Abstract
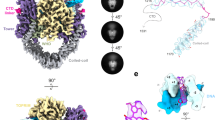
Type IIA topoisomerases control DNA supercoiling and disentangle chromosomes through a complex ATP-dependent strand-passage mechanism. Although a general framework exists for type IIA topoisomerase function, the architecture of the full-length enzyme has remained undefined. Here we present the structure of a fully catalytic Saccharomyces cerevisiae topoisomerase II homodimer complexed with DNA and a nonhydrolyzable ATP analog. The enzyme adopts a domain-swapped configuration wherein the ATPase domain of one protomer sits atop the nucleolytic region of its partner subunit. This organization produces an unexpected interaction between bound DNA and a conformational transducing element in the ATPase domain, which we show is critical for both DNA-stimulated ATP hydrolysis and global topoisomerase activity. Our data indicate that the ATPase domains pivot about each other to ensure unidirectional strand passage and that this state senses bound DNA to promote ATP turnover and enzyme reset.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Bustamante, C., Cheng, W. & Mejia, Y.X. Revisiting the central dogma one molecule at a time. Cell 144, 480–497 (2011).
Schoeffler, A.J. & Berger, J.M. DNA topoisomerases: harnessing and constraining energy to govern chromosome topology. Q. Rev. Biophys. 41, 41–101 (2008).
Roca, J. & Wang, J.C. The capture of a DNA double helix by an ATP-dependent protein clamp: a key step in DNA transport by type II DNA topoisomerases. Cell 71, 833–840 (1992).
Roca, J., Berger, J.M., Harrison, S.C. & Wang, J.C. DNA transport by a type II topoisomerase: direct evidence for a two-gate mechanism. Proc. Natl. Acad. Sci. USA 93, 4057–4062 (1996).
Wigley, D.B., Davies, G.J., Dodson, E.J., Maxwell, A. & Dodson, G. Crystal structure of an N-terminal fragment of the DNA gyrase B protein. Nature 351, 624–629 (1991).
Berger, J.M., Gamblin, S.J., Harrison, S.C. & Wang, J.C. Structure and mechanism of DNA topoisomerase II. Nature 379, 225–232 (1996).
Morais Cabral, J.H. et al. Crystal structure of the breakage-reunion domain of DNA gyrase. Nature 388, 903–906 (1997).
Roca, J. & Wang, J.C. DNA transport by a type II DNA topoisomerase: evidence in favor of a two-gate mechanism. Cell 77, 609–616 (1994).
Worland, S.T. & Wang, J.C. Inducible overexpression, purification, and active site mapping of DNA topoisomerase II from the yeast Saccharomyces cerevisiae. J. Biol. Chem. 264, 4412–4416 (1989).
Tse, Y.C., Kirkegaard, K. & Wang, J.C. Covalent bonds between protein and DNA. Formation of phosphotyrosine linkage between certain DNA topoisomerases and DNA. J. Biol. Chem. 255, 5560–5565 (1980).
Aravind, L., Leipe, D.D. & Koonin, E.V. Toprim–a conserved catalytic domain in type IA and II topoisomerases, DnaG-type primases, OLD family nucleases and RecR proteins. Nucleic Acids Res. 26, 4205–4213 (1998).
Lindsley, J.E. & Wang, J.C. Study of allosteric communication between protomers by immunotagging. Nature 361, 749–750 (1993).
Gellert, M., Fisher, L.M. & O'Dea, M.H. DNA gyrase: purification and catalytic properties of a fragment of gyrase B protein. Proc. Natl. Acad. Sci. USA 76, 6289–6293 (1979).
Brown, P.O., Peebles, C.L. & Cozzarelli, N.R. A topoisomerase from Escherichia coli related to DNA gyrase. Proc. Natl. Acad. Sci. USA 76, 6110–6114 (1979).
