Abstract
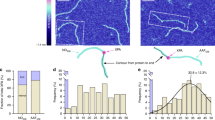
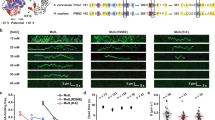
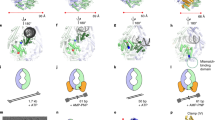
Single-molecule trajectory analysis has suggested DNA repair proteins may carry out a one-dimensional (1D) search on naked DNA encompassing >10,000 nucleotides. Organized cellular DNA (chromatin) presents substantial barriers to such lengthy searches. Using dynamic single-molecule fluorescence resonance energy transfer, we determined that the mismatch repair (MMR) initiation protein MutS forms a transient clamp that scans duplex DNA for mismatched nucleotides by 1D diffusion for 1 s (~700 base pairs) while in continuous rotational contact with the DNA. Mismatch identification provokes ATP binding (3 s) that induces distinctly different MutS sliding clamps with unusual stability on DNA (~600 s), which may be released by adjacent single-stranded DNA (ssDNA). These observations suggest that ATP transforms short-lived MutS lesion scanning clamps into highly stable MMR signaling clamps that are capable of competing with chromatin and recruiting MMR machinery, yet are recycled by ssDNA excision tracts.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
Change history
13 February 2011
In the version of this article initially published online, the x axis for the right-hand graph in Figure 5c and the x axis for the graph in Figure 5d should have read 'Dwell time (s)'. The error has been corrected in all versions of this article.
References
Kolodner, R.D. & Marsischky, G.T. Eukaryotic DNA mismatch repair. Curr. Opin. Genet. Dev. 9, 89–96 (1999).
Modrich, P. & Lahue, R. Mismatch repair in replication fidelity, genetic recombination, and cancer biology. Annu. Rev. Biochem. 65, 101–133 (1996).
Kolodner, R.D., Mendillo, M.L. & Putnam, C.D. Coupling distant sites in DNA during DNA mismatch repair. Proc. Natl. Acad. Sci. USA 104, 12953–12954 (2007).
Acharya, S., Foster, P.L., Brooks, P. & Fishel, R. The coordinated functions of the E. coli MutS and MutL proteins in mismatch repair. Mol. Cell 12, 233–246 (2003).
Gradia, S., Acharya, S. & Fishel, R. The role of mismatched nucleotides in activating the hMSH2-hMSH6 molecular switch. J. Biol. Chem. 275, 3922–3930 (2000).
Berg, O.G., Winter, R.B. & von Hippel, P.H. Diffusion-driven mechanisms of protein translocation on nucleic acids. 1. Models and theory. Biochemistry 20, 6929–6948 (1981).
Riggs, A.D., Bourgeois, S. & Cohn, M. The lac repressor-operator interaction. 3. Kinetic studies. J. Mol. Biol. 53, 401–417 (1970).
Blainey, P.C., van Oijen, A.M., Banerjee, A., Verdine, G.L. & Xie, X.S. A base-excision DNA-repair protein finds intrahelical lesion bases by fast sliding in contact with DNA. Proc. Natl. Acad. Sci. USA 103, 5752–5757 (2006).
Gorman, J. et al. Dynamic basis for one-dimensional DNA scanning by the mismatch repair complex Msh2–Msh6. Mol. Cell 28, 359–370 (2007).
Wolffe, A.P. & Guschin, D. Review: chromatin structural features and targets that regulate transcription. J. Struct. Biol. 129, 102–122 (2000).
Fox, R.F. Rectified Brownian movement in molecular and cell biology. Phys. Rev. E Stat. Phys. Plasmas Fluids Relat. Interdiscip. Topics 57, 2177–2203 (1998).
Fox, R.F. & Choi, M.H. Rectified Brownian motion and kinesin motion along microtubules. Phys. Rev. E 63, 051901 (2001).
Roy, R., Hohng, S. & Ha, T. A practical guide to single-molecule FRET. Nat. Methods 5, 507–516 (2008).
Biswas, I. & Hsieh, P. Identification and characterization of a thermostable MutS homolog from Thermus aquaticus. J. Biol. Chem. 271, 5040–5048 (1996).
