Abstract
Various technical tools have been developed to probe the functions of microRNAs (miRNAs), yet their application has been limited by low efficacy and specificity. To overcome the limitations, we used transcription activator–like effector nucleases (TALENs) to knock out human miRNA genes. We designed and produced a library of 540 pairs of TALENs for 274 miRNA loci, focusing on potentially important miRNAs. The knockout procedure takes only 2–4 weeks and can be applied to any cell type. As a case study, we generated knockout cells for two related miRNAs, miR-141 and miR-200c, which belong to the highly conserved miR-200 family. Interestingly, miR-141 and miR-200c, despite their overall similarity, suppress largely nonoverlapping groups of targets, thus suggesting that functional miRNA-target interaction requires strict seed-pairing. Our study illustrates the potency of TALEN technology and provides useful resources for miRNA research.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Bartel, D.P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Kim, V.N., Han, J. & Siomi, M.C. Biogenesis of small RNAs in animals. Nat. Rev. Mol. Cell Biol. 10, 126–139 (2009).
Kozomara, A. & Griffiths-Jones, S. miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res. 39, D152–D157 (2011).
Thomas, M., Lieberman, J. & Lal, A. Desperately seeking microRNA targets. Nat. Struct. Mol. Biol. 17, 1169–1174 (2010).
Khan, A.A. et al. Transfection of small RNAs globally perturbs gene regulation by endogenous microRNAs. Nat. Biotechnol. 27, 549–555 (2009).
Stenvang, J., Petri, A., Lindow, M., Obad, S. & Kauppinen, S. Inhibition of microRNA function by antimiR oligonucleotides. Silence 3, 1 (2012).
Ebert, M.S. & Sharp, P.A. MicroRNA sponges: progress and possibilities. RNA 16, 2043–2050 (2010).
Prosser, H.M., Koike-Yusa, H., Cooper, J.D., Law, F.C. & Bradley, A. A resource of vectors and ES cells for targeted deletion of microRNAs in mice. Nat. Biotechnol. 29, 840–845 (2011).
Park, C.Y. et al. A resource for the conditional ablation of microRNAs in the mouse. Cell Rep. 1, 385–391 (2012).
Bibikova, M., Beumer, K., Trautman, J.K. & Carroll, D. Enhancing gene targeting with designed zinc finger nucleases. Science 300, 764 (2003).
Urnov, F.D. et al. Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 435, 646–651 (2005).
Miller, J.C. et al. A TALE nuclease architecture for efficient genome editing. Nat. Biotechnol. 29, 143–148 (2011).
Cho, S.W., Kim, S., Kim, J.M. & Kim, J.S. Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat. Biotechnol. 31, 230–232 (2013).
Kim, H.J., Lee, H.J., Kim, H., Cho, S.W. & Kim, J.S. Targeted genome editing in human cells with zinc finger nucleases constructed via modular assembly. Genome Res. 19, 1279–1288 (2009).
Kim, S., Lee, M.J., Kim, H., Kang, M. & Kim, J.S. Preassembled zinc-finger arrays for rapid construction of ZFNs. Nat. Methods 8, 7 (2011).
Kim, Y. et al. A library of TAL effector nucleases spanning the human genome. Nat. Biotechnol. 31, 251–258 (2013).
Sung, Y.H. et al. Knockout mice created by TALEN-mediated gene targeting. Nat. Biotechnol. 31, 23–24 (2013).
Mojica, F.J., Diez-Villasenor, C., Garcia-Martinez, J. & Almendros, C. Short motif sequences determine the targets of the prokaryotic CRISPR defence system. Microbiology 155, 733–740 (2009).
Miyoshi, K., Miyoshi, T. & Siomi, H. Many ways to generate microRNA-like small RNAs: non-canonical pathways for microRNA production. Mol. Genet. Genomics 284, 95–103 (2010).
Chiang, H.R. et al. Mammalian microRNAs: experimental evaluation of novel and previously annotated genes. Genes Dev. 24, 992–1009 (2010).
