Abstract
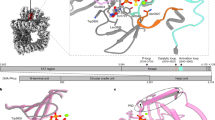
The DNA-repair enzyme Tdp2 resolves 5′-phosphotyrosyl DNA adducts and mediates resistance to anticancer drugs that target covalent topoisomerase–DNA complexes. Tdp2 also participates in key signaling pathways during development and tumorigenesis and cleaves a protein-RNA linkage during picornavirus replication. The crystal structure of zebrafish Tdp2 bound to DNA reveals a deep, narrow basic groove that selectively accommodates the 5′ end of single-stranded DNA in a stretched conformation. The crystal structure of the full-length Caenorhabditis elegans Tdp2 shows that this groove can also accommodate an acidic peptide stretch in vitro, with glutamate and aspartate side chains occupying the DNA backbone phosphate–binding sites. This extensive molecular mimicry suggests a potential mechanism for autoregulation and interaction of Tdp2 with phosphorylated proteins in signaling. Our study provides a framework to interrogate functions of Tdp2 and develop inhibitors for chemotherapeutic and antiviral applications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Pommier, Y., Leo, E., Zhang, H. & Marchand, C. DNA topoisomerases and their poisoning by anticancer and antibacterial drugs. Chem. Biol. 17, 421–433 (2010).
Wang, J.C. Cellular roles of DNA topoisomerases: a molecular perspective. Nat. Rev. Mol. Cell Biol. 3, 430–440 (2002).
Champoux, J.J. DNA topoisomerases: structure, function, and mechanism. Annu. Rev. Biochem. 70, 369–413 (2001).
Corbett, K.D. & Berger, J.M. Structure, molecular mechanisms, and evolutionary relationships in DNA topoisomerases. Annu. Rev. Biophys. Biomol. Struct. 33, 95–118 (2004).
Haffner, M.C. et al. Androgen-induced TOP2B-mediated double-strand breaks and prostate cancer gene rearrangements. Nat. Genet. 42, 668–675 (2010).
Deweese, J.E. & Osheroff, N. The DNA cleavage reaction of topoisomerase II: wolf in sheep's clothing. Nucleic Acids Res. 37, 738–748 (2009).
Nitiss, J.L. Targeting DNA topoisomerase II in cancer chemotherapy. Nat. Rev. Cancer 9, 338–350 (2009).
Wu, C.C. et al. Structural basis of type II topoisomerase inhibition by the anticancer drug etoposide. Science 333, 459–462 (2011).
Cortes Ledesma, F., El Khamisy, S.F., Zuma, M.C., Osborn, K. & Caldecott, K.W. A human 5′-tyrosyl DNA phosphodiesterase that repairs topoisomerase-mediated DNA damage. Nature 461, 674–678 (2009).
Zeng, Z., Cortes-Ledesma, F., El Khamisy, S.F. & Caldecott, K.W. Tdp2/TTRAP is the major 5′-tyrosyl DNA phosphodiesterase activity in vertebrate cells and is critical for cellular resistance to topoisomerase II-induced DNA damage. J. Biol. Chem. 286, 403–409 (2011).
Gao, R., Huang, S.Y., Marchand, C. & Pommier, Y. Biochemical characterization of human tyrosyl DNA phosphodiesterase 2 (Tdp2/TTRAP): a Mg2+/Mn2+-dependent phosphodiesterase specific for the repair of topoisomerase cleavage complexes. J. Biol. Chem. 287, 30842–30852 (2012).
Do, P.M. et al. Mutant p53 cooperates with ETS2 to promote etoposide resistance. Genes Dev. 26, 830–845 (2012).
Yang, S.W. et al. A eukaryotic enzyme that can disjoin dead-end covalent complexes between DNA and type I topoisomerases. Proc. Natl. Acad. Sci. USA 93, 11534–11539 (1996).
Raymond, A.C. & Burgin, A.B. Jr. Tyrosyl-DNA phosphodiesterase (Tdp1) (3′-phosphotyrosyl DNA phosphodiesterase). Methods Enzymol. 409, 511–524 (2006).
Davies, D.R., Interthal, H., Champoux, J.J. & Hol, W.G. Crystal structure of a transition state mimic for Tdp1 assembled from vanadate, DNA, and a topoisomerase I-derived peptide. Chem. Biol. 10, 139–147 (2003).
Pype, S. et al. TTRAP, a novel protein that associates with CD40, tumor necrosis factor (TNF) receptor-75 and TNF receptor-associated factors (TRAFs), and that inhibits nuclear factor-κB activation. J. Biol. Chem. 275, 18586–18593 (2000).
Esguerra, C.V. et al. Ttrap is an essential modulator of Smad3-dependent Nodal signaling during zebrafish gastrulation and left-right axis determination. Development 134, 4381–4393 (2007).
Pei, H. et al. EAPII interacts with ETS1 and modulates its transcriptional function. Oncogene 22, 2699–2709 (2003).
Li, C. et al. Oncogenic role of EAPII in lung cancer development and its activation of the MAPK-ERK pathway. Oncogene 30, 3802–3812 (2011).
Li, C., Sun, S.Y., Khuri, F.R. & Li, R. Pleiotropic functions of EAPII/TTRAP/Tdp2: cancer development, chemoresistance and beyond. Cell Cycle 10, 3274–3283 (2011).
