Abstract
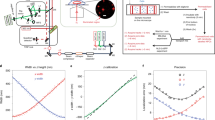
The nuclear pore complex (NPC) perforates the nuclear envelope to facilitate selective transport between nucleus and cytoplasm. The NPC is composed of multiple copies of ∼30 different proteins, termed nucleoporins, whose arrangement within the NPC is an important unsolved puzzle in structural biology. Various alternative models for NPC architecture have been proposed but not tested experimentally in intact NPCs. We present a method using polarized fluorescence microscopy to investigate nucleoporin orientation in live yeast and mammalian cells. Our results support an arrangement of both yeast Nic96 and human Nup133–Nup107 in which their long axes are approximately parallel to the nuclear envelope plane. The method we developed can complement X-ray crystallography and electron microscopy to generate a high-resolution map of the entire NPC, and may be able to monitor nucleoporin rearrangements during nucleocytoplasmic transport and NPC assembly. This strategy can also be adapted for other macromolecular machines.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Lim, R.Y., Ullman, K.S. & Fahrenkrog, B. Biology and biophysics of the nuclear pore complex and its components. Int. Rev. Cell Mol. Biol. 267, 299–342 (2008).
Beck, M., Lucic, V., Forster, F., Baumeister, W. & Medalia, O. Snapshots of nuclear pore complexes in action captured by cryo-electron tomography. Nature 449, 611–615 (2007).
Brohawn, S.G., Partridge, J.R., Whittle, J.R. & Schwartz, T.U. The nuclear pore complex has entered the atomic age. Structure 17, 1156–1168 (2009).
Kampmann, M. & Blobel, G. Three-dimensional structure and flexibility of a membrane-coating module of the nuclear pore complex. Nat. Struct. Mol. Biol. 16, 782–788 (2009).
Alber, F. et al. The molecular architecture of the nuclear pore complex. Nature 450, 695–701 (2007).
Schrader, N. et al. Structural basis of the nic96 subcomplex organization in the nuclear pore channel. Mol. Cell 29, 46–55 (2008).
Brohawn, S.G. & Schwartz, T.U. Molecular architecture of the Nup84–Nup145C–Sec13 edge element in the nuclear pore complex lattice. Nat. Struct. Mol. Biol. 16, 1173–1177 (2009).
Brohawn, S.G., Leksa, N.C., Spear, E.D., Rajashankar, K.R. & Schwartz, T.U. Structural evidence for common ancestry of the nuclear pore complex and vesicle coats. Science 322, 1369–1373 (2008).
Seo, H.S. et al. Structural and functional analysis of Nup120 suggests ring formation of the Nup84 complex. Proc. Natl. Acad. Sci. USA 106, 14281–14286 (2009).
Debler, E.W. et al. A fence-like coat for the nuclear pore membrane. Mol. Cell 32, 815–826 (2008).
Hsia, K.C., Stavropoulos, P., Blobel, G. & Hoelz, A. Architecture of a coat for the nuclear pore membrane. Cell 131, 1313–1326 (2007).
Ajtai, K., Toft, D.J. & Burghardt, T.P. Path and extent of cross-bridge rotation during muscle contraction. Biochemistry 33, 5382–5391 (1994).
Axelrod, D. Carbocyanine dye orientation in red cell membrane studied by microscopic fluorescence polarization. Biophys. J. 26, 557–573 (1979).
Rocheleau, J.V., Edidin, M. & Piston, D.W. Intrasequence GFP in class I MHC molecules, a rigid probe for fluorescence anisotropy measurements of the membrane environment. Biophys. J. 84, 4078–4086 (2003).
Vrabioiu, A.M. & Mitchison, T.J. Structural insights into yeast septin organization from polarized fluorescence microscopy. Nature 443, 466–469 (2006).
Vrabioiu, A.M. & Mitchison, T.J. Symmetry of septin hourglass and ring structures. J. Mol. Biol. 372, 37–49 (2007).
Corrie, J.E. et al. Dynamic measurement of myosin light-chain-domain tilt and twist in muscle contraction. Nature 400, 425–430 (1999).
Mattheyses, A.L., Kampmann, M., Atkinson, C.E. & Simon, S.M. Fluorescence anisotropy reveals order and disorder of protein domains in the nuclear pore complex. Biophys. J. 99, 1706–1717 (2010).
Rosell, F.I. & Boxer, S.G. Polarized absorption spectra of green fluorescent protein single crystals: transition dipole moment directions. Biochemistry 42, 177–183 (2003).
Jeudy, S. & Schwartz, T.U. Crystal structure of nucleoporin Nic96 reveals a novel, intricate helical domain architecture. J. Biol. Chem. 282, 34904–34912 (2007).
