Abstract
The mechanism of activation of the alternative lengthening of telomeres (ALT) pathway of mammalian chromosome-end maintenance has been unclear. We have now discovered that co-depletion of the histone chaperones ASF1a and ASF1b in human cells induced all hallmarks of ALT in both primary and cancer cells. These included the formation of ALT-associated PML (promyelocytic leukemia) bodies (APBs), the presence of extrachromosomal telomeric DNA species, an elevated frequency of telomeric sister chromatid exchanges (t-SCE) events and intertelomeric exchange of an integrated tag. The induction of ALT characteristics in this setting led to the simultaneous suppression of telomerase. We determined that ALT induction is positively regulated by the proteins RAD17 and BLM and negatively regulated by EXO1 and DNA2. The induction of ALT phenotypes as a consequence of ASF1 depletion strongly supports the hypothesis that ALT is a consequence of histone management dysfunction.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Cesare, A.J. & Reddel, R.R. Alternative lengthening of telomeres: models, mechanisms and implications. Nat. Rev. Genet. 11, 319–330 (2010).
Heaphy, C.M.C. et al. Prevalence of the alternative lengthening of telomeres telomere maintenance mechanism in human cancer subtypes. Am. J. Pathol. 179, 1608–1615 (2011).
Henson, J.D. & Reddel, R.R. Assaying and investigating Alternative Lengthening of Telomeres activity in human cells and cancers. FEBS Lett. 584, 3800–3811 (2010).
Londoño-Vallejo, J.A., Der-Sarkissian, H., Cazes, L., Bacchetti, S. & Reddel, R.R. Alternative lengthening of telomeres is characterized by high rates of telomeric exchange. Cancer Res. 64, 2324–2327 (2004).
Yeager, T.R.T. et al. Telomerase-negative immortalized human cells contain a novel type of promyelocytic leukemia (PML) body. Cancer Res. 59, 4175–4179 (1999).
Cesare, A.J.A. & Griffith, J.D.J. Telomeric DNA in ALT cells is characterized by free telomeric circles and heterogeneous t-loops. Mol. Cell. Biol. 24, 9948–9957 (2004).
Heaphy, C.M. et al. Altered telomeres in tumors with ATRX and DAXX mutations. Science 333, 425 (2011).
Schwartzentruber, J. et al. Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 482, 226–231 (2012).
Lovejoy, C.A.C. et al. Loss of ATRX, genome instability, and an altered DNA damage response are hallmarks of the alternative lengthening of telomeres pathway. PLoS Genet. 8, e1002772 (2012).
Bower, K. et al. Loss of wild-type ATRX expression in somatic cell hybrids segregates with activation of Alternative Lengthening of Telomeres. PLoS ONE 7, e50062 2012).
Tyler, J.K. et al. The RCAF complex mediates chromatin assembly during DNA replication and repair. Nature 402, 555–560 (1999).
Tagami, H., Ray-Gallet, D., Almouzni, G. & Nakatani, Y. Histone H3.1 and H3.3 complexes mediate nucleosome assembly pathways dependent or independent of DNA synthesis. Cell 116, 51–61 (2004).
Green, E.M.E. et al. Replication-independent histone deposition by the HIR complex and Asf1. Curr. Biol. 15, 2044–2049 (2005).
Tang, Y. et al. Structure of a human ASF1a-HIRA complex and insights into specificity of histone chaperone complex assembly. Nat. Struct. Mol. Biol. 13, 921–929 (2006).
Groth, A. et al. Regulation of Replication Fork Progression Through Histone Supply and Demand. Science 318, 1928–1931 (2007).
Groth, A. et al. Human Asf1 regulates the flow of S phase histones during replicational stress. Mol. Cell 17, 301–311 (2005).
Jasencakova, Z. et al. Replication stress interferes with histone recycling and predeposition marking of new histones. Mol. Cell 37, 736–743 (2010).
