Abstract
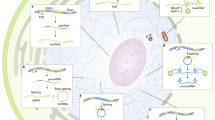
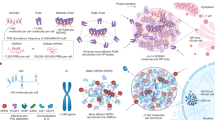
Genomes of complex organisms encode an abundance and diversity of long noncoding RNAs (lncRNAs) that are expressed throughout the cell and fulfill a wide variety of regulatory roles at almost every stage of gene expression. These roles, which encompass sensory, guiding, scaffolding and allosteric capacities, derive from folded modular domains in lncRNAs. In this diverse functional repertoire, we focus on the well-characterized ability for lncRNAs to function as epigenetic modulators. Many lncRNAs bind to chromatin-modifying proteins and recruit their catalytic activity to specific sites in the genome, thereby modulating chromatin states and impacting gene expression. Considering this regulatory potential in combination with the abundance of lncRNAs suggests that lncRNAs may be part of a broad epigenetic regulatory network.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Carninci, P. et al. The transcriptional landscape of the mammalian genome. Science 309, 1559–1563 (2005). One of the earliest analyses of large-scale cDNA sequencing, this work reveals the abundance of lncRNAs and complexity of transcriptional organization in the eukaryotic genome.
Mattick, J.S. A new paradigm for developmental biology. J. Exp. Biol. 210, 1526–1547 (2007).
Amaral, P.P., Clark, M.B., Gascoigne, D.K., Dinger, M.E. & Mattick, J.S. lncRNAdb: a reference database for long noncoding RNAs. Nucleic Acids Res. 39, D146–D151 (2011).
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009).
Kapranov, P. et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science 316, 1484–1488 (2007).
Dieci, G., Fiorino, G., Castelnuovo, M., Teichmann, M. & Pagano, A. The expanding RNA polymerase III transcriptome. Trends Genet. 23, 614–622 (2007).
Yin, Q.F. et al. Long noncoding RNAs with snoRNA ends. Mol. Cell 48, 219–230 (2012).
Dinger, M.E., Pang, K.C., Mercer, T.R. & Mattick, J.S. Differentiating protein-coding and noncoding RNA: challenges and ambiguities. PLOS Comput. Biol. 4, e1000176 (2008).
Lin, M.F., Jungreis, I. & Kellis, M. PhyloCSF: a comparative genomics method to distinguish protein coding and non-coding regions. Bioinformatics 27, i275–i282 (2011).
Galindo, M.I., Pueyo, J.I., Fouix, S., Bishop, S.A. & Couso, J.P. Peptides encoded by short ORFs control development and define a new eukaryotic gene family. PLoS Biol. 5, e106 (2007).
Ingolia, N.T., Lareau, L.F. & Weissman, J.S. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147, 789–802 (2011).
Banfai, B. et al. Long noncoding RNAs are rarely translated in two human cell lines. Genome Res. 22, 1646–1657 (2012).
Dinger, M.E., Gascoigne, D.K. & Mattick, J.S. The evolution of RNAs with multiple functions. Biochimie 93, 2013–2018 (2011).
Poliseno, L. et al. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature 465, 1033–1038 (2010).
Carvunis, A.R. et al. Proto-genes and de novo gene birth. Nature 487, 370–374 (2012).
Zheng, D. et al. Pseudogenes in the ENCODE regions: consensus annotation, analysis of transcription, and evolution. Genome Res. 17, 839–851 (2007).
Duret, L., Chureau, C., Samain, S., Weissenbach, J. & Avner, P. The Xist RNA gene evolved in eutherians by pseudogenization of a protein-coding gene. Science 312, 1653–1655 (2006).
Lee, J.T. The X as model for RNA's niche in epigenomic regulation. Cold Spring Harb. Perspect. Biol. 2, a003749 (2010).
Gerstein, M.B. et al. What is a gene, post-ENCODE? History and updated definition. Genome Res. 17, 669–681 (2007).
Denoeud, F. et al. Prominent use of distal 5′ transcription start sites and discovery of a large number of additional exons in ENCODE regions. Genome Res. 17, 746–759 (2007).
Djebali, S., Davis, C.A., LaGarde, J. & Gingeras, T. Landscape of transcription in human cell lines. Nature 489, 101–108 (2012).
Mazumder, B., Seshadri, V. & Fox, P.L. Translational control by the 3′-UTR: the ends specify the means. Trends Biochem. Sci. 28, 91–98 (2003).