Berke, I.C., Boehmer, T., Blobel, G. & Schwartz, T.U. Structural and functional analysis of Nup133 domains reveals modular building blocks of the nuclear pore complex. J. Cell Biol. 167, 591–597 (2004).
Boehmer, T., Jeudy, S., Berke, I.C. & Schwartz, T.U. Structural and functional studies of Nup107/Nup133 interaction and its implications for the architecture of the nuclear pore complex. Mol. Cell 30, 721–731 (2008).
Whittle, J.R. & Schwartz, T.U. Architectural nucleoporins Nup157/170 and Nup133 are structurally related and descend from a second ancestral element. J. Biol. Chem. 284, 28442–28452 (2009).
Choe, S. & Sun, S.X. The elasticity of α-helices. J. Chem. Phys. 122, 244912 (2005).
Devos, D. et al. Simple fold composition and modular architecture of the nuclear pore complex. Proc. Natl. Acad. Sci. USA 103, 2172–2177 (2006).
Melcák, I., Hoelz, A. & Blobel, G. Structure of Nup58/45 suggests flexible nuclear pore diameter by intermolecular sliding. Science 315, 1729–1732 (2007).
Shulga, N. et al. In vivo nuclear transport kinetics in Saccharomyces cerevisiae: a role for heat shock protein 70 during targeting and translocation. J. Cell Biol. 135, 329–339 (1996).
Brachmann, C.B. et al. Designer deletion strains derived from Saccharomyces cerevisiae S288C: a useful set of strains and plasmids for PCR-mediated gene disruption and other applications. Yeast 14, 115–132 (1998).
Sheff, M.A. & Thorn, K.S. Optimized cassettes for fluorescent protein tagging in Saccharomyces cerevisiae. Yeast 21, 661–670 (2004).
Sherman, F. Getting started with yeast. Methods Enzymol. 350, 3–41 (2002).
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Pettersen, E.F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Elsliger, M.A., Wachter, R.M., Hanson, G.T., Kallio, K. & Remington, S.J. Structural and spectral response of green fluorescent protein variants to changes in pH. Biochemistry 38, 5296–5301 (1999).
Maegawa, Y. et al. Structure of the catalytic nucleotide-binding subunit A of A-type ATP synthase from Pyrococcus horikoshii reveals a novel domain related to the peripheral stalk. Acta Crystallogr. D Biol. Crystallogr. 62, 483–488 (2006).
Diepholz, M. et al. A different conformation for EGC stator subcomplex in solution and in the assembled yeast V-ATPase: possible implications for regulatory disassembly. Structure 16, 1789–1798 (2008).
Shi, J., Blundell, T.L. & Mizuguchi, K. FUGUE: sequence-structure homology recognition using environment-specific substitution tables and structure-dependent gap penalties. J. Mol. Biol. 310, 243–257 (2001).
Acknowledgements
We thank G. Blobel (Rockefeller University, Howard Hughes Medical Institute) for his generous support of this work, partially carried out in his laboratory. We thank D. Johnson and D. Muzzey for comments on the manuscript and C. Lue for technical assistance. M.K. was supported by a Howard Hughes Medical Institute Predoctoral Fellowship. A.L.M. was supported by a Rockefeller University Women & Science Postdoctoral Fellowship. C.E.A., A.L.M. and S.M.S. were supported by US National Science Foundation grant BES-0620813 and National Institutes of Health grant R01 GM087977 to S.M.S.
Author information
Authors and Affiliations
Contributions
M.K., C.E.A. and A.L.M. did all experimentation, and M.K., C.E.A., A.L.M. and S.M.S. shared the design, analysis and writing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Table 1 and Supplementary Note (PDF 5939 kb)
Rights and permissions
About this article
Cite this article
Kampmann, M., Atkinson, C., Mattheyses, A. et al. Mapping the orientation of nuclear pore proteins in living cells with polarized fluorescence microscopy. Nat Struct Mol Biol 18, 643–649 (2011). https://doi.org/10.1038/nsmb.2056
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2056
This article is cited by
-
Extending fluorescence anisotropy to large complexes using reversibly switchable proteins
Nature Biotechnology (2023)
-
Polarization modulation with optical lock-in detection reveals universal fluorescence anisotropy of subcellular structures in live cells
Light: Science & Applications (2022)
-
Polarisation optics for biomedical and clinical applications: a review
Light: Science & Applications (2021)
-
Quantitative linear dichroism imaging of molecular processes in living cells made simple by open software tools
Communications Biology (2021)
-
Super-resolution imaging of fluorescent dipoles via polarized structured illumination microscopy
Nature Communications (2019)