Dunham, M.A., Neumann, A.A., Fasching, C.L. & Reddel, R.R. Telomere maintenance by recombination in human cells. Nat. Genet. 26, 447–450 (2000).
Pickett, H.A., Cesare, AJ., Johnston, R.L., Neumann, A.A. & Reddel, R.R. Control of telomere length by a trimming mechanism that involves generation of t-circles. EMBO J. 28, 799–809 (2009).
Henson, J.D. et al. DNA C-circles are specific and quantifiable markers of alternative-lengthening-of-telomeres activity. Nat. Biotechnol. 27, 1181–1185 (2009).
Zellinger, B., Akimcheva, S., Puizina, J., Schirato, M. & Riha, K. Ku suppresses formation of telomeric circles and alternative telomere lengthening in Arabidopsis. Mol. Cell 27, 163–169 (2007).
Damm, K. et al. A highly selective telomerase inhibitor limiting human cancer cell proliferation. EMBO J. 20, 6958–6968 (2001).
Lin, S.Y. & Elledge, S.J. Multiple tumor suppressor pathways negatively regulate telomerase. Cell 113, 881–889 (2003).
Zhang, H. & Cohen, S.N.S. Smurf2 up-regulation activates telomere-dependent senescence. Genes Dev. 18, 3028–3040 (2004).
Wu, K.J.K. et al. Direct activation of TERT transcription by c-MYC. Nat. Genet. 21, 220–224 (1999).
Palm, W. & de Lange, T. How shelterin protects mammalian telomeres. Annu. Rev. Genet. 42, 301–334 (2008).
Stracker, T.H. & Petrini, J.H.J. The MRE11 complex: starting from the ends. Nat. Rev. Mol. Cell Biol. 12, 90–103 (2011).
Toledo, L.I. et al. A cell-based screen identifies ATR inhibitors with synthetic lethal properties for cancer-associated mutations. Nat. Struct. Mol. Biol. 18, 721–727 (2011).
Hickson, I. et al. Identification and characterization of a novel and specific inhibitor of the ataxia-telangiectasia mutated kinase ATM. Cancer Res. 64, 9152–9159 (2004).
Leahy, J.J.J. et al. Identification of a highly potent and selective DNA-dependent protein kinase (DNA-PK) inhibitor (NU7441) by screening of chromenone libraries. Bioorg. Med. Chem. Lett. 14, 6083–6087 (2004).
Vannier, J.-B.J., Pavicic-Kaltenbrunner, V.V., Petalcorin, M.I.R.M., Ding, H.H. & Boulton, S.J.S. RTEL1 dismantles T loops and counteracts telomeric G4-DNA to maintain telomere integrity. Cell 149, 795–806 (2012).
Sarkies, P., Reams, C., Simpson, L.J. & Sale, J.E. Epigenetic instability due to defective replication of structured DNA. Mol. Cell 40, 703–713 (2010).
Burrell, R.A. et al. Replication stress links structural and numerical cancer chromosomal instability. Nature 494, 492–496 (2013).
Jiang, W.-Q., Zhong, Z.-H., Henson, J.D. & Reddel, R.R. Identification of candidate alternative lengthening of telomeres genes by methionine restriction and RNA interference. Oncogene 26, 4635–4647 (2007).
Draskovic, I. et al. Probing PML body function in ALT cells reveals spatiotemporal requirements for telomere recombination. Proc. Natl. Acad. Sci. USA 106, 15726–15731 (2009).
Muntoni, A. & Reddel, R.R. The first molecular details of ALT in human tumor cells. Hum. Mol. Genet. 14 (Spec. No. 2), R191–R196 (2005).
Wang, X.X. et al. Rad17 phosphorylation is required for claspin recruitment and Chk1 activation in response to replication stress. Mol. Cell 23, 331–441 (2006).
