E. & Dalby, K.N. Kinetic mechanism for p38 MAP kinase α. FEBS J. 272, 4631–4645 (2005).
Akella, R., Min, X., Wu, Q., Gardner, K.H. & Goldsmith, E.J. The third conformation of p38α MAP kinase observed in phosphorylated p38α and in solution. Structure 18, 1571–1578 (2010).
Jacobs, D., Glossip, D., Xing, H., Muslin, A.J. & Kornfeld, K. Multiple docking sites on substrate proteins form a modular system that mediates recognition by ERK MAP kinase. Genes Dev. 13, 163–175 (1999).
Lee, S. et al. A model of a MAPKsubstrate complex in an active conformation: a computational and experimental approach. PLoS ONE 6, e18594 (2011).
Masterson, L.R., Mascioni, A., Traaseth, N.J., Taylor, S.S. & Veglia, G. Allosteric cooperativity in protein kinase A. Proc. Natl. Acad. Sci. USA 105, 506–511 (2008).
Zimmerman, S.B. & Trach, S.O. Estimation of macromolecule concentrations and excluded volume effects for the cytoplasm of Escherichia coli. J. Mol. Biol. 222, 599–620 (1991).
Lugo, T.G., Pendergast, A.M., Muller, A.J. & Witte, O.N. Tyrosine kinase activity and transformation potency of bcr-abl oncogene products. Science 247, 1079–1082 (1990).
Heisterkamp, N. et al. Acute leukaemia in bcr/abl transgenic mice. Nature 344, 251–253 (1990).
Dennis, P.B. et al. Mammalian TOR: a homeostatic ATP sensor. Science 294, 1102–1105 (2001).
Knight, Z.A. & Shokat, K.M. Features of selective kinase inhibitors. Chem. Biol. 12, 621–637 (2005).
Steenbergen, C., Murphy, E., Watts, J.A. & London, R.E. Correlation between cytosolic free calcium, contracture, ATP, and irreversible ischemic injury in perfused rat heart. Circ. Res. 66, 135–146 (1990).
Schütt, F., Aretz, S., Auffarth, G.U. & Kopitz, J. Moderately reduced ATP levels promote oxidative stress and debilitate autophagic and phagocytic capacities in human RPE cells. Invest. Ophthalmol. Vis. Sci. 53, 5354–5361 (2012).
Kumphune, S., Chattipakorn, S. & Chattipakorn, N. Role of p38 inhibition in cardiac ischemia/reperfusion injury. Eur. J. Clin. Pharmacol. 68, 513–524 (2012).
Nielsen, G. & Schwalbe, H. NMR spectroscopic investigations of the activated p38α mitogen-activated protein kinase. ChemBioChem 12, 2599–2607 (2011).
Francis, D.M. et al. Structural basis of p38α regulation by hematopoietic tyrosine phosphatase. Nat. Chem. Biol. 7, 916–924 (2011).
Piserchio, A. et al. Docking interactions of hematopoietic tyrosine phosphatase with MAP kinases ERK2 and p38α. Biochemistry 51, 8047–8049 (2012).
Arnold, K., Bordoli, L., Kopp, J. & Schwede, T. The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22, 195–201 (2006).
Wang, Z. et al. Structural basis of inhibitor selectivity in MAP kinases. Structure 6, 1117–1128 (1998).
Gardner, K.H., Rosen, M.K. & Kay, L.E. Global folds of highly deuterated, methyl-protonated proteins by multidimensional NMR. Biochemistry 36, 1389–1401 (1997).
Rosen, M.K. et al. Selective methyl group protonation of perdeuterated proteins. J. Mol. Biol. 263, 627–636 (1996).
Underwood, K.W. et al. Catalytically active MAP KAP kinase 2 structures in complex with staurosporine and ADP reveal differences with the autoinhibited enzyme. Structure 11, 627–636 (2003).
Kinoshita, E., Takahashi, M., Takeda, H., Shiro, M. & Koike, T. Recognition of phosphate monoester dianion by an alkoxide-bridged dinuclear zinc(II) complex. Dalton Trans. 21, 1189–1193 (2004).