Williams, N.L. & Maxwell, A. Locking the DNA gate of DNA gyrase: investigating the effects on DNA cleavage and ATP hydrolysis. Biochemistry 38, 14157–14164 (1999).
Williams, N.L. & Maxwell, A. Probing the two-gate mechanism of DNA gyrase using cysteine cross-linking. Biochemistry 38, 13502–13511 (1999).
Classen, S., Olland, S. & Berger, J.M. Structure of the topoisomerase II ATPase region and its mechanism of inhibition by the chemotherapeutic agent ICRF-187. Proc. Natl. Acad. Sci. USA 100, 10629–10634 (2003).
Mizuuchi, K., O'Dea, M.H. & Gellert, M. DNA gyrase: subunit structure and ATPase activity of the purified enzyme. Proc. Natl. Acad. Sci. USA 75, 5960–5963 (1978).
Lindsley, J.E. & Wang, J.C. On the coupling between ATP usage and DNA transport by yeast DNA topoisomerase II. J. Biol. Chem. 268, 8096–8104 (1993).
Osheroff, N., Shelton, E.R. & Brutlag, D.L. DNA topoisomerase II from Drosophila melanogaster. Relaxation of supercoiled DNA. J. Biol. Chem. 258, 9536–9543 (1983).
Harkins, T.T., Lewis, T.J. & Lindsley, J.E. Pre-steady-state analysis of ATP hydrolysis by Saccharomyces cerevisiae DNA topoisomerase II. 2. Kinetic mechanism for the sequential hydrolysis of two ATP. Biochemistry 37, 7299–7312 (1998).
Dutta, R. & Inouye, M. GHKL, an emergent ATPase/kinase superfamily. Trends Biochem. Sci. 25, 24–28 (2000).
Laponogov, I. et al. Structural basis of gate-DNA breakage and resealing by type II topoisomerases. PLoS ONE 5, e11338 (2010).
Laponogov, I. et al. Structural insight into the quinolone-DNA cleavage complex of type IIA topoisomerases. Nat. Struct. Mol. Biol. 16, 667–669 (2009).
Wohlkonig, A. et al. Structural basis of quinolone inhibition of type IIA topoisomerases and target-mediated resistance. Nat. Struct. Mol. Biol. 17, 1152–1153 (2010).
Bax, B.D. et al. Type IIA topoisomerase inhibition by a new class of antibacterial agents. Nature 466, 935–940 (2010).
Schmidt, B.H., Burgin, A.B., Deweese, J.E., Osheroff, N. & Berger, J.M. A novel and unified two-metal mechanism for DNA cleavage by type II and IA topoisomerases. Nature 465, 641–644 (2010).
Wu, C.C. et al. Structural basis of type II topoisomerase inhibition by the anticancer drug etoposide. Science 333, 459–462 (2011).
Wendorff, T.J., Schmidt, B.H., Heslop, P., Austin, C.A. & Berger, J.M. The structure of DNA-bound human topoisomerase II alpha: conformational mechanisms for coordinating inter-subunit interactions with DNA cleavage. J. Mol. Biol. published online, doi:10.1016/j.jmb.2012.07.014 (25 July 2012).
Kampranis, S.C., Bates, A.D. & Maxwell, A. A model for the mechanism of strand passage by DNA gyrase. Proc. Natl. Acad. Sci. USA 96, 8414–8419 (1999).
Schultz, P., Olland, S., Oudet, P. & Hancock, R. Structure and conformational changes of DNA topoisomerase II visualized by electron microscopy. Proc. Natl. Acad. Sci. USA 93, 5936–5940 (1996).
Lamour, V., Hoermann, L., Jeltsch, J.M., Oudet, P. & Moras, D. An open conformation of the Thermus thermophilus gyrase B ATP-binding domain. J. Biol. Chem. 277, 18947–18953 (2002).