Friedman, R.C., Farh, K.K.H., Burge, C.B. & Bartel, D.P. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 19, 92–105 (2009).
Han, J. et al. Molecular basis for the recognition of primary microRNAs by the Drosha-DGCR8 complex. Cell 125, 887–901 (2006).
Lewis, B.P., Burge, C.B. & Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005).
Krek, A. et al. Combinatorial microRNA target predictions. Nat. Genet. 37, 495–500 (2005).
Gregory, P.A., Bracken, C.P., Bert, A.G. & Goodall, G.J. MicroRNAs as regulators of epithelial-mesenchymal transition. Cell Cycle 7, 3112–3118 (2008).
Park, S.M., Gaur, A.B., Lengyel, E. & Peter, M.E. The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev. 22, 894–907 (2008).
Hyun, S. et al. Conserved microRNA miR-8/miR-200 and its target USH/FOG2 control growth by regulating PI3K. Cell 139, 1096–1108 (2009).
Burk, U. et al. A reciprocal repression between ZEB1 and members of the miR-200 family promotes EMT and invasion in cancer cells. EMBO Rep. 9, 582–589 (2008).
Kim, H. et al. Magnetic separation and antibiotics selection enable enrichment of cells with ZFN/TALEN-induced mutations. PLoS ONE 8, e56476 (2013).
Zeng, Y., Yi, R. & Cullen, B.R. Recognition and cleavage of primary microRNA precursors by the nuclear processing enzyme Drosha. EMBO J. 24, 138–148 (2005).
Hu, R., Wallace, J., Dahlem, T.J., Grunwald, D.J. & O'Connell, R.M. Targeting human microRNA genes using engineered Tal-effector nucleases (TALENs). PLoS ONE 8, e63074 (2013).
Elson-Schwab, I., Lorentzen, A. & Marshall, C.J. MicroRNA-200 family members differentially regulate morphological plasticity and mode of melanoma cell invasion. PLoS ONE 5, e13176 (2010).
Uhlmann, S. et al. miR-200bc/429 cluster targets PLCgγ1 and differentially regulates proliferation and EGF-driven invasion than miR-200a/141 in breast cancer. Oncogene 29, 4297–4306 (2010).
Doench, J.G. & Sharp, P.A. Specificity of microRNA target selection in translational repression. Genes Dev. 18, 504–511 (2004).
Brennecke, J., Stark, A., Russell, R.B. & Cohen, S.M. Principles of microRNA-target recognition. PLoS Biol. 3, e85 (2005).
Baek, D. et al. The impact of microRNAs on protein output. Nature 455, 64–71 (2008).
Didiano, D. & Hobert, O. Perfect seed pairing is not a generally reliable predictor for miRNA-target interactions. Nat. Struct. Mol. Biol. 13, 849–851 (2006).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Guo, J., Gaj, T. & Barbas, C.F. III. Directed evolution of an enhanced and highly efficient FokI cleavage domain for zinc finger nucleases. J. Mol. Biol. 400, 96–107 (2010).
Pall, G.S. & Hamilton, A.J. Improved northern blot method for enhanced detection of small RNA. Nat. Protoc. 3, 1077–1084 (2008).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Grimson, A. et al. MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol. Cell 27, 91–105 (2007).
Acknowledgements
We thank S. Kim (ToolGen) for the help with TALEN design and construction. We gratefully acknowledge S. Ryu and S. Bae for their technical help. We are grateful to members of our laboratories, particularly Y. Kim, for discussion and reading the manuscript. This work was supported by the Research Center Program of the Institute for Basic Science (EM1202) (Y.-K.K., J.P., D.B. and V.N.K.) and Basic Science Research Program of the National Research Foundation funded by the Ministry of Education, Science and Technology of Korea (20110014523) (J.K. and D.B.). J.-S.K. was supported by the National Research Foundation of Korea (2013000718).
Author information
Authors and Affiliations
Contributions
Y.-K.K., G.W., J.-S.K. and V.N.K. designed the project; Y.-K.K. and G.W. performed the experiments and analyzed the data; J.P., J.K. and D.B. performed the bioinformatics analysis; Y.-K.K. and V.N.K. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 T7E1 assay to test the efficacy of TALEN constructs.