Nimonkar, A.V. et al. BLM-DNA2-RPA-MRN and EXO1-BLM-RPA-MRN constitute two DNA end resection machineries for human DNA break repair. Genes Dev. 25, 350–362 (2011).
Lin, W. et al. Mammalian DNA2 helicase/nuclease cleaves G-quadruplex DNA and is required for telomere integrity. EMBO J. 32, 1425–1439 (2013).
Davies, S.L., North, P.S. & Hickson, I.D. Role for BLM in replication-fork restart and suppression of origin firing after replicative stress. Nat. Struct. Mol. Biol. 14, 677–679 (2007).
Gravel, S., Chapman, J.R., Magill, C. & Jackson, S.P. DNA helicases Sgs1 and BLM promote DNA double-strand break resection. Genes Dev. 22, 2767–2772 (2008).
Cejka, P. et al. DNA end resection by Dna2-Sgs1-RPA and its stimulation by Top3-Rmi1 and Mre11-Rad50-Xrs2. Nature 467, 112–116 (2010).
Davies, S.L.S., North, P.S.P., Dart, A.A., Lakin, N.D.N. & Hickson, I.D.I. Phosphorylation of the Bloom's syndrome helicase and its role in recovery from S-phase arrest. Mol. Cell. Biol. 24, 1279–1291 (2004).
Chan, K.L., Palmai-Pallag, T., Ying, S. & Hickson, I.D. Replication stress induces sister-chromatid bridging at fragile site loci in mitosis. Nat. Cell Biol. 11, 753–760 (2009).
Barefield, C. & Karlseder, J. The BLM helicase contributes to telomere maintenance through processing of late-replicating intermediate structures. Nucleic Acids Res. 40, 7358–7367 (2012).
Sfeir, A. et al. Mammalian telomeres resemble fragile sites and require TRF1 for efficient replication. Cell 138, 90–103 (2009).
Ralf, C., Hickson, I.D. & Wu, L. The Bloom's syndrome helicase can promote the regression of a model replication fork. J. Biol. Chem. 281, 22839–22846 (2006).
Nguyen, G.H. et al. A small molecule inhibitor of the BLM helicase modulates chromosome stability in human cells. Chem. Biol. 20, 55–62 (2013).
Hu, J. et al. Antitelomerase therapy provokes ALT and mitochondrial adaptive mechanisms in cancer. Cell 148, 651–663 (2012).
Martínez, P. et al. Increased telomere fragility and fusions resulting from TRF1 deficiency lead to degenerative pathologies and increased cancer in mice. Genes Dev. 23, 2060–2075 (2009).
Makarov, V.L., Lejnine, S., Bedoyan, J. & Langmore, J.P. Nucleosomal organization of telomere-specific chromatin in rat. Cell 73, 775–787 (1993).
Pisano, S. et al. Telomeric nucleosomes are intrinsically mobile. J. Mol. Biol. 369, 1153–1162 (2007).
O'Sullivan, R.J., Kubicek, S., Schreiber, S.L. & Karlseder, J. Reduced histone biosynthesis and chromatin changes arising from a damage signal at telomeres. Nat. Struct. Mol. Biol. 17, 1218–1225 (2010).
Conomos, D. et al. Variant repeats are interspersed throughout the telomeres and recruit nuclear receptors in ALT cells. J. Cell Biol. 199, 893–906 (2012).
de Wilde, R.F. et al. Loss of ATRX or DAXX expression and concomitant acquisition of the alternative lengthening of telomeres phenotype are late events in a small subset of MEN-1 syndrome pancreatic neuroendocrine tumors. Mod. Pathol. 25, 1033–1039 (2012).
Corpet, A. et al. Asf1b, the necessary Asf1 isoform for proliferation, is predictive of outcome in breast cancer. EMBO J. 30, 480–493 (2011).
Hartford, S.A.S. et al. Minichromosome maintenance helicase paralog MCM9 is dispensible for DNA replication but functions in germ-line stem cells and tumor suppression. Proc. Natl. Acad. Sci. USA 108, 17702–17707 (2011).