Vogtherr, M. et al. NMR backbone assignment of the mitogen-activated protein (MAP) kinase p38. J. Biomol. NMR 32, 175 (2005).
Tugarinov, V., Hwang, P.M., Ollerenshaw, J.E. & Kay, L.E. Cross-correlated relaxation enhanced 1H-13C NMR spectroscopy of methyl groups in very high molecular weight proteins and protein complexes. J. Am. Chem. Soc. 125, 10420–10428 (2003).
Goto, N.K., Gardner, K.H., Mueller, G.A., Willis, R.C. & Kay, L.E. A robust and cost-effective method for the production of Val, Leu, Ile (δ1) methyl-protonated 15N-, 13C-, 2H-labeled proteins. J. Biomol. NMR 13, 369–374 (1999).
Amero, C. et al. Fast two-dimensional NMR spectroscopy of high molecular weight protein assemblies. J. Am. Chem. Soc. 131, 3448–3449 (2009).
Maruyama, Y. et al. Human Gene and Protein Database (HGPD): a novel database presenting a large quantity of experiment-based results in human proteomics. Nucleic Acids Res. 37, D762–D766 (2009).
Maruyama, Y. et al. HGPD: Human Gene and Protein Database, 2012 update. Nucleic Acids Res. 40, D924–D929 (2012).
Schanda, P., Kupce, E. & Brutscher, B. SOFAST-HMQC experiments for recording two-dimensional heteronuclear correlation spectra of proteins within a few seconds. J. Biomol. NMR 33, 199–211 (2005).
Selenko, P. et al. In situ observation of protein phosphorylation by high-resolution NMR spectroscopy. Nat. Struct. Mol. Biol. 15, 321–329 (2008).
Acknowledgements
We would like to thank H. Hanzawa (Daiichi Sankyo Co.) for providing the expression plasmid for p38α. We are also indebted to N. Goshima (Japanese National Institute of Advanced Industrial Science and Technology) for providing cDNA clones for the phosphatases PPM1A and HePTP. This work was funded by grants from the Japan New Energy and Industrial Technology Development Organization (NEDO). Funding was also provided by Grants-in-Aid for Scientific Research on Innovative Areas (25121743 to K.T.) from the Japanese Ministry of Education, Culture, Sports, Science and Technology (MEXT) and Japan Society for the Promotion of Science (JSPS).
Author information
Authors and Affiliations
Contributions
Y.T., K.T., H.T. and I.S. conceived the project. Y.T. performed the experiments. Y.T., K.T., H.T. and I.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Characterization of p38α and the model substrates.
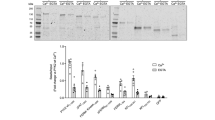
(a) Phos-tagTM SDS-PAGE of p38α, which is dually phosphorylated by MKK6DD (lane 2). The sample was then dephosphorylated by either HePTP, a phosphatase that is specific to phospho-Tyr182 (lanes 3 and 4), or PPM1A, a phosphatase that is specific to phospho-Thr180 (lanes 7 and 8). In lanes 5, 6, 9, and 10, the p38α-2P was treated with both of the phosphtases in different orders. The mobility of p38α-2P treated with HePTP increased (p38α-1PT), indicating the dephosphorylation of Tyr182 (lanes 3 and 4). Thr180, however, remained phosphorylated, as the mobility is still slower than unphosphorylated p38α. Further treatment of p38α-1PT with PPM1A led to the complete regression to unphosphorylated p38α (lanes 5 and 6). This indicated that p38α was successfully dually phosphorylated by MKK6DD. Although p38α-2P first treated with PPM1A exhibited almost the same mobility as p38α-2P (p38α-1PY; lanes 7 and 8), the subsequent treatment with HePTP resulted in the same mobility as unphosphorylated p38α. It seems that the mobilities of p38α-1PY and p38α-2P are indiscernible. The details of the reaction conditions