Mercer, T.R. et al. Expression of distinct RNAs from 3′ untranslated regions. Nucleic Acids Res. 39, 2393–2403 (2011).
Okazaki, Y. et al. Analysis of the mouse transcriptome based on functional annotation of 60,770 full-length cDNAs. Nature 420, 563–573 (2002).
Rinn, J.L. et al. The transcriptional activity of human chromosome 22. Genes Dev. 17, 529–540 (2003).
Derrien, T. et al. The GENCODE v7 catalog of human long noncoding RNAs: Analysis of their gene structure, evolution, and expression. Genome Res. 22, 1775–1789 (2012).
Cabili, M.N. et al. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev. 25, 1915–1927 (2011).
Mercer, T.R. et al. Targeted RNA sequencing reveals the deep complexity of the human transcriptome. Nat. Biotechnol. 30, 99–104 (2012).
Kapranov, P., Willingham, A.T. & Gingeras, T.R. Genome-wide transcription and the implications for genomic organization. Nat. Rev. Genet. 8, 413–423 (2007).
Kapranov, P. et al. Examples of the complex architecture of the human transcriptome revealed by RACE and high-density tiling arrays. Genome Res. 15, 987–997 (2005). This systematic study reveals the complex organization of gene loci that is a common feature within the genome's modular architecture.
Katayama, S. et al. Antisense transcription in the mammalian transcriptome. Science 309, 1564–1566 (2005).
Ohhata, T., Hoki, Y., Sasaki, H. & Sado, T. Crucial role of antisense transcription across the Xist promoter in Tsix-mediated Xist chromatin modification. Development 135, 227–235 (2008).
Sun, B.K., Deaton, A.M. & Lee, J.T. A transient heterochromatic state in Xist preempts X inactivation choice without RNA stabilization. Mol. Cell 21, 617–628 (2006).
Zhao, J., Sun, B.K., Erwin, J.A., Song, J.J. & Lee, J.T. Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 322, 750–756 (2008).
Sado, T., Okano, M., Li, E. & Sasaki, H. De novo DNA methylation is dispensable for the initiation and propagation of X chromosome inactivation. Development 131, 975–982 (2004).
Ogawa, Y., Sun, B.K. & Lee, J.T. Intersection of the RNA interference and X-inactivation pathways. Science 320, 1336–1341 (2008).
Lee, J.T. Regulation of X-chromosome counting by Tsix and Xite sequences. Science 309, 768–771 (2005).
Cantara, W.A. et al. The RNA Modification Database, RNAMDB: 2011 update. Nucleic Acids Res. 39, D195–D201 (2011).
Helm, M. Post-transcriptional nucleotide modification and alternative folding of RNA. Nucleic Acids Res. 34, 721–733 (2006).
Kellner, S., Burhenne, J. & Helm, M. Detection of RNA modifications. RNA Biol. 7, 237–247 (2010).
Squires, J.E. et al. Widespread occurrence of 5-methylcytosine in human coding and non-coding RNA. Nucleic Acids Res. 40, 5023–5033 (2012).
Jia, G. et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat. Chem. Biol. 7, 885–887 (2011).
He, C. Grand challenge commentary: RNA epigenetics? Nat. Chem. Biol. 6, 863–865 (2010).
Cruz, J.A. & Westhof, E. The dynamic landscapes of RNA architecture. Cell 136, 604–609 (2009).
Lescoute, A. & Westhof, E. The interaction networks of structured RNAs. Nucleic Acids Res. 34, 6587–6604 (2006).
Talkington, M.W., Siuzdak, G. & Williamson, J.R. An assembly landscape for the 30S ribosomal subunit. Nature 438, 628–632 (2005).
Zhang, X. et al. Maternally expressed gene 3 (MEG3) noncoding ribonucleic acid: isoform structure, expression, and functions. Endocrinology 151, 939–947 (2010).
Steitz, T.A. A structural understanding of the dynamic ribosome machine. Nat. Rev. Mol. Cell Biol. 9, 242–253 (2008).
Novikova, I.V., Hennelly, S.P. & Sanbonmatsu, K.Y. Structural architecture of the human long non-coding RNA, steroid receptor RNA activator. Nucleic Acids Res. 40, 5034–5051 (2012).
Underwood, J.G. et al. FragSeq: transcriptome-wide RNA structure probing using high-throughput sequencing. Nat. Methods 7, 995–1001 (2010).