Jiang, W.-Q., Nguyen, A., Cao, Y., Chang, A.C.-M. & Reddel, R.R. HP1-mediated formation of alternative lengthening of telomeres-associated PML bodies requires HIRA but not ASF1a. PLoS ONE 6, e17036 (2011).
Elsässer, S.J. et al. DAXX envelops an H3.3–H4 dimer for H3.3-specific recognition. Nature 491, 560–565 (2012).
Liu, P. et al. Chromosome catastrophes involve replication mechanisms generating complex genomic rearrangements. Cell 146, 889–903 (2011).
Moffat, J. et al. A lentiviral RNAi library for human and mouse genes applied to an arrayed viral high-content screen. Cell 124, 1283–1298 (2006).
Cesare, A.J., Hayashi, M.T., Crabbe, L. & Karlseder, J. The telomere deprotection response is functionally distinct from the genomic DNA damage response. Mol. Cell 51, 141–155 (2013).
Everett, R.D. et al. PML contributes to a cellular mechanism of repression of herpes simplex virus type 1 infection that is inactivated by ICP0. J. Virol. 80, 7995–8005 (2006).
You, Z. et al. CtIP links DNA double-strand break sensing to resection. Mol. Cell 36, 954–969 (2009).
Duxin, J.P. et al. Okazaki fragment processing-independent role for human Dna2 enzyme during DNA replication. J. Biol. Chem. 287, 21980–21991 (2012).
Crabbe, L., Verdun, R., Haggblom, C. & Karlseder, J. Defective telomere lagging strand synthesis in cells lacking WRN helicase activity. Science 306, 1951–1953 (2004).
Theunissen, J.-W.F. & Petrini, J.H.J. Methods for studying the cellular response to DNA damage: influence of the Mre11 complex on chromosome metabolism. Methods Enzymol. 409, 251–284 (2006).
Lewis, P.W., Elsaesser, S.J., Noh, K.-M., Stadler, S.C. & Allis, C.D. Daxx is an H3.3-specific histone chaperone and cooperates with ATRX in replication-independent chromatin assembly at telomeres. Proc. Natl. Acad. Sci. USA 107, 14075–14080 (2010).
Hayashi, M.T.M., Cesare, A.J.A., Fitzpatrick, J.A.J.J., Lazzerini-Denchi, E.E. & Karlseder, J.J. A telomere-dependent DNA damage checkpoint induced by prolonged mitotic arrest. Nat. Struct. Mol. Biol. 19, 387–394 (2012).
Acknowledgements
We are indebted to H. Pickett (CMRI, Sydney, Australia) for generously sharing HT1080-hTR cells, reagents and for expert advice. We are grateful to Z. You (Washington University, St. Louis), J. Campbell (California Institute of Technology, Pasadena, California, USA), R. Everett (University of Glasgow), Z. Gurard-Levin (Institut Curie, Paris) and O. Fernandez-Capetillo (CNIO, Madrid, Spain) for sharing reagents. We thank the Salk Institute's J. Fitzpatrick of the Waitt Advanced Biophotonics Center (La Jolla, California, USA) for imaging assistance. R.J.O. is supported by the American Foundation for Aging Research (AFAR) and the Glenn Center for Research on Aging. N.A. is supported by a Human Frontier Science Program (HFSP) fellowship. D.H.L. and L.O. are supported by the Glenn Center for Research on Aging. G.A. is supported by La Ligue Nationale contre le Cancer (Equipe labellisée Ligue), PIC Programs, the European Commission Network of Excellence EpiGeneSys (HEALTH-F4-2010-257082), the European Commission ITN FP7-PEOPLE-2008-238176 “Nucleosome 4D,” ERC Advanced Grant 2009-AdG_20090506 “Eccentric,” the European Commission large-scale integrating project FP7_HEALTH-2010-259743 “MODHEP,” ANR “ChromaTin” ANR-10-BLAN-1326-03, ANR-11-LABX-0044_DEEP and ANR-10-IDEX-0001-02 PSL*, ANR “CHAPINHIB” ANR-12-BSV5-0022-02 and Aviesan-ITMO cancer project “Epigenomics of breast cancer.” J.K. is supported by the Salk Institute Cancer Center Core Grant (P30CA014195), the US National Institutes of Health (R01GM087476, R01CA174942), the Sabo Trust, the Fritz B. Burns Foundation, Philip Messinger and the Highland Street Foundation.