are described in the Online Methods. (b-i) Characterization of the phosphorylation states of p38α based on the spectral pattern of the Thr-γ2 region of the 1H-13C HSQC spectra. Panels (b) to (e) show the spectra of p38α, p38α-2P, p38α-1PT, and p38α-1PY, respectively. The p38α-1PT and p38α-1PY proteins were prepared by treating p38α-2P with HePTP and PPM1A, respectively. Methyl resonances with distinctive chemical shifts in the respective phosphorylation states are highlighted by the blue rectangles. Panels (f) to (i) are the spectra of p38α phosphorylated with 1/70 mol. eq. of MKK6DD at 14 °C for the indicated times. A 20 hour reaction rendered most of the p38α dually phosphorylated. (j) Phos-tagTM SDS-PAGE of MK2 (47–400) phosphorylated by p38α-2P. Lane 1 is the reference which is not treated with p38α-2P, and lanes 2 to 7 show the timecourse of the phosphorylation of 5 μM MK2 (47–400) by 0.5 μM p38α-2P in the presence of 2 mM ATP at 25 °C. The time-dependent, discrete decrease in the mobility is indicative of the multisite phosphorylation of MK2 (47–400) by p38α-2P. (k) The retention volumes in the SEC analyses of p38α-2P in the absence and presence of 1 mM ATP. Samples were prepared as 240 μL solution of 25 μM p38α and were analyzed by chromatography on a Superdex 200 GL 10 300 column connected to AKTA Explorer 10S (GE Healthcare) at 4 °C. The retention volumes and the error bars are the averages and the standard deviations of five repeated runs. The larger error bar in the presence of ATP is due to the rough baseline caused by the high background absorbance at 280 nm by ATP. Nevertheless, the retention volume in the presence of ATP is larger than that in the absence of ATP, suggesting that the conformation of p38α-2P is more compact (i.e. “closed”) when ATP is loaded. (l) Competition experiment between the 334D-peptide and MK2 (47-400). Western blot of p38α-2P, detected with 1:1000 dilution of the anti-p38 monoclonal antibody. The N-terminally GB1-fused 334D-peptide was immobilized on IgG Sepharose beads, and then p38α-2P was further immobilized on them (input; lane 6). After washing the beads, p38α-2P was eluted with the longer MK2 fragment (47–400) (0.3, 0.6, and 1.2 μM; lanes 2–4). The intensity of the p38α-2P band increased in proportion to the MK2 concentration, which indicates that the longer MK2 fragment competes with the GB1-334D-peptide for the same site on p38α-2P. The dissociation of p38α-2P from the beads by washing is negligible, as the band is very weak when the elution buffer without the longer MK2 fragment is used (lane 1). The non-specific binding of p38α-2P on the beads is also negligible, as shown in the experiment using GB1 instead of the GB1-334D-peptide (lane 5). The detailed experimental conditions are provided in the Online Methods. (m) Phosphorylations of the 334D-peptide and the 334-peptide by p38α-2P were monitored by time-dependent increase of the phospho-334D-peptide (red circles) and the phospho-334-peptide (cyan squares). The reactions were tracked by successive measurements of 1H-15N SOFAST-HMQC spectra. The concentration of the phosphorylated product was estimated from the intensity of the phospho-Thr334 resonance for each peptide. The peptides (100 μM) were phosphorylated by 20 nM of p38α-2P, in the presence of 2 mM ATP at 25 °C. The error bars correspond to the level of noise in each spectrum.
Supplementary Figure 2 Effect of phosphorylation and ATP titration on p38α.