Kertesz, M. et al. Genome-wide measurement of RNA secondary structure in yeast. Nature 467, 103–107 (2010).
Liang, J.C., Bloom, R.J. & Smolke, C.D. Engineering biological systems with synthetic RNA molecules. Mol. Cell 43, 915–926 (2011).
Win, M.N. & Smolke, C.D. Higher-order cellular information processing with synthetic RNA devices. Science 322, 456–460 (2008).
Carrieri, C. et al. Long non-coding antisense RNA controls Uchl1 translation through an embedded SINEB2 repeat. Nature 491, 454–457 (2012).
Cisse, I.I., Kim, H. & Ha, T. A rule of seven in Watson-Crick base-pairing of mismatched sequences. Nat. Struct. Mol. Biol. 19, 623–627 (2012).
Affymetrix ENCODE Transcriptome Project & Cold Spring Harbor Laboratory ENCODE Transcriptome Project. Post-transcriptional processing generates a diversity of 5′-modified long and short RNAs. Nature 457, 1028–1032 (2009).
Wilusz, J.E., Freier, S.M. & Spector, D.L. 3′ end processing of a long nuclear-retained noncoding RNA yields a tRNA-like cytoplasmic RNA. Cell 135, 919–932 (2008).
Djupedal, I. & Ekwall, K. Epigenetics: heterochromatin meets RNAi. Cell Res. 19, 282–295 (2009).
Talini, G., Branciamore, S. & Gallori, E. Ribozymes: flexible molecular devices at work. Biochimie 93, 1998–2005 (2011).
Hogan, D.J., Riordan, D.P., Gerber, A.P., Herschlag, D. & Brown, P.O. Diverse RNA-binding proteins interact with functionally related sets of RNAs, suggesting an extensive regulatory system. PLoS Biol. 6, e255 (2008).
Lunde, B.M., Moore, C. & Varani, G. RNA-binding proteins: modular design for efficient function. Nat. Rev. Mol. Cell Biol. 8, 479–490 (2007).
Castello, A. et al. Insights into RNA biology from an atlas of mammalian mRNA-binding proteins. Cell 149, 1393–1406 (2012).
Ciesla, J. Metabolic enzymes that bind RNA: yet another level of cellular regulatory network? Acta Biochim. Pol. 53, 11–32 (2006).
Baltz, A.G. et al. The mRNA-bound proteome and its global occupancy profile on protein-coding transcripts. Mol. Cell 46, 674–690 (2012).
Scheibe, M., Butter, F., Hafner, M., Tuschl, T. & Mann, M. Quantitative mass spectrometry and PAR-CLIP to identify RNA-protein interactions. Nucleic Acids Res. 40, 9897–9902 (2012).
Reiter, N.J., Chan, C.W. & Mondragon, A. Emerging structural themes in large RNA molecules. Curr. Opin. Struct. Biol. 21, 319–326 (2011).
Buske, F.A., Mattick, J.S. & Bailey, T.L. Potential in vivo roles of nucleic acid triple-helices. RNA Biol. 8, 427–439 (2011).
Aguilera, A. & Garcia-Muse, T. R loops: from transcription byproducts to threats to genome stability. Mol. Cell 46, 115–124 (2012).
Schmitz, K.M., Mayer, C., Postepska, A. & Grummt, I. Interaction of noncoding RNA with the rDNA promoter mediates recruitment of DNMT3b and silencing of rRNA genes. Genes Dev. 24, 2264–2269 (2010).
Chu, C., Qu, K., Zhong, F.L., Artandi, S.E. & Chang, H.Y. Genomic maps of long noncoding RNA occupancy reveal principles of RNA-chromatin interactions. Mol. Cell 44, 667–678 (2011).
Jeon, Y. & Lee, J.T. YY1 tethers Xist RNA to the inactive X nucleation center. Cell 146, 119–133 (2011).
Ray, P.S. et al. A stress-responsive RNA switch regulates VEGFA expression. Nature 457, 915–919 (2009).
Delebecque, C.J., Lindner, A.B., Silver, P.A. & Aldaye, F.A. Organization of intracellular reactions with rationally designed RNA assemblies. Science 333, 470–474 (2011).
Yap, K.L. et al. Molecular interplay of the noncoding RNA ANRIL and methylated histone H3 lysine 27 by polycomb CBX7 in transcriptional silencing of INK4a. Mol. Cell 38, 662–674 (2010).