Author information
Authors and Affiliations
Contributions
R.J.O. designed and carried out the experiments and wrote the manuscript. N.A., L.O. and C.H. designed and carried out experiments. D.H.L. analyzed microarray data. A.C. provided essential reagents and contributed to the original concept of the project. G.A. provided essential reagents, contributed to the original concept of the project and provided additional support throughout. J.K. designed experiments, supervised the work and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Effects of ASF1 suppression.
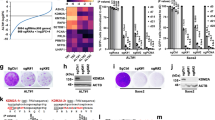
(a) Western blot analysis of ASF1 expression and RPA2 S4/S8 phosphorylation following transfection with control and ASF1 siRNAs in IMR90, WI38 and HeLa cells. γTubulin was used as loading control. (b) FACS analysis of control and ASF1 transfected Hela LT and IMR90-hTERT cells. Average percentage of cells in S-phase are indicated from 3 independent experiments. (c) Chromatin Immuno-precipitation (ChIP) of TRF2 and histone H3 at telomeres in control (c) and ASF1 (ASF1) siRNA treated HeLa cells. Numbers displayed beneath the panels are % telomeric DNA precipitated relative to the IgG control IP. (d) Micrococcal Nucleasae (MNase) digestion of chromatin from control siRNA and ASF1 siRNA treated HeLa LT cells. Increasing amounts of MNase (2, 4 and 8 units) were used in 10 min digestions. 5 μg of purified digested DNA was electrophoresed on a 1.2% TAE gel. (e) Telomere length and telomerase activity of HeLa 1.2.11, Hela LT, ST and VST cells. Telomerase activity was determined by TRAP assay using 1 μg of input soluble protein extract. (f) Quantification of FACS analysis of BrdU incorporation in control and ASF1 transfected Hela LT and IMR90 cells. Percentages of BrdU positive cells are shown and derived from 3 independent experiments.
Supplementary Figure 2 ASF1 suppression causes APB formation.
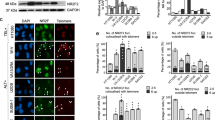
(a) Confocal IF-FISH imaging of RPA2 (Red), PML (Cyan), TTAGGG-FISH (Green) co-localization in control siRNA and ASF1 siRNA treated WI38-hTERT and WI38-hTERT-E6-E7. Images are maximum intensity projections of ∼10 stacks captured with a 63X objective lens. All panels include the merged channels with DAPI. (b) Confocal IF-FISH imaging of RPA2 (Red), BLM (Cyan), TTAGGG-FISH (Green) co-localization in control siRNA and ASF1 siRNA treated IMR90-hTERT and HeLa LT. Images are maximum intensity projections of ∼10 stacks captured with a 63X objective lens. All panels include the merged channels with DAPI and enlarged sections of the merge with DAPI. (c) Confocal-IF imaging of TRF2 (Green) and RPA2 (Red) co-localization in control siRNA and ASF1 siRNA treated IMR90-hTERT, IMR90-hTERT-E6-E7, HeLa LT, ST and VST cells. As in Fig 1A, TRF2/RPA2 co-localization in large foci in IMR90 hTERT, IMR90-hTERT-E6-E7 occurs in ∼5% of the cellular population. Otherwise, images of all control siRNA treated cells and ASF1 depleted HeLa ST represent the general cell population.