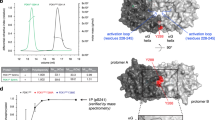
(a) Overlay of the annotated Leu/Val methyl 1H-13C SOFAST-HMQC spectra of p38α-2P in the absence (black) and presence (red) of 5 mM ATP analog. Large chemical shift changes are indicated by blue arrows. (b) Left, center: Overlay of the Ile(δ1) and Leu/Val methyl 1H-13C SOFAST-HMQC spectra of 20 μM [ILV-methyl] p38α-2P in the presence (red) and absence (black) of 5 mM ATP. Large chemical shift changes in the Ile(δ1) region are indicated by blue arrows. Spectra were measured with an 800 MHz 1H resonance frequency spectrometer at 10 °C. Right: Methyl sites that exhibited chemical shift changes larger than the linewidth or were not assigned only in the ATP-bound state are indicated as red spheres on the crystal structure (PDB code: 1A9U51). The ATP-binding site is highlighted in the orange oval. (c) Overlay of the annotated Leu/Val methyl 1H-13C SOFAST-HMQC spectra of p38α-2P (blue) and unphosphorylated p38α (black) in the apo state. (d) Comparison of the Ile(δ1) and Leu/Val methyl regions of 1H-13C HMQC spectra of 80 μM [ILVM-methyl] unphosphorylated p38α, in the presence (green) and absence (red) of 4 mM ATP. Spectra were measured at 800 MHz (1H frequency) at 25 °C. For the methyl resonances of Leu and Val residues, only those dispersed in the spectra are labeled, for clarification. (e) Met methyl regions of 1H-13C HMQC spectra of unphosphorylated p38α titrated with various concentration of ATP. The resonance of Met109, which was traced during the titration to determine the affinity of unphosphorylated p38α for ATP, is expanded as an inset, in which all of the titration points are overlaid. The dissociation constant of unphosphorylated p38α for ATP was estimated to be larger than 15 mM, based on the CSPs of the Met109 methyl 1H and 13C resonances. Since the chemical shift changes did not reach a plateau at the highest ATP concentration (15 mM), a unique dissociation constant was not obtained.
Supplementary Figure 3 Changes in 1H-15N TROSY spectra of p38α upon dual phosphorylation.
Comparison of 1H-15N TROSY spectra of unphosphorylated p38α (p38α-0P; black) and p38α-2P (red) in the absence of ATP. The regions containing the resonances of Met179, Thr180, Gly181, Tyr182, and Val183, which are located in the activation loop, are expanded to indicate that these resonances disappeared upon dual phosphorylation. In the lower left panel, the signal from p38α-2P, which overlaps with the Met179 signal from p38α-0P, does not belong to Met179 and is presumably from the unassigned resonance (n.a.) below.
Supplementary Figure 4 Effects of substrate binding on p38α NMR spectra.
(a) Overlay of the annotated Leu/Val methyl 1H-13C SOFAST-HMQC spectra of p38α-2P without the ATP analog, in the absence (black) and presence (red) of an equimolar amount of the 334D-peptide. Large chemical shift changes are indicated by blue arrows. (b) The same as (a), except that p38α-2P is in complex with the ATP analog.
Supplementary Figure 5 Interaction between the substrate phosphoacceptor site and p38α-2P upon ATP loading.
