Abstract
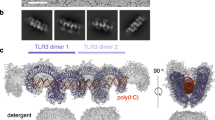
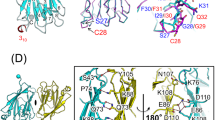
RP105–MD-1 modulates the TLR4–MD-2-mediated, innate immune response against bacterial lipopolysaccharide (LPS). The crystal structure of the bovine 1:1 RP105–MD-1 complex bound to a putative endogenous lipid at 2.9 Å resolution shares a similar overall architecture to its homolog TLR4–MD-2 but assembles into an unusual 2:2 homodimer that differs from any other known TLR-ligand assembly. The homodimer is assembled in a head-to-head orientation that juxtaposes the N-terminal leucine-rich repeats (LRRs) of the two RP105 chains, rather than the usual tail-to-tail configuration of C-terminal LRRs in ligand-activated TLR dimers, such as TLR1–TRL2, TLR2–TLR6, TLR3–TLR3 and TLR4–TLR4. Another unusual interaction is mediated by an RP105-specific asparagine-linked glycan, which wedges MD-1 into the co-receptor binding concavity on RP105. This unique mode of assembly represents a new paradigm for TLR complexes and suggests a molecular mechanism for regulating LPS responses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Bell, J.K. et al. Leucine-rich repeats and pathogen recognition in Toll-like receptors. Trends Immunol. 24, 528–533 (2003).
Takeda, K., Kaisho, T. & Akira, S. Toll-like receptors. Annu. Rev. Immunol. 21, 335–376 (2003).
Proell, M., Riedl, S.J., Fritz, J.H., Rojas, A.M. & Schwarzenbacher, R. The Nod-like receptor (NLR) family: a tale of similarities and differences. PLoS ONE 3, e2119 (2008).
Nakhaei, P., Genin, P., Civas, A. & Hiscott, J. RIG-I-like receptors: sensing and responding to RNA virus infection. Semin. Immunol. 21, 215–222 (2009).
Kim, H.M. et al. Crystal structure of the TLR4-MD-2 complex with bound endotoxin antagonist Eritoran. Cell 130, 906–917 (2007).
Jin, M.S. et al. Crystal structure of the TLR1–TLR2 heterodimer induced by binding of a tri-acylated lipopeptide. Cell 130, 1071–1082 (2007).
Choe, J., Kelker, M.S. & Wilson, I.A. Crystal structure of human toll-like receptor 3 (TLR3) ectodomain. Science 309, 581–585 (2005).
Kang, J.Y. et al. Recognition of lipopeptide patterns by Toll-like receptor 2-Toll-like receptor 6 heterodimer. Immunity 31, 873–884 (2009).
Bell, J.K. et al. The molecular structure of the Toll-like receptor 3 ligand-binding domain. Proc. Natl. Acad. Sci. USA 102, 10976–10980 (2005).
Park, B.S. et al. The structural basis of lipopolysaccharide recognition by the TLR4-MD-2 complex. Nature 458, 1191–1195 (2009).
Liu, L. et al. Structural basis of toll-like receptor 3 signaling with double-stranded RNA. Science 320, 379–381 (2008).
Shimazu, R. et al. MD-2, a molecule that confers lipopolysaccharide responsiveness on Toll-like receptor 4. J. Exp. Med. 189, 1777–1782 (1999).
Nagai, Y. et al. Essential role of MD-2 in LPS responsiveness and TLR4 distribution. Nat. Immunol. 3, 667–672 (2002).
Poltorak, A. et al. Defective LPS signaling in C3H/HeJ and C57BL/10ScCr mice: mutations in Tlr4 gene. Science 282, 2085–2088 (1998).
Ohto, U., Fukase, K., Miyake, K. & Satow, Y. Crystal structures of human MD-2 and its complex with antiendotoxic lipid IVa. Science 316, 1632–1634 (2007).
Mullarkey, M. et al. Inhibition of endotoxin response by e5564, a novel Toll-like receptor 4-directed endotoxin antagonist. J. Pharmacol. Exp. Ther. 304, 1093–1102 (2003).
Miyake, K., Yamashita, Y., Ogata, M., Sudo, T. & Kimoto, M. RP105, a novel B cell surface molecule implicated in B cell activation, is a member of the leucine-rich repeat protein family. J. Immunol. 154, 3333–3340 (1995).
Miura, Y. et al. Molecular cloning of a human RP105 homologue and chromosomal localization of the mouse and human RP105 genes (Ly64 and LY64). Genomics 38, 299–304 (1996).
Fugier-Vivier, I. et al. Molecular cloning of human RP105. Eur. J. Immunol. 27, 1824–1827 (1997).
Divanovic, S. et al. Negative regulation of Toll-like receptor 4 signaling by the Toll-like receptor homolog RP105. Nat. Immunol. 6, 571–578 (2005).
Miura, Y. et al. RP105 is associated with MD-1 and transmits an activation signal in human B cells. Blood 92, 2815–2822 (1998).
Miyake, K. et al. Mouse MD-1, a molecule that is physically associated with RP105 and positively regulates its expression. J. Immunol. 161, 1348–1353 (1998).