Supplementary Figure 3 ASF1 suppression leads to ECTR generation.
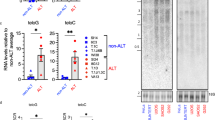
(a) C-circle assay in control and ASF1 depleted human primary IMR90/WI38-Empty Vector (EV), IMR90/WI38-E6-E7, IMR90/WI38-hTERT and IMR90/WI38-hTERT-E6-E7 cells. Note that the transformed IMR90 cells displayed here were derived from independent retroviral infections and differ from those shown in Fig. 2. Negative controls are reactions lacking φ29 and DNA. (b) Quantification of C-circles in control (black bar) and ASF1 depleted (light grey bar) cells shown in (a). The ALT positive U2OS control was used as reference for C-circles (dark grey bar), against which IMR90 and WI38 data was calculated. Data represent means ±SDs of at least 3 experiments. (c) T-circle assay in following ASF1 depletion. Increasing amounts (0.25, 0.5 and 1 μg) of digested genomic DNA from control and ASF1 depleted human primary IMR90, IMR90 E6-E7, IMR90-hTERT and IMR90-hTERT E6-E7 cells. Arrows indicate T-circle products. Negative controls are reactions lacking φ29.
Supplementary Figure 4 ASF1 suppression leads to C-circle formation.
(a) Confocal IF-FISH imaging of RPA2 (Cyan), PML (green), ssTTAGGG-FISH (Red) co-localization in control siRNA and ASF1 siRNA treated HeLa LT. Images are maximum intensity projections of ∼10 stacks captured with a 63X objective lens. All panels include the merged channels with and without DAPI. (b) As in (a) with the exception that the coverslips were denatured so as to allow detection of dsTTAGGG repeats.
Supplementary Figure 5 ASF1 suppression inhibits telomerase activity.
(a) Telomerase activity was determined by TRAP assay using 1 μg of input soluble protein extract. Lanes i-vi show: (i) No extract control (ii) DMSO added to extract (iii) 10 μM BIBR-1532 (TERTi) added to extract (iv) heat denatured control (v) RNase I treated extract (vi) positive control (mock treated HeLa LT extract). (b) Confocal IF-FISH imaging of RPA2 (Red), PML (Cyan), TTAGGG-FISH (Green) co-localization in ASF1 siRNA transfected HeLa LT and IMR90 hTERT cells treated with 10μM BIBR-1532 (TERTi). Images are maximum intensity projections of ∼10 stacks captured with a 63X objective lens. All panels include the merged channels with DAPI. (c) Microarray heatmap of differentially expressed genes in ASF1 depleted HeLa LT. Red indicates up-regulated genes and blue indicates down-regulated genes. Selected enriched functional pathways are shown. (d) Expression of secretory cytokines (IL1A, IL1B, IL6, IL8), activators of NFKB signaling (IKBKG (Nemo), NFKB2 (p100)), and the TGFβ pathway. Data represent means ±SDs of 3 independent experiments. (e) qPCR of gene expression of regulators of TERT expression. Gene expression is normalized to that of the RPLO (ribosomal protein L15) gene. Data represent means of 3 independent experiments in which each PCR was conducted in triplicate.
Supplementary Figure 6 Effects of ASF1 suppression.