(a) Extensive interaction between the 334D-peptide and p38α-2P upon the ATP analog loading. Comparison of the 1H-15N TROSY spectra of a stoichiometric complex of 180 μM [U-15N] 334D-peptide and unlabeled p38α-2P, in the absence (black) and presence (red) of 5 mM ATP analog. Several resonances, including that of the phosphoacceptor residue, Thr334, lost intensity when the ATP analog was added, reflecting the increase in the transverse relaxation rate. This would be explained by the increase in the effective local rotational correlation times due to the binding of Thr334 to the active site of p38α-2P and/or the generation of chemical exchange with relatively long (msec – μsec) time scale associating with the binding. This suggested that an extensive interaction is formed by the loading of the ATP analog on p38α-2P. (b) Spectral perturbations induced upon the addition of the T334A mutants of 334D-peptide to p38α-2P. Comparison of the regions of the 1H-13C HMQC spectra of 40 μM [ILV-methyl] p38α-2P that contain the resonance of either Ile229 or Ile259, which is located around the P+1 site, in the absence (black) and presence (red) of an equimolar amount of the T334A mutant of the 334D-peptide. The panels on the left and right are the spectra in the absence of presence of ATP (not analog; 5 mM). The addition of the unlabeled peptide induced CSPs in the P+1 site resonances only in the presence of ATP. The addition of the peptide induced CSPs to the P+1 site resonances of p38α-2P in the presence of ATP. Spectra were recorded at 10 °C, with an 800 MHz 1H resonance frequency spectrometer. (c, d) Comparison of the Ile-δ1 regions of 1H-13C HMQC spectra of 40 μM [ILV-methyl] p38α-2P, in the absence (black) and presence (red) of a 10-fold molar excess of the 334-peptide without (c) and with (d) a saturating amount of the ATP analog (5 mM). Regions containing the resonances of Ile229 and Ile259, which are located near the P+1 site, are enlarged. In the presence of the ATP analog, the resonance from Ile229 disappeared upon the addition of the 334-peptide. (e) Spectral perturbations induced upon the addition of the T334A mutants 334-peptide to p38α-2P. The experimental conditions are the same as (b), except that the titrated peptide was a 10-fold molar excess of the unlabeled 334-peptide with the T334A mutation. (f, g) Extensive interaction between the 334-peptide and p38α-2P induced by the ATP analog. The methyl region of the one dimensional (1D) proton NMR spectrum of 100 μM unlabeled 334-peptide in the free form (f, blue) was compared to that with 50 μM p38α-2P in the absence (g, black) and presence of the ATP analog (4 mM; g, red). The signals from the 334-peptide were not significantly affected by the presence of p38α-2P itself, while the broad signals derived from p38α-2P are also observed (g, black). However, considerable broadening of the 334-peptide signals were observed upon the addition of 4 mM ATP analog (g, red), indicating that the transverse relaxation rates of the methyl proton resonances of the 334-peptide increased. This would result from the increased effective rotational correlation time of the 334-peptide upon the binding to p38α-2P, as well as the relaxation enhancement due to the exchange between free and bound states with distinct chemical shifts. Thus, the 334-peptide interacts with p38α-2P, which is loaded with the ATP analog.
Supplementary Figure 6 Allosteric effects induced by the docking interaction to p38α-2P.
(a) Comparison of the ILV-methyl regions of the 1H-13C HMQC spectra of p38α-2P in the absence (black) and presence (magenta) of a 1.1 molar equivalent of the D-peptide. Spectra were recorded in the presence of 5 mM ATP analog. Regions containing the resonances of Val89, located in the ATP-binding site, and Ile259, located in the P+1 site, are shown in orange and cyan squares, respectively. (b) Mapping of the methyl groups that showed CSPs larger than 0.02 ppm, upon the addition of the D-peptide in the presence of 5 mM ATP analog, shown in red, while those that are not assigned only in the presence of the ATP analog are shown in black. The docking sequence of MK2 is shown as purple ribbon. The structure figure was generated from the X-ray structure of p38α in complex with the MK2 docking-peptide (PDB code: 2OKR29).
Supplementary Figure 7 Enhancements of catalytic steps of p38α by the docking interaction.
