Abstract
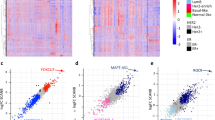
Despite growing appreciation of the importance of long noncoding RNAs (lncRNAs) in normal physiology and disease, our knowledge of cancer-related lncRNAs remains limited. By repurposing microarray probes, we constructed expression profiles of 10,207 lncRNA genes in approximately 1,300 tumors over four different cancer types. Through integrative analysis of the lncRNA expression profiles with clinical outcome and somatic copy-number alterations, we identified lncRNAs that are associated with cancer subtypes and clinical prognosis and predicted those that are potential drivers of cancer progression. We validated our predictions by experimentally confirming prostate cancer cell growth dependence on two newly identified lncRNAs. Our analysis provides a resource of clinically relevant lncRNAs for the development of lncRNA biomarkers and the identification of lncRNA therapeutic targets. It also demonstrates the power of integrating publically available genomic data sets and clinical information for discovering disease-associated lncRNAs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Accessions
Ensembl
Gene Expression Omnibus
References
Ota, T. et al. Complete sequencing and characterization of 21,243 full-length human cDNAs. Nat. Genet. 36, 40–45 (2004).
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009).
Khalil, A.M. et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc. Natl. Acad. Sci. USA 106, 11667–11672 (2009).
Guttman, M. et al. Ab initio reconstruction of cell type-specific transcriptomes in mouse reveals the conserved multi-exonic structure of lincRNAs. Nat. Biotechnol. 28, 503–510 (2010).
Cabili, M.N. et al. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev. 25, 1915–1927 (2011).
Prensner, J.R. & Chinnaiyan, A.M. The emergence of lncRNAs in cancer biology. Cancer Discov. 1, 391–407 (2011).
Wapinski, O. & Chang, H.Y. Long noncoding RNAs and human disease. Trends Cell Biol. 21, 354–361 (2011).
Lee, G.L., Dobi, A. & Srivastava, S. Prostate cancer: diagnostic performance of the PCA3 urine test. Nat. Rev. Urol. 8, 123–124 (2011).
Liao, Q. et al. Large-scale prediction of long non-coding RNA functions in a coding–non-coding gene co-expression network. Nucleic Acids Res. 39, 3864–3878 (2011).
Mercer, T.R., Dinger, M.E., Sunkin, S.M., Mehler, M.F. & Mattick, J.S. Specific expression of long noncoding RNAs in the mouse brain. Proc. Natl. Acad. Sci. USA 105, 716–721 (2008).
Michelhaugh, S.K. et al. Mining Affymetrix microarray data for long non-coding RNAs: altered expression in the nucleus accumbens of heroin abusers. J. Neurochem. 116, 459–466 (2011).
Gellert, P., Ponomareva, Y., Braun, T. & Uchida, S. Noncoder: a web interface for exon array-based detection of long non-coding RNAs. Nucleic Acids Res. 41, e20 (2013).
Johnson, R. Long non-coding RNAs in Huntington's disease neurodegeneration. Neurobiol. Dis. 46, 245–254 (2012).
Zhang, X. et al. Long non-coding RNA expression profiles predict clinical phenotypes in glioma. Neurobiol. Dis. 48, 1–8 (2012).
Raghavachari, N. et al. A systematic comparison and evaluation of high density exon arrays and RNA-seq technology used to unravel the peripheral blood transcriptome of sickle cell disease. BMC Med. Genomics 5, 28 (2012).
Xu, W. et al. Human transcriptome array for high-throughput clinical studies. Proc. Natl. Acad. Sci. USA 108, 3707–3712 (2011).
Levin, J.Z. et al. Comprehensive comparative analysis of strand-specific RNA sequencing methods. Nat. Methods 7, 709–715 (2010).
Taylor, B.S. et al. Integrative genomic profiling of human prostate cancer. Cancer Cell 18, 11–22 (2010).