(a) Western blot analysis of ASF1 and RPA2 expression and RPA2 S4/S8 phosphorylation following transfection with individual and mixed single siRNAs and Smartpool siRNAs against ASF1a and ASF1b in HeLa LT cells. Control siRNAs were also used in transfections. γTubulin was used as loading control. Black triangles above panels indicate 2X and 1X loading of whole cell extracts. (b) C-circle assays from siRNA transfected HeLa LT cells in (a). Negative controls are reactions lacking φ29 and DNA. Control siRNAs were also used in transfections. (c) Confocal-IF imaging of RPA2 (Red), PML (Cyan), TTAGGG-FISH (Green) co-localization in control siRNA and single ASF1 siRNA treated HeLa LT cells. Images are maximum intensity projections of ∼10 stacks captured with a 63X objective lens. All panels include the merged channels with DAPI and enlarged sections of the merge with DAPI. (d) Western analysis to confirm siRNA knockdown. The target gene is indicated on the left, adjacent to each panel. To the right of each panel, the S-phase index (%) at the time of harvest (72 hrs) of the transfections is indicated. (e) Standard IF of TRF2 (Green) and RPA2 (Red) localization after synchronization of HeLa LT cells in S-phase and chronic treatment with hydroxyurea (HU) (3 mM) and aphidicolin (APH) (5 μM) over 48 hrs. Images. All panels were captured using a 63X objective lens on a Zeiss Axioplan II microscope and include the merged channels with DAPI and enlarged sections of the merge with DAPI.
Supplementary Figure 7 ALT pathway analysis.
(a) Quantitation of S-phase index of shRNA siC/ASF1 siRNA transfected cells. Data represents the average total S-phase population in cell cultures at 72 hrs post siRNA transfection. (b) Western analysis of ASF1, phospho-RPA2 S4/S8, RPA2, γH2AX and H2AX. shRNA target is indicated above panels. γTUB is general loading control. (c) Confocal IF for RPA2 (red), PML (cyan) and TTAGGG FISH (green) in shRNA infected HeLa LT cells. From top to bottom the images are from shScramble and control siRNA, shScramble and ASF1 siRNA, BLM shRNA and ASF1 siRNA, WRN shRNA and ASF1 siRNA, EXO1 shRNA and ASF1 siRNA and RAD17 shRNA and ASF1 siRNA HeLa LT cells. Images shown are representative of those cells containing APB like structures that are observed as indicated in Fig. 4. For BLM shRNA and ASF1 siRNA where few APBs are observed the images shown are representative of the cell population. Note the absence of focal RPA2 formation in the absence of BLM. Images in are maximum intensity projections of ∼10 stacks captured with a 63X objective lens. All panels include the merged channels with DAPI and enlarged sections of the merge with DAPI.
Supplementary Figure 8 Original blots of westerns in main figures.
(a) Complete blots of westerns for ASF1, RPA2 S4S8 and γTubulin shown in Figure 5b. The molecular weight of the target protein is indicated. (b) Complete blots of westerns for ASF1, RPA2 S4S8, γH2AX and γTubulin shown in Figure 6b. The molecular weight of the target protein is indicated. (c) Complete blots of westerns confirming shRNA knockdown of factors shown in shown in Figure 7a. Top panel; left to right; PML, BLM, CtIP, EXO1, DNA2 and bottom panel; left to right; TRF2, RAD51, MRE11, RAD17 and WRN. The molecular weight of the target protein is indicated. The type and percent gradient of acrylamide in each gel is indicated beneath the image.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 17028 kb)
Rights and permissions
About this article
Cite this article
O'Sullivan, R., Arnoult, N., Lackner, D. et al. Rapid induction of alternative lengthening of telomeres by depletion of the histone chaperone ASF1. Nat Struct Mol Biol 21, 167–174 (2014). https://doi.org/10.1038/nsmb.2754
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2754
This article is cited by
-
Combining old and new concepts in targeting telomerase for cancer therapy: transient, immediate, complete and combinatory attack (TICCA)
Cancer Cell International (2023)
-
Alternative lengthening of telomeres (ALT) cells viability is dependent on C-rich telomeric RNAs
Nature Communications (2023)
-
Histone demethylase KDM2A is a selective vulnerability of cancers relying on alternative telomere maintenance
Nature Communications (2023)
-
A non-genetic switch triggers alternative telomere lengthening and cellular immortalization in ATRX deficient cells
Nature Communications (2023)
-
Telomere-to-mitochondria signalling by ZBP1 mediates replicative crisis
Nature (2023)