Guttman, M. et al. lincRNAs act in the circuitry controlling pluripotency and differentiation. Nature 477, 295–300 (2011).
Koziol, M.J. & Rinn, J.L. RNA traffic control of chromatin complexes. Curr. Opin. Genet. Dev. 20, 142–148 (2010).
Pandey, R.R. et al. Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol. Cell 32, 232–246 (2008).
Tsai, M.C. et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science 329, 689–693 (2010).
Plath, K. et al. Role of histone H3 lysine 27 methylation in X inactivation. Science 300, 131–135 (2003). This work shows the recruitment of the Eed–Ezh2 complex to mediate H3K27 methylation at the inactive X chromosome by Xist.
Wutz, A., Rasmussen, T.P. & Jaenisch, R. Chromosomal silencing and localization are mediated by different domains of Xist RNA. Nat. Genet. 30, 167–174 (2002).
Hasegawa, Y., Brockdorff, N., Kawano, S., Tsutui, K. & Nakagawa, S. The matrix protein hnRNP U is required for chromosomal localization of Xist RNA. Dev. Cell 19, 469–476 (2010).
Grant, J. et al. Rsx is a metatherian RNA with Xist-like properties in X-chromosome inactivation. Nature 487, 254–258 (2012).
Gong, C. & Maquat, L.E. lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3′ UTRs via Alu elements. Nature 470, 284–288 (2011).
Mariner, P.D. et al. Human Alu RNA is a modular transacting repressor of mRNA transcription during heat shock. Mol. Cell 29, 499–509 (2008).
Khalil, A.M. et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc. Natl. Acad. Sci. USA 106, 11667–11672 (2009).
Sanchez-Elsner, T., Gou, D., Kremmer, E. & Sauer, F. Noncoding RNAs of Trithorax response elements recruit Drosophila Ash1 to Ultrabithorax. Science 311, 1118–1123 (2006). A model for the local recruitment of the epigenetic regulator Ash1 by lncRNAs is proposed.
Rinn, J.L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007). This work is a demonstration of a lncRNA, HOTAIR, that targets Polycomb proteins in trans to the HOXD loci sites in the human genome.
Mohammad, F., Mondal, T. & Kanduri, C. Epigenetics of imprinted long noncoding RNAs. Epigenetics 4, 277–286 (2009).
Nagano, T. et al. The Air noncoding RNA epigenetically silences transcription by targeting G9a to chromatin. Science 322, 1717–1720 (2008).
Engstrom, P.G. et al. Complex Loci in human and mouse genomes. PLoS Genet. 2, e47 (2006).
Pinter, S.F. et al. Spreading of X chromosome inactivation via a hierarchy of defined Polycomb stations. Genome Res. 22, 1864–1876 (2012).
Wutz, A. Gene silencing in X-chromosome inactivation: advances in understanding facultative heterochromatin formation. Nat. Rev. Genet. 12, 542–553 (2011).
Almouzni, G. & Probst, A.V. Heterochromatin maintenance and establishment: lessons from the mouse pericentromere. Nucleus 2, 332–338 (2011).
Maison, C. et al. SUMOylation promotes de novo targeting of HP1alpha to pericentric heterochromatin. Nat. Genet. 43, 220–227 (2011).
Wang, K.C. et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature 472, 120–124 (2011).
Tripathi, V. et al. The nuclear-retained noncoding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol. Cell 39, 925–938 (2010).
Mao, Y.S., Sunwoo, H., Zhang, B. & Spector, D.L. Direct visualization of the co-transcriptional assembly of a nuclear body by noncoding RNAs. Nat. Cell Biol. 13, 95–101 (2011).
Shevtsov, S.P. & Dundr, M. Nucleation of nuclear bodies by RNA. Nat. Cell Biol. 13, 167–173 (2011).
Bantignies, F. et al. Polycomb-dependent regulatory contacts between distant Hox loci in Drosophila. Cell 144, 214–226 (2011).
Lanzuolo, C., Roure, V., Dekker, J., Bantignies, F. & Orlando, V. Polycomb response elements mediate the formation of chromosome higher-order structures in the bithorax complex. Nat. Cell Biol. 9, 1167–1174 (2007).
Yang, L. et al. ncRNA- and Pc2 methylation-dependent gene relocation between nuclear structures mediates gene activation programs. Cell 147, 773–788 (2011).