The Cancer Genome Atlas Research Network. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455, 1061–1068 (2008).
Derrien, T. et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 22, 1775–1789 (2012).
Kapur, K., Xing, Y., Ouyang, Z. & Wong, W.H. Exon arrays provide accurate assessments of gene expression. Genome Biol. 8, R82 (2007).
Prensner, J.R. et al. Transcriptome sequencing across a prostate cancer cohort identifies PCAT-1, an unannotated lincRNA implicated in disease progression. Nat. Biotechnol. 29, 742–749 (2011).
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: a revolutionary tool for transcriptomics. Nat. Rev. Genet. 10, 57–63 (2009).
Petrovics, G. et al. Elevated expression of PCGEM1, a prostate-specific gene with cell growth-promoting function, is associated with high-risk prostate cancer patients. Oncogene 23, 605–611 (2004).
Mourtada-Maarabouni, M., Pickard, M.R., Hedge, V.L., Farzaneh, F. & Williams, G.T. GAS5, a non-protein-coding RNA, controls apoptosis and is downregulated in breast cancer. Oncogene 28, 195–208 (2009).
Clemson, C.M. et al. An architectural role for a nuclear noncoding RNA: NEAT1 RNA is essential for the structure of paraspeckles. Mol. Cell 33, 717–726 (2009).
Kretz, M. et al. Suppression of progenitor differentiation requires the long noncoding RNA ANCR. Genes Dev. 26, 338–343 (2012).
Wang, K.C. et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature 472, 120–124 (2011).
Szegedi, K. et al. The anti-apoptotic protein G1P3 is overexpressed in psoriasis and regulated by the non-coding RNA, PRINS. Exp. Dermatol. 19, 269–278 (2010).
Wagner, L.A. et al. EGO, a novel, noncoding RNA gene, regulates eosinophil granule protein transcript expression. Blood 109, 5191–5198 (2007).
The Cancer Genome Atlas Research Network. Integrated genomic analyses of ovarian carcinoma. Nature 474, 609–615 (2011).
Hammerman, P.S. et al. Comprehensive genomic characterization of squamous cell lung cancers. Nature 489, 519–525 (2012).
Ishii, N. et al. Identification of a novel non-coding RNA, MIAT, that confers risk of myocardial infarction. J. Hum. Genet. 51, 1087–1099 (2006).
Rapicavoli, N.A., Poth, E.M. & Blackshaw, S. The long noncoding RNA RNCR2 directs mouse retinal cell specification. BMC Dev. Biol. 10, 49 (2010).
Chan, A.S., Thorner, P.S., Squire, J.A. & Zielenska, M. Identification of a novel gene NCRMS on chromosome 12q21 with differential expression between rhabdomyosarcoma subtypes. Oncogene 21, 3029–3037 (2002).
Gupta, R.A. et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 464, 1071–1076 (2010).
Rinn, J.L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007).
Kogo, R. et al. Long noncoding RNA HOTAIR regulates polycomb-dependent chromatin modification and is associated with poor prognosis in colorectal cancers. Cancer Res. 71, 6320–6326 (2011).
Beroukhim, R. et al. The landscape of somatic copy-number alteration across human cancers. Nature 463, 899–905 (2010).
Garraway, L.A. et al. Integrative genomic analyses identify MITF as a lineage survival oncogene amplified in malignant melanoma. Nature 436, 117–122 (2005).
Akavia, U.D. et al. An integrated approach to uncover drivers of cancer. Cell 143, 1005–1017 (2010).
Tran, V.G. et al. H19 antisense RNA can up-regulate Igf2 transcription by activation of a novel promoter in mouse myoblasts. PLoS ONE 7, e37923 (2012).
Califano, A., Butte, A.J., Friend, S., Ideker, T. & Schadt, E. Leveraging models of cell regulation and GWAS data in integrative network-based association studies. Nat. Genet. 44, 841–847 (2012).