Nagai, Y. et al. Requirement for MD-1 in cell surface expression of RP105/CD180 and B-cell responsiveness to lipopolysaccharide. Blood 99, 1699–1705 (2002).
Nagai, Y. et al. The radioprotective 105/MD-1 complex links TLR2 and TLR4/MD-2 in antibody response to microbial membranes. J. Immunol. 174, 7043–7049 (2005).
Ogata, H. et al. The toll-like receptor protein RP105 regulates lipopolysaccharide signaling in B cells. J. Exp. Med. 192, 23–29 (2000).
Divanovic, S. et al. Regulation of TLR4 signaling and the host interface with pathogens and danger: the role of RP105. J. Leukoc. Biol. 82, 265–271 (2007).
Divanovic, S. et al. Inhibition of TLR-4/MD-2 signaling by RP105/MD-1. J. Endotoxin Res. 11, 363–368 (2005).
Blumenthal, A. et al. RP105 facilitates macrophage activation by Mycobacterium tuberculosis lipoproteins. Cell Host Microbe 5, 35–46 (2009).
Hindle, K.L., Bella, J. & Lovell, S.C. Quantitative analysis and prediction of curvature in leucine-rich repeat proteins. Proteins 77, 342–358 (2009).
Bublitz, M. et al. Crystal structure and standardized geometric analysis of InlJ, a listerial virulence factor and leucine-rich repeat protein with a novel cysteine ladder. J. Mol. Biol. 378, 87–96 (2008).
Enkhbayar, P., Kamiya, M., Osaki, M., Matsumoto, T. & Matsushima, N. Structural principles of leucine-rich repeat (LRR) proteins. Proteins 54, 394–403 (2004).
Harada, H., Ohto, U. & Satow, Y. Crystal structure of mouse MD-1 with endogenous phospholipid bound in its cavity. J. Mol. Biol. 400, 838–846 (2010).
Yoon, S.I., Hong, M., Han, G.W. & Wilson, I.A. Crystal structure of soluble MD-1 and its interaction with lipid IVa. Proc. Natl. Acad. Sci. USA 107, 10990–10995 (2010).
Altmann, F., Staudacher, E., Wilson, I.B. & Marz, L. Insect cells as hosts for the expression of recombinant glycoproteins. Glycoconj. J. 16, 109–123 (1999).
Aeed, P.A. & Elhammer, A.P. Glycosylation of recombinant prorenin in insect cells: the insect cell line Sf9 does not express the mannose 6-phosphate recognition signal. Biochemistry 33, 8793–8797 (1994).
Lawrence, M.C. & Colman, P.M. Shape complementarity at protein/protein interfaces. J. Mol. Biol. 234, 946–950 (1993).
Walter, T.S. et al. Lysine methylation as a routine rescue strategy for protein crystallization. Structure 14, 1617–1622 (2006).
Jentoft, N. & Dearborn, D.G. Protein labeling by reductive alkylation. Methods Enzymol. 91, 570–579 (1983).
Jin, M.S. & Lee, J.O. Application of hybrid LRR technique to protein crystallization. BMB Rep 41, 353–357 (2008).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Sheldrick, G.M. A short history of SHELX. Acta Crystallogr. A 64, 112–122 (2008).
Cowtan, K. ′dm′: an automated procedure for phase improvement by density modification. Joint CCP4 and ESF-EACBM Newslett. on Protein Crystallogr. 31, 34–38 (1994).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crysttallogr. 40, 658–674 (2007).
Acknowledgements
We thank R.L. Stanfield (The Scripps Research Institute) and Y.S. Choo (Sanford-Burnham Medical Research Institute) for critical comments on the manuscript, R.L. Stanfield (The Scripps Research Institute), H. Tien and D. Marciano (The Joint Center for Structural Genomics) for automated crystal screening, and X. Dai and M.A. Elsliger (The Scripps Research Institute) for expert technical assistance. The work was supported by US National Institutes of Health grant AI042266 (to I.A.W.) and the Skaggs Institute for Chemical Biology. X-ray diffraction datasets were collected at the Stanford Synchrotron Radiation Lightsource beamline 9-2 and the Advanced Photon Source beamline 23ID-B. This is manuscript no. 20749 from The Scripps Research Institute.
Author information
Authors and Affiliations
Contributions
S.Y. and I.A.W. designed experiments. S.Y. and M.H. conducted experiments. S.Y., M.H. and I.A.W. analyzed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 5704 kb)
Rights and permissions
About this article
Cite this article
Yoon, Si., Hong, M. & Wilson, I. An unusual dimeric structure and assembly for TLR4 regulator RP105–MD-1. Nat Struct Mol Biol 18, 1028–1035 (2011). https://doi.org/10.1038/nsmb.2106
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2106
This article is cited by
-
The role of CD180 in hematological malignancies and inflammatory disorders
Molecular Medicine (2023)
-
Expression of CD180 in common B cell neoplasms using immunohistochemistry
Journal of Hematopathology (2020)
-
Novel Protective Role of Myeloid Differentiation 1 in Pathological Cardiac Remodelling
Scientific Reports (2017)
-
Secreted tryptophanyl-tRNA synthetase as a primary defence system against infection
Nature Microbiology (2016)
-
Recognition of lipopolysaccharide pattern by TLR4 complexes
Experimental & Molecular Medicine (2013)