Blanchard, S.C., Kim, H.D., Gonzalez, R.L., Jr., Puglisi, J.D. & Chu, S. tRNA dynamics on the ribosome during translation. Proc. Natl. Acad. Sci. USA 101, 12893–12898 (2004).
Murphy, M.C., Rasnik, I., Cheng, W., Lohman, T.M. & Ha, T. Probing single-stranded DNA conformational flexibility using fluorescence spectroscopy. Biophys. J. 86, 2530–2537 (2004).
Karran, P. & Hampson, R. Genomic instability and tolerance to alkylating agents. Cancer Surv. 28, 69–85 (1996).
Karran, P. & Marinus, M.G. Mismatch correction at O6-methylguanine residues in E. coli DNA. Nature 296, 868–869 (1982).
Blainey, P.C. et al. Nonspecifically bound proteins spin while diffusing along DNA. Nat. Struct. Mol. Biol. 16, 1224–1229 (2009).
Tessmer, I. et al. Mechanism of MutS searching for DNA mismatches and signaling repair. J. Biol. Chem. 283, 36646–36654 (2008).
Schofield, M.J., Nayak, S., Scott, T.H., Du, C. & Hsieh, P. Interaction of Escherichia coli MutS and MutL at a DNA mismatch. J. Biol. Chem. 276, 28291–28299 (2001).
Yang, Y., Sass, L.E., Du, C., Hsieh, P. & Erie, D.A. Determination of protein-DNA binding constants and specificities from statistical analyses of single molecules: MutS-DNA interactions. Nucleic Acids Res. 33, 4322–4334 (2005).
Abeliovich, H. An empirical extremum principle for the hill coefficient in ligand-protein interactions showing negative cooperativity. Biophys. J. 89, 76–79 (2005).
Lamers, M.H., Winterwerp, H.H. & Sixma, T.K. The alternating ATPase domains of MutS control DNA mismatch repair. EMBO J. 22, 746–756 (2003).
Gradia, S. et al. hMSH2-hMSH6 forms a hydrolysis-independent sliding clamp on mismatched DNA. Mol. Cell 3, 255–261 (1999).
Javaid, S. et al. Nucleosome remodeling by hMSH2-hMSH6. Mol. Cell 36, 1086–1094 (2009).
Laurence, T.A. et al. Motion of a DNA sliding clamp observed by single molecule fluorescence spectroscopy. J. Biol. Chem. 283, 22895–22906 (2008).
Mather, W.H. & Fox, R.F. Kinesin's biased stepping mechanism: amplification of neck linker zippering. Biophys. J. 91, 2416–2426 (2006).
Gradia, S., Acharya, S. & Fishel, R. The human mismatch recognition complex hMSH2–hMSH6 functions as a novel molecular switch. Cell 91, 995–1005 (1997).
Fishel, R. Mismatch repair, molecular switches, and signal transduction. Genes Dev. 12, 2096–2101 (1998).
Fishel, R. Signaling mismatch repair in cancer. Nat. Med. 5, 1239–1241 (1999).
Fishel, R. The selection for mismatch repair defects in hereditary nonpolyposis colorectal cancer: revising the mutator hypothesis. Cancer Res. 61, 7369–7374 (2001).
Hampel, H. et al. Screening for the Lynch syndrome (hereditary nonpolyposis colorectal cancer). N. Engl. J. Med. 352, 1851–1860 (2005).
Li, G.-W., Berg, O.G. & Elf, J. Effects of macromolecular crowding and DNA looping on gene regulation kinetics. Nat. Phys. 5, 294–297 (2009).
Constantin, N., Dzantiev, L., Kadyrov, F.A. & Modrich, P. Human mismatch repair: reconstitution of a nick-directed bidirectional reaction. J. Biol. Chem. 280, 39752–39761 (2005).
Lahue, R.S., Au, K.G. & Modrich, P. DNA mismatch correction in a defined system. Science 245, 160–164 (1989).
Blackwell, L.J., Martik, D., Bjornson, K.P., Bjornson, E.S. & Modrich, P. Nucleotide-promoted release of hMutS from heteroduplex DNA is consistent with an ATP-dependent translocation mechanism. J. Biol. Chem. 273, 32055–32062 (1998).