Pe'er, D. & Hacohen, N. Principles and strategies for developing network models in cancer. Cell 144, 864–873 (2011).
Zhao, J. et al. Genome-wide identification of polycomb-associated RNAs by RIP-seq. Mol. Cell 40, 939–953 (2010).
Syvänen, A.C. Accessing genetic variation: genotyping single nucleotide polymorphisms. Nat. Rev. Genet. 2, 930–942 (2001).
Meyerson, M., Gabriel, S. & Getz, G. Advances in understanding cancer genomes through second-generation sequencing. Nat. Rev. Genet. 11, 685–696 (2010).
Flicek, P. et al. Ensembl 2012. Nucleic Acids Res. 40, D84–D90 (2012).
Jiang, H. & Wong, W.H. SeqMap: mapping massive amount of oligonucleotides to the genome. Bioinformatics 24, 2395–2396 (2008).
Kuhn, R.M., Haussler, D. & Kent, W.J. The UCSC genome browser and associated tools. Brief. Bioinform. 14, 144–161 (2013).
Seok, J., Xu, W., Gao, H., Davis, R.W. & Xiao, W. JETTA: junction and exon toolkits for transcriptome analysis. Bioinformatics 28, 1274–1275 (2012).
Johnson, W.E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Beroukhim, R. et al. Assessing the significance of chromosomal aberrations in cancer: methodology and application to glioma. Proc. Natl. Acad. Sci. USA 104, 20007–20012 (2007).
Mermel, C.H. et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 12, R41 (2011).
Taylor, B.S. et al. Functional copy-number alterations in cancer. PLoS ONE 3, e3179 (2008).
Lin, M.F., Jungreis, I. & Kellis, M. PhyloCSF: a comparative genomics method to distinguish protein coding and non-coding regions. Bioinformatica 27, 275–282 (2011).
Lindblad-Toh, K. et al. A high-resolution map of human evolutionary constraint using 29 mammals. Nature 478, 476–482 (2011).
Acknowledgements
This work was partially funded by the National Natural Science Foundation of China (31028011) (X.S.L.), the National Basic Research (973) Program of China (2010CB944904; Y.Z.) and US National Institutes of Health grant GM099409 (X.S.L.).
Author information
Authors and Affiliations
Contributions
Y.C. conceived the project. Z.D. and Y.C. designed the algorithms and performed computational analyses. R.G.W.V. contributed to the subtype analyses of ovarian cancer. T.F. performed all the experimental validation. Z.D., T.F., Z.S., Y.Z., M.B., Y.C. and X.S.L. participated in the discussions and contributed to the analysis of the intermediate results throughout the project. Y.C., M.B. and X.S.L. supervised the project. Z.D., T.F., Y.C. and X.S.L. wrote the manuscript with the help from other coauthors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Tables 1–5 (PDF 661 kb)
Rights and permissions
About this article
Cite this article
Du, Z., Fei, T., Verhaak, R. et al. Integrative genomic analyses reveal clinically relevant long noncoding RNAs in human cancer. Nat Struct Mol Biol 20, 908–913 (2013). https://doi.org/10.1038/nsmb.2591
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2591
This article is cited by
-
Modeling CRISPR-Cas13d on-target and off-target effects using machine learning approaches
Nature Communications (2023)
-
Intrinsic targeting of host RNA by Cas13 constrains its utility
Nature Biomedical Engineering (2023)
-
LncRNA PCAT6 promotes proliferation, migration, invasion, and epithelial-mesenchymal transition of lung adenocarcinoma cell by targeting miR-545-3p
Molecular Biology Reports (2023)
-
LncRNA weighted gene co-expression network analysis reveals novel biomarkers related to prostate cancer metastasis
BMC Medical Genomics (2022)
-
CircSCAF8 promotes growth and metastasis of prostate cancer through the circSCAF8-miR-140-3p/miR-335-LIF pathway
Cell Death & Disease (2022)