Barabasi, A.L. & Oltvai, Z.N. Network biology: understanding the cell′s functional organization. Nat. Rev. Genet. 5, 101–113 (2004).
Mattick, J.S. The genetic signatures of noncoding RNAs. PLoS Genet. 5, e1000459 (2009).
Schorderet, P. & Duboule, D. Structural and functional differences in the long non-coding RNA hotair in mouse and human. PLoS Genet. 7, e1002071 (2011).
Zhang, B. et al. The lncRNA Malat1 is dispensable for mouse development but its transcription plays a cis-regulatory role in the adult. Cell Rep. 2, 111–123 (2012).
On, T. et al. The evolutionary landscape of the chromatin modification machinery reveals lineage specific gains, expansions, and losses. Proteins 78, 2075–2089 (2010).
Fog, C.K., Jensen, K.T. & Lund, A.H. Chromatin-modifying proteins in cancer. APMIS 115, 1060–1089 (2007).
Kutter, C. et al. Rapid turnover of long noncoding RNAs and the evolution of gene expression. PLoS Genet. 8, e1002841 (2012).
Pollard, K.S. et al. An RNA gene expressed during cortical development evolved rapidly in humans. Nature 443, 167–172 (2006).
Pollard, K.S. et al. Forces shaping the fastest evolving regions in the human genome. PLoS Genet. 2, e168 (2006).
Ferrada, E. & Wagner, A. A comparison of genotype-phenotype maps for RNA and proteins. Biophys. J. 102, 1916–1925 (2012).
Amaral, P.P. et al. Complex architecture and regulated expression of the Sox2ot locus during vertebrate development. RNA 15, 2013–2027 (2009).
Willingham, A.T. et al. A strategy for probing the function of noncoding RNAs finds a repressor of NFAT. Science 309, 1570–1573 (2005).
Schuster, P., Fontana, W., Stadler, P.F. & Hofacker, I.L. From sequences to shapes and back: a case study in RNA secondary structures. Proc. Biol. Sci. 255, 279–284 (1994).
Saks, M.E., Sampson, J.R. & Abelson, J. Evolution of a transfer RNA gene through a point mutation in the anticodon. Science 279, 1665–1670 (1998).
Jorg, T., Martin, O.C. & Wagner, A. Neutral network sizes of biological RNA molecules can be computed and are not atypically small. BMC Bioinformatics 9, 464 (2008).
Yomo, T. et al. Properties of artificial proteins with random sequences. Ann. NY Acad. Sci. 864, 131–135 (1998).
Schultes, E.A., Spasic, A., Mohanty, U. & Bartel, D.P. Compact and ordered collapse of randomly generated RNA sequences. Nat. Struct. Mol. Biol. 12, 1130–1136 (2005).
Wagner, A. The role of robustness in phenotypic adaptation and innovation. Proc. Biol. Sci. 279, 1249–1258 (2012).
Hayden, E.J., Ferrada, E. & Wagner, A. Cryptic genetic variation promotes rapid evolutionary adaptation in an RNA enzyme. Nature 474, 92–95 (2011). This work is an empirical demonstration that variation in an RNA genotype network can drive rapid phenotypic adaptation.
Pigliucci, M. Is evolvability evolvable? Nat. Rev. Genet. 9, 75–82 (2008).
Acknowledgements
We thank the following funding sources: Australian National Health and Medical Research Council Australia Fellowship (631668 to J.S.M. and T.R.M.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Mercer, T., Mattick, J. Structure and function of long noncoding RNAs in epigenetic regulation. Nat Struct Mol Biol 20, 300–307 (2013). https://doi.org/10.1038/nsmb.2480
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2480
This article is cited by
-
GCNFORMER: graph convolutional network and transformer for predicting lncRNA-disease associations
BMC Bioinformatics (2024)
-
Mechanisms of norcantharidin against renal tubulointerstitial fibrosis
Pharmacological Reports (2024)
-
Cardiovascular complications of diabetes: role of non-coding RNAs in the crosstalk between immune and cardiovascular systems
Cardiovascular Diabetology (2023)
-
Genome instability-related LINC02577, LINC01133 and AC107464.2 are lncRNA prognostic markers correlated with immune microenvironment in pancreatic adenocarcinoma
BMC Cancer (2023)
-
LncRNA SNHG6 role in clinicopathological parameters in cancers
European Journal of Medical Research (2023)