Lau, P.J. & Kolodner, R.D. Transfer of the MSH2.MSH6 complex from proliferating cell nuclear antigen to mispaired bases in DNA. J. Biol. Chem. 278, 14–17 (2003).
Shell, S.S., Putnam, C.D. & Kolodner, R.D. The N terminus of Saccharomyces cerevisiae Msh6 is an unstructured tether to PCNA. Mol. Cell 26, 565–578 (2007).
Mazurek, A., Johnson, C.N., Germann, M.W. & Fishel, R. Sequence context effect for hMSH2–hMSH6 mismatch–dependent activation. Proc. Natl. Acad. Sci. USA 106, 4177–4182 (2009).
Lamers, M.H. et al. The crystal structure of DNA mismatch repair protein MutS binding to a G x T mismatch. Nature 407, 711–717 (2000).
Obmolova, G., Ban, C., Hsieh, P. & Yang, W. Crystal structures of mismatch repair protein MutS and its complex with a substrate DNA. Nature 407, 703–710 (2000).
Warren, J.J. et al. Structure of the human MutSα DNA lesion recognition complex. Mol. Cell 26, 579–592 (2007).
Mendillo, M.L., Mazur, D.J. & Kolodner, R.D. Analysis of the interaction between the Saccharomyces cerevisiae MSH2–MSH6 and MLH1–PMS1 complexes with DNA using a reversible DNA end–blocking system. J. Biol. Chem. 280, 22245–22257 (2005).
Heo, S.D., Cho, M., Ku, J.K. & Ban, C. Steady–state ATPase activity of E. coli MutS modulated by its dissociation from heteroduplex DNA. Biochem. Biophys. Res. Commun. 364, 264–269 (2007).
Cho, M., Chung, S., Heo, S.D., Ku, J. & Ban, C. A simple fluorescent method for detecting mismatched DNAs using a MutS-fluorophore conjugate. Biosens. Bioelectron. 22, 1376–1381 (2007).
Rasnik, I., Myong, S., Cheng, W., Lohman, T.M. & Ha, T. DNA-binding orientation and domain conformation of the E. coli rep helicase monomer bound to a partial duplex junction: single-molecule studies of fluorescently labeled enzymes. J. Mol. Biol. 336, 395–408 (2004).
Joo, C. & Ha, T. Single-molecule FRET with total internal reflection microscopy. in Single-Molecule Techniques: A Laboratory Manual (eds. Selvin, P.R. & Ha, T.) 3–36 (Cold Spring Harbor Laboratory Press, 2008).
Acknowledgements
We thank J.-H. Park for helping with the FRET experiments. This work was supported by National Research Foundation of Korea grants funded by the Korean government (MEST) (no. 2008-0061211, no. 2009-351-C00118; C.J. postdoctoral fellowship), the Brain Korea 21 project, a POSTECH Basic Science Research Institute Grant (J.-B.L.), HRF-2008-314-C00218 (C.B.) and US National Institutes of Health grant CA067007 (R.F.).
Author information
Authors and Affiliations
Contributions
C.J., R.F. and J.-B.L. designed the experiments. C.B. and K.-M.S. contributed essential reagents. C.J. and W.-K.C. carried out single-molecule analysis. C.C. carried out bulk analysis. C.J., W.-K.C., T.-Y.Y., C.B., R.F. and J.-B.L. analyzed the data. C.J., J.-B.L. and R.F. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Tables 1 and 2 (PDF 1998 kb)
Rights and permissions
About this article
Cite this article
Jeong, C., Cho, WK., Song, KM. et al. MutS switches between two fundamentally distinct clamps during mismatch repair. Nat Struct Mol Biol 18, 379–385 (2011). https://doi.org/10.1038/nsmb.2009
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2009
This article is cited by
-
Evolutionary origin of germline pathogenic variants in human DNA mismatch repair genes
Human Genomics (2024)
-
DNA strand breaks and gaps target retroviral intasome binding and integration
Nature Communications (2023)
-
Tandem regulation of MutS activity by ATP and DNA during MMR initiation
Nature Structural & Molecular Biology (2022)
-
MutS functions as a clamp loader by positioning MutL on the DNA during mismatch repair
Nature Communications (2022)
-
MutS and MutL sliding clamps in DNA mismatch repair
Genome Instability & Disease (2022)