Brino, L. et al. Dimerization of Escherichia coli DNA-gyrase B provides a structural mechanism for activating the ATPase catalytic center. J. Biol. Chem. 275, 9468–9475 (2000).
Wei, H., Ruthenburg, A.J., Bechis, S.K. & Verdine, G.L. Nucleotide-dependent domain movement in the ATPase domain of a human type IIA DNA topoisomerase. J. Biol. Chem. 280, 37041–37047 (2005).
Bjergbaek, L. et al. Communication between the ATPase and cleavage/religation domains of human topoisomerase IIα. J. Biol. Chem. 275, 13041–13048 (2000).
Liu, L.F., Liu, C.C. & Alberts, B.M. Type II DNA topoisomerases: enzymes that can unknot a topologically knotted DNA molecule via a reversible double-strand break. Cell 19, 697–707 (1980).
Marini, J.C., Miller, K.G. & Englund, P.T. Decatenation of kinetoplast DNA by topoisomerases. J. Biol. Chem. 255, 4976–4979 (1980).
Osheroff, N. & Zechiedrich, E.L. Calcium-promoted DNA cleavage by eukaryotic topoisomerase II: trapping the covalent enzyme-DNA complex in an active form. Biochemistry 26, 4303–4309 (1987).
Lindsley, J.E. Use of a real-time, coupled assay to measure the ATPase activity of DNA topoisomerase II. Methods Mol. Biol. 95, 57–64 (2001).
Tamura, J.K. & Gellert, M. Characterization of the ATP binding site on Escherichia coli DNA gyrase. Affinity labeling of Lys-103 and Lys-110 of the B subunit by pyridoxal 5′-diphospho-5′-adenosine. J. Biol. Chem. 265, 21342–21349 (1990).
Maxwell, A. & Gellert, M. The DNA dependence of the ATPase activity of DNA gyrase. J. Biol. Chem. 259, 14472–14480 (1984).
Hammonds, T.R. & Maxwell, A. The DNA dependence of the ATPase activity of human DNA topoisomerase IIα. J. Biol. Chem. 272, 32696–32703 (1997).
West, K.L., Turnbull, R.M., Willmore, E., Lakey, J.H. & Austin, C.A. Characterisation of the DNA-dependent ATPase activity of human DNA topoisomerase IIbeta: mutation of Ser165 in the ATPase domain reduces the ATPase activity and abolishes the in vivo complementation ability. Nucleic Acids Res. 30, 5416–5424 (2002).
Mueller-Planitz, F. & Herschlag, D. DNA topoisomerase II selects DNA cleavage sites based on reactivity rather than binding affinity. Nucleic Acids Res. 35, 3764–3773 (2007).
Shiozaki, K. & Yanagida, M. Functional dissection of the phosphorylated termini of fission yeast DNA topoisomerase II. J. Cell Biol. 119, 1023–1036 (1992).
Cardenas, M.E., Dang, Q., Glover, C.V. & Gasser, S.M. Casein kinase II phosphorylates the eukaryote-specific C-terminal domain of topoisomerase II in vivo. EMBO J. 11, 1785–1796 (1992).
McClendon, A.K. et al. Bimodal recognition of DNA geometry by human topoisomerase II α: preferential relaxation of positively supercoiled DNA requires elements in the C-terminal domain. Biochemistry 47, 13169–13178 (2008).
Caron, P.R., Watt, P. & Wang, J.C. The C-terminal domain of Saccharomyces cerevisiae DNA topoisomerase II. Mol. Cell. Biol. 14, 3197–3207 (1994).
Roca, J. The path of the DNA along the dimer interface of topoisomerase II. J. Biol. Chem. 279, 25783–25788 (2004).
Baird, C.L., Harkins, T.T., Morris, S.K. & Lindsley, J.E. Topoisomerase II drives DNA transport by hydrolyzing one ATP. Proc. Natl. Acad. Sci. USA 96, 13685–13690 (1999).