To confirm the genome-editing activity of TALEN constructs, we transfected each TALEN pair into HEK293T (for all miRNAs except for mir-99b and mir-222) and K562 (for mir-99b and mir-222) cells, and carried out T7E1 assay using total genomic DNA. The asterisks indicate cleaved DNA produced by T7E1 enzyme that is specific to heteroduplex DNA caused by genome editing. The mutation frequency was calculated from the proportion of cut bands intensity to total bands intensity.
Supplementary Figure 2 TALEN-mediated knockout of miR-141 and miR-200c.
(a) Expression of the mir-200 family in human breast cell lines. miRNA level was measured by qRT-PCR and normalized against that of U6 snRNA. The genomic organization of the mir-200 family loci is shown. (b) Effective enrichment of mutated cells by using surrogate reporter system. The structure of surrogate reporter plasmid for the enrichment of mutated cells (upper panel). In the plasmid, a cassette of enhanced green fluorescent protein (eGFP) fused to the H-2Kk cell surface antigen is placed downstream of monomeric red fluorescent protein (mRFP) sequence out of frame (indicated with a stop sign). In between mRFP and eGFP sequences, the target site for the cotransfected TALEN construct is inserted. Therefore, if the transfected TALEN works effectively in cells, small indel is made at the target sequence randomly, and the eGFP-H-2Kk fusion protein is expressed when the mutation corrects the reading frame by chance. By magnetic separation procedure or FACS, those cells expressing H-2Kk antigen can be enriched. (Middle and lower panel.) With the reporter, TALEN constructs for mir-141 (TALEN L1R1) or mir-200c (TALEN L4R4) were transfected to generate mir-141 KO (middle left panel), mir-200c KO (middle right panel), or DKO cell line (lower panel), and T7E1 assay was carried out. Compared to the control (lane 1), the DNA from TALEN-transfected cells (lane 2) shows clear cut bands. After the magnetic separation, the proportion of mutated DNA is significantly increased in the enriched cells (lane 4) compared to that from the remaining cells (lane 3). Note that electroporation increased transfection efficiency in double KO experiment (lower panel). (c) T7E1 assay to check mutations at potential off-target sites. Potential off-target sites were selected based on the sequence similarity to the TALEN target sequences (Supplementary Tables 5, 6, 7). Using the genomic DNA from reporter-enriched cells (upper panel) or KO cell colonies (lower panel), T7E1 assay was carried out against each site. Note that none of the samples shows cut band. (d) T7E1 assay to detect DNA mutation. After the enrichment of mutant cells, single cells were separated and grown into cell colonies, and genomic DNA was extracted (mir-141 KO clones in upper panel, mir-200c KO clones in middle panel, and DKO clones in lower panel). The DNA fragment encompassing the targeted region was amplified by PCR, treated with T7E1, and separated by gel electrophoresis. (e) Fluorescent PCR analysis to detect KO clones. Among the mutant clones which were confirmed by T7E1 assay in d, we selected some of them to test whether mutation occurred at all alleles by fluorescent PCR. The results from the cell lines used in this study are shown. Note that, compared to the size of PCR product from parental cells, increased or decreased size of PCR products were detected from mutant cells. (f) Summary of knockout procedure. The experimental procedure for the production of mir-141 and mir-200c KO clones is summarized. For each step, the efficiency/frequency of DNA mutation was monitored by the method indicated in the first column. The values in the first two rows indicate overall efficiency of DNA mutation in the cell population from b. In the following three rows, the frequency of the mutated cell colonies out of all colonies tested are shown (indicated in the parenthesis) (Fig. 2c and Supplementary Fig. 2d, e). Note that T7E1 assay detects the clones with mutation in at least one allele whereas F-PCR is used for the selection of clones with all alleles mutated.
Supplementary Figure 3 The relative levels of miRNAs in KO cells.