Virgen-Slane, R. et al. An RNA virus hijacks an incognito function of a DNA repair enzyme. Proc. Natl. Acad. Sci. USA 109, 14634–14639 (2012).
Weston, S.A., Lahm, A. & Suck, D. X-ray structure of the DNase I-d(GGTATACC)2 complex at 2.3 Å resolution. J. Mol. Biol. 226, 1237–1256 (1992).
Mol, C.D., Izumi, T., Mitra, S. & Tainer, J.A. DNA-bound structures and mutants reveal abasic DNA binding by APE1 and DNA repair coordination. Nature 403, 451–456 (2000).
Adhikari, S. et al. Characterization of magnesium requirement of human 5′-tyrosyl DNA phosphodiesterase mediated reaction. BMC Res Notes 5, 134 (2012).
Tsutakawa, S.E. et al. Human flap endonuclease structures, DNA double-base flipping, and a unified understanding of the FEN1 superfamily. Cell 145, 198–211 (2011).
Adhikari, S. et al. Development of a novel assay for human tyrosyl DNA phosphodiesterase 2. Anal. Biochem. 416, 112–116 (2011).
Putnam, C.D., Hammel, M., Hura, G.L. & Tainer, J.A. X-ray solution scattering (SAXS) combined with crystallography and computation: defining accurate macromolecular structures, conformations and assemblies in solution. Q. Rev. Biophys. 40, 191–285 (2007).
Hura, G.L. et al. Robust, high-throughput solution structural analyses by small angle X-ray scattering (SAXS). Nat. Methods 6, 606–612 (2009).
Putnam, C.D. & Tainer, J.A. Protein mimicry of DNA and pathway regulation. DNA Repair (Amst.) 4, 1410–1420 (2005).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Adams, P.D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D Biol. Crystallogr. 58, 1948–1954 (2002).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Collaborative Computational Project. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
The PyMOL Molecular Graphics System. http://www.pymol.org/.
Baker, N.A., Sept, D., Joseph, S., Holst, M.J. & McCammon, J.A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl. Acad. Sci. USA 98, 10037–10041 (2001).
Brünger, A.T. et al. Crystallography & NMR System (CNS), A new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Classen, S. et al. Software for the high-throughput collection of SAXS data using an enhanced Blu-Ice/DCS control system. J. Synchrotron Radiat. 17, 774–781 (2010).
Konarev, P.V., Petoukhov, M.V., Volkov, V.V. & Svergun, D.I. ATSAS 2.1, a program package for small-angle scattering data analysis. J. Appl. Crystallogr. 39, 277–286 (2006).
Acknowledgements
We thank the beamline staff at Sector-24 of the Advanced Photon Source (APS) and J. Nix of the Molecular Biology Consortium at the Advanced Light Source (ALS) for help in data collection and H. Hiasa, D. Grandgenett and J. Lee for comments on the manuscript. Computer resources were provided by the Basic Sciences Computing Laboratory of the University of Minnesota Supercomputing Institute. The work conducted at the APS NE-CAT beamlines was supported by award RR15301 from the National Center for Research Resources at the US National Institutes of Health (NIH). Use of the APS, an Office of Science User Facility operated for the US Department of Energy (DOE) Office of Science by Argonne National Laboratory, was supported by the US DOE under contract no. DE-AC02-06CH11357. This work was supported by NIH grants GM095558 and AI087098 (H.A.), the Center for Cancer Research (Z01 BC 006150-19), the Intramural Program of the US National Cancer Institute (R.G. and Y.P.) and NCI P01 CA092584 and GM046312 (J.A.T.). SAXS data were collected at the ALS SIBYLS beamline 12.3.1 supported by the IDAT DOE program DE-AC02-05CH11231 and by NIH R01GM105404.
Author information
Authors and Affiliations
Contributions
K.K. purified the proteins and performed the activity assay on a substrate mimic. K.K., K.S. and H.A. crystallized the proteins. K.S. collected X-ray diffraction data and determined all crystal structures. R.G. and Y.P. examined enzyme activities on the 5′-phosphotyrosyl DNA substrate. S.E.T. and J.A.T. collected and analyzed the SAXS data. H.A. wrote the manuscript. All authors contributed in editing and figure preparation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 (PDF 3534 kb)
Rights and permissions
About this article
Cite this article
Shi, K., Kurahashi, K., Gao, R. et al. Structural basis for recognition of 5′-phosphotyrosine adducts by Tdp2. Nat Struct Mol Biol 19, 1372–1377 (2012). https://doi.org/10.1038/nsmb.2423
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2423
This article is cited by
-
Inactivating TDP2 missense mutation in siblings with congenital abnormalities reminiscent of fanconi anemia
Human Genetics (2023)
-
Molecular mechanisms of topoisomerase 2 DNA–protein crosslink resolution
Cellular and Molecular Life Sciences (2020)
-
High-fidelity DNA replication in Mycobacterium tuberculosis relies on a trinuclear zinc center
Nature Communications (2017)
-
Topoisomerase-mediated chromosomal break repair: an emerging player in many games
Nature Reviews Cancer (2015)
-
TDP2 protects transcription from abortive topoisomerase activity and is required for normal neural function
Nature Genetics (2014)