(a) Determination of the affinity of p38α-2P to the ATP analog. Ile84-δ1 signals in the 1H-13C HMQC spectra of 40 μM [ILV-methyl] p38α-2P, without (upper) and with (lower) a 1.1 molar excess of the D-peptide. Both samples contained 200 μM of the ATP analog. High field (upper right) and low field resonances (lower left) are from the ATP analog unbound and bound states, respectively. 1D slices in the 1H dimension are overlaid on the 2D signals. The intensity of the resonance from the ATP analog-bound state becomes stronger in the presence of the D-peptide. (b) Determination of the affinity of p38α-2P to ATP. Left; Expanded regions of the spectra for the probe resonances. The upper and lower rows correspond to the conditions without and with a 1.1 molar equivalent of the D-peptide, respectively. From left to right, the conditions represent 5 mM ATP analog, the mixture of 2.5 mM ATP analog and 2.5 mM ATP, and 5 mM ATP, respectively. Right; The intensities of the probe resonances with the different concentrations of ATP and the ATP analog. The intensities of the resonances in each condition were normalized to the reference spectra for the ATP-bound form. The upper and lower panels are the data in the absence and presence of the D-peptide, respectively. The intensities for the ATP and ATP analog bound resonances were plotted as the orange squares and green diamonds, respectively, and were used to determine the affinity of ATP, as described in the Supplemental Materials and Methods. The error bars correspond to the level of noise in each spectrum. (c) Titration of the 334-peptide to p38α-2P. Overlays of the Ile-δ1 regions of 1H-13C HMQC spectra of 40 μM [ILV-methyl] p38α-2P with different concentrations of the 334-peptide, in the absence (left) and presence (right) of a 1.1 molar equivalent of the D-peptide. Black, orange, and red spectra correspond to 0, 120, and 400 μM concentrations of the 334-peptide, respectively. Only three spectra of the five titration points are shown for visual clarity. The resonances of Ile259 near the P+1 site are indicated by the cyan rectangles, and are enlarged in the insets. The dose-dependent CSPs of the Ile259 methyl resonance were used to determining the affinity of ATP-loaded p38α-2P for the 334-peptide. (d) Phosphorylation of the 334-peptide by p38α-2P. Left; Expanded region of the phosphorylated Thr334 resonance in the 1H-15N HSQC spectra of 100 μM [U-15N] 334-peptide, before (black) and after (red) phosphorylation by p38α-2P. Only after reaction, the resonance of the phosphorylated Thr334 is observed, with the low-field chemical shifts characteristic of phosphorylated residues64. Right; Time-dependent increase of the phosphorylated 334-peptide produced by p38α-2P, in the absence (black) and presence (magenta) of the D-peptide. The concentration of the product was estimated from the intensity of the phospho-Thr334 resonance. The detailed experimental conditions are provided in the Online Methods.
Supplementary Figure 8 Determination of the affinities of the docking fragments for p38α-2P.
(a-d) NMR titration experiments of the docking fragment of MEF2A to p38α-2P, in the absence and presence of the ATP analog. (a, b) Comparison of the 1H-13C HMQC spectra (Ile-δ1 region) of 20 μM [ILV-methyl] p38α-2P with different concentrations of the MEF2A docking fragment in the absence (a) and presence (b) of 4 mM ATP analog. The arrows trace the chemical shift changes of Ile116, which exhibited the largest CSP values. (c, d) CSPs induced in the methyl resonances of Val83, Ile116, Val117, Val127, Ile131, Ile134, and Val158, which are located in and near the docking site, by titration of the MEF2A docking fragment. The dissociation constants are indicated in the plots with standard deviation estimated from fitting errors. (e, f) ITC experiments of the D-peptide to p38α-2P in the absence (e) and presence (f) of the ATP analog. The dissociation constants are indicated in the plots. The detailed experimental conditions are provided in the Online Methods.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Table 1 (PDF 2186 kb)
Rights and permissions
About this article
Cite this article
Tokunaga, Y., Takeuchi, K., Takahashi, H. et al. Allosteric enhancement of MAP kinase p38α's activity and substrate selectivity by docking interactions. Nat Struct Mol Biol 21, 704–711 (2014). https://doi.org/10.1038/nsmb.2861
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2861
This article is cited by
-
Diversity and versatility of p38 kinase signalling in health and disease
Nature Reviews Molecular Cell Biology (2021)
-
The role of NMR in leveraging dynamics and entropy in drug design
Journal of Biomolecular NMR (2020)
-
The methyl 13C-edited/13C-filtered transferred NOE for studying protein interactions with short linear motifs
Journal of Biomolecular NMR (2020)
-
Identification and classification of small molecule kinases: insights into substrate recognition and specificity
BMC Evolutionary Biology (2016)
-
An Allosteric Cross-Talk Between the Activation Loop and the ATP Binding Site Regulates the Activation of Src Kinase
Scientific Reports (2016)