(a) The expression levels of miRNAs were measured by qRT-PCR, normalized against that of U6 snRNA, and normalized again to the miRNA level from parental cells. Data are presented as mean ± s.e. of biological replicates (n=3). (b) The expression levels of miRNAs were measured by qRT-PCR, and normalized against that of U6 snRNA. The same data was used to calculate the relative level of miRNAs in KO cells compared to the level from parental cells (Fig. 3c). For those samples that the miRNA expression level is below detection limit, asterisks are indicated. Data are presented as mean ± s.e. of biological replicates (n=4) for miR-141 and miR-200c or as mean ± range for the other miRNAs (n=2).
Supplementary Figure 4 Cellular and molecular impact of miRNA KO.
(a) Cell cycle analysis of KO cells. The same number (250,000 cells) of exponentially growing parental and KO cells were seeded on a 35 mm dish. Two days later, floating and attached cells were collected together, and cell cycle distribution was profiled using FACSCalibur (BD Biosciences) and analyzed with Flowing Software 2 (http://www.flowingsoftware.com/). The average of percentages from two different trials is shown. (b) Median fold change of target gene groups with miR-141 or miR-200c motif. To calculate the median fold change of target gene groups with miR-141 or miR-200c motif, we collected the transcripts which have single 7-mer-m8 motif in their 3'-UTR sequences. The median value of fold changes was normalized by that of transcripts which do not have the 6-mer motif of corresponding miRNA in their entire sequence. P value was calculated by Wilcoxon rank-sum test. (c) The miRNA recognition elements in the 3' UTRs of PPT2 and FOG2. Luciferase constructs harboring the 3' UTR of PPT2 (upper panel) and FOG2 (lower panel) are shown. The target sites of miRNAs were predicted by TargetScan (http://www.targetscan.org). The base-pairing structures between miRNA and target mRNA sequences were predicted from mfold web server (http://mfold.rna.albany.edu/?q=mfold). For each pair, mRNA and miRNA sequences are presented with blue and red letters, respectively. The sequences in seed match region are indicated by underline. To create mutant reporter constructs, four nucleotides in the rectangle were changed into those indicated with arrows. (d) Phylogenetic distribution of mir-200 family. The mir-200 family miRNA sequences were collected from miRBase release 19. The mir-200 family miRNAs are grouped into two subfamilies based on the seed sequence. For each taxon, representative species was selected. The presence of miRNAs in each subfamily is indicated with filled circles. Note that the mir-141, -200a subfamily is found only in chordates. (e) Sequence alignment of mir-200 family miRNAs. Mature sequences of miR-8 from D. melanogaster, miR-236 from C. elegans, and miR-141, -200a, -200b, -200c, and -429 from H. sapiens, were used for the alignment.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Tables 4–8 (PDF 3321 kb)
Supplementary Table 1
Sequencing libraries used for miRNA selection. (XLSX 14 kb)
Supplementary Table 2
List of miRNAs for TALEN library. (XLSX 28 kb)
Supplementary Table 3
Information of TALEN constructs for miRNA KO. (XLSX 44 kb)
Rights and permissions
About this article
Cite this article
Kim, YK., Wee, G., Park, J. et al. TALEN-based knockout library for human microRNAs. Nat Struct Mol Biol 20, 1458–1464 (2013). https://doi.org/10.1038/nsmb.2701
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2701
This article is cited by
-
Diverse roles of noncoding RNAs in vascular calcification
Archives of Pharmacal Research (2019)
-
A proteomic analysis of an in vitro knock-out of miR-200c
Scientific Reports (2018)
-
MicroRNA Metabolism and Dysregulation in Amyotrophic Lateral Sclerosis
Molecular Neurobiology (2018)
-
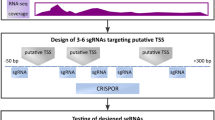
Precise mapping of the transcription start sites of human microRNAs using DROSHA knockout cells
BMC Genomics (2016)
-
CRISPR/cas9, a novel genomic tool to knock down microRNA in vitro and in vivo
Scientific Reports (2016)