Osheroff, N. Eukaryotic topoisomerase II. Characterization of enzyme turnover. J. Biol. Chem. 261, 9944–9950 (1986).
Tingey, A.P. & Maxwell, A. Probing the role of the ATP-operated clamp in the strand-passage reaction of DNA gyrase. Nucleic Acids Res. 24, 4868–4873 (1996).
Li, W. & Wang, J.C. Footprinting of yeast DNA topoisomerase II lysyl side chains involved in substrate binding and interdomainal interactions. J. Biol. Chem. 272, 31190–31195 (1997).
Choudhary, C. et al. Lysine acetylation targets protein complexes and co-regulates major cellular functions. Science 325, 834–840 (2009).
Bates, A.D., Berger, J.M. & Maxwell, A. The ancestral role of ATP hydrolysis in type II topoisomerases: prevention of DNA double-strand breaks. Nucleic Acids Res. 39, 6327–6339 (2011).
Ali, M.M. et al. Crystal structure of an Hsp90-nucleotide-p23/Sba1 closed chaperone complex. Nature 440, 1013–1017 (2006).
Gellert, M., Mizuuchi, K., O'Dea, M.H. & Nash, H.A. DNA gyrase: an enzyme that introduces superhelical turns into DNA. Proc. Natl. Acad. Sci. USA 73, 3872–3876 (1976).
Lee, S. et al. DNA cleavage and opening reactions of human topoisomerase IIα are regulated via Mg2+-mediated dynamic bending of gate-DNA. Proc. Natl. Acad. Sci. USA 109, 2925–2930 (2012).
Skouboe, C. et al. A human topoisomerase II α heterodimer with only one ATP binding site can go through successive catalytic cycles. J. Biol. Chem. 278, 5768–5774 (2003).
Göttler, T. & Klostermeier, D. Dissection of the nucleotide cycle of B. subtilis DNA gyrase and its modulation by DNA. J. Mol. Biol. 367, 1392–1404 (2007).
Deweese, J.E., Burgin, A.B. & Osheroff, N. Using 3′-bridging phosphorothiolates to isolate the forward DNA cleavage reaction of human topoisomerase IIα. Biochemistry 47, 4129–4140 (2008).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
The PyMOL Molecular Graphics System. Version 1.5.0.1 Schrödinger, LLC.
Acknowledgements
The authors thank A. Burgin, at Emerald BioStructures, Bainbridge Island, Washington, USA, for synthesizing the phosphorothioamidite reagent, as well as members of the Berger Lab for helpful discussions. This work was supported by a US National Institutes of Health training grant (GM08295 to B.H.S.) and the US National Cancer Institute (CA077373 to J.M.B.) and the US National Institutes of Health (GM33944 to N.O.).
Author information
Authors and Affiliations
Contributions
B.H.S., J.M.B. and N.O. designed the experiments. B.H.S. performed all of the experiments. B.H.S. and J.M.B. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 1009 kb)
Rights and permissions
About this article
Cite this article
Schmidt, B., Osheroff, N. & Berger, J. Structure of a topoisomerase II–DNA–nucleotide complex reveals a new control mechanism for ATPase activity. Nat Struct Mol Biol 19, 1147–1154 (2012). https://doi.org/10.1038/nsmb.2388
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2388
This article is cited by
-
Etoposide promotes DNA loop trapping and barrier formation by topoisomerase II
Nature Chemical Biology (2023)
-
Evybactin is a DNA gyrase inhibitor that selectively kills Mycobacterium tuberculosis
Nature Chemical Biology (2022)
-
Molecular mechanisms of topoisomerase 2 DNA–protein crosslink resolution
Cellular and Molecular Life Sciences (2020)
-
Dynamic coupling between conformations and nucleotide states in DNA gyrase
Nature Chemical Biology (2018)
-
Structural insights into the gating of DNA passage by the topoisomerase II DNA-gate
